
20.5.2026A cell researcher's new tools
10.4.2026The Mystery of Brain Development
27.2.2026Modelling helps to identify a rare childhood neurological disorder caused by missense mutations of SynGAP1 protein
23.6.2025Too clean is unhealthy
8.5.2025Alkaloids derived from tree bark destroy cancer cells
27.3.2025Vinca alkaloids: Madagascar’s gift to cancer treatment
6.2.2025Genetic testing improves medication safety and effectiveness
26.12.2024The ComPatAI consortium uses large datasets to create an AI learning model for pathology
14.11.2024Microbiota affects the immune system
21.10.2024The skin’s wide range of microbiota improves the immune system
30.9.2024New drug targets from RNA-binding proteins
31.8.2024New machine learning method speeds up drug screening hundred-fold
22.7.2024Mapping the coffee genome to improve disease resistance
25.6.2024Why do some get the severe form of COVID-19?
30.5.2024An AI model that understands health data warns of future diseases
29.4.2024An infrastructure for genomic data
1.4.2024European research community preparing for next pandemic
8.3.2024Evolutionary dynamics of viruses and other microbes affect human health
2.3.2024A million European genomes
20.2.2024Efficient transfer and analysis of biological image data through web interfaces
23.1.2024Improving breast cancer treatment prognoses with liquid biopsy
15.12.2023The European Health Data Space: health data moves across borders for research purposes
16.11.2023New method for measuring gut microbiota
31.10.2023Purifying mining wastewater with plant-associated microbes
29.9.2023Artificial intelligence helps researchers find suitable drugs based on patient’s genetic data and cancer cell samples
1.9.2023Combining data from different sources for personalised treatment
15.8.2023Better treatments for leukaemia
10.6.2023MicroRNAs may reveal type 1 diabetes
16.5.2023Single-cell RNA sequencing enabling individual disease treatment
12.4.2023Tissue samples analysed with Sensitive Data (SD) services provide new information on celiac disease and other autoimmune diseases
20.3.2023DNA isolated from Baltic Sea sediment shedding light on climate change and biodiversity
27.2.2023Organoids grown from stem cells boost cancer research
19.12.2022Sensitive Data (SD) services for Research: with a few clicks a researcher can launch a personal secure computing environment
30.11.2022Microbiota in permafrost play an important role in climate change
20.10.2022Reusable, accurately described and high-quality data – tools created by the research community for agile data management
29.9.2022Gene sequencing used for study of structure and functioning of microbial communities in oceans
1.9.2022Antibiotic-resistant bacteria are a global problem
23.8.2022Personalised medicine against cancer and viruses
30.6.2022Studying the human microbiome is a key towards holistic understanding of our health
23.5.2022FINRISK: one of the world’s longest-running population survey time series
8.4.2022Combining biobank data with data from health registers enables research towards personalised treatment
3.3.2022Finnish research team sequences the genomes of thousands of individuals with diabetes to look for genetic risk factors
10.2.2022BIGPICTURE helps pathology go digital
30.12.2021Sensitive data infrastructure
23.11.2021In the future, an algorithm may diagnose glaucoma from fundus photos
26.10.2021Patient data creating better artificial intelligence models
15.9.2021Teaching an algorithm to identify cancer from sequence data
3.12.2020Efficient processing and sharing of data improving disease diagnosis and treatment
10.11.2020Bioinformatics to revolutionise healthcare: Efficient data processing speeds up diagnoses and enables personalised drug treatments
27.8.2020Tissue samples into digital images, interpreted by artificial intelligence
9.6.2020Digital pathology speeds up diagnosis
18.5.2020Searching markers for breast cancer by machine learning
8.4.2020Metabolomics measures and analyses metabolic changes caused by illness, diet or medication
1.3.2020Deep learning algorithms help in breast cancer screening
13.2.2020All breast cancer risk factors evaluated with AI
6.2.2020A dog can smell diseases
2.12.2019ELIXIR Compute Platform for life and health sciences
18.11.2019New bioinformatics methods and measurement technologies call for continuously updated courses and analysis software
30.10.2019No need to turn up personally: SisuID improves electronic authentication
30.9.2019Risk assessment of cardiovascular diseases for all citizens
20.8.2019Federated user ID management: a single identity giving access to numerous bioinformatics services
4.9.2019Targeted treatment for venous diseases with vascular system modelling
4.7.2019Research on rare genetic disorders can be utilised in understanding the mechanisms behind even more common diseases
3.6.2019VEIL.AI: patient data in a veil
20.5.2019Biocenter Oulu: technology services for biomedical research
23.4.2019Mouse models provide insights into the causal mechanisms of diseases
4.3.2019Euro-BioImaging: imaging infrastructure
26.2.2019Imaging helps to highlight significance of data
14.1.2019Data harmony and standards: data must be processed, described and stored by uniform means
10.12.2018Hundreds of genes could lie behind a single disease
5.11.2018Help from the Finnish genome for the prevention of cardiovascular diseases
8.10.2018Disease prediction models are becoming more accurate thanks to the computational methods
11.9.2018Genetic data under control and in the desired format
23.8.2018Massive data management project: Finns’ heredity is collected and safeguarded
14.6.2018Half of all drug ingredients affect only three protein families
12.6.2018Looking for a good drug
29.5.2018Quick DNA analysis of patient samples with artificial intelligence
7.5.2018Secrets of the intestines
4.4.2018Algorithm determines the appropriate drug
19.3.2018Bank of million patient samples
20.2.2018Mapping the genomes of all organisms enables the development of new vaccines and medicines
7.2.2018Ordered and secured
2.11.2017Striving for a national service to utilise genomic data in health care
11.8.2017Better harvests on the horizon? Data will also be harvested
19.6.2017Microbes and climate change
21.5.2017Storing the whole genome of the Finnish population? The data will benefit disease research
6.4.2017”Smart life insurances” offered: human biological data is only useful when interpreted correctly
15.1.2016New drug molecules through determining the structure of proteins
26.10.2015BBMRI.fi: an IT Infrastructure for shared biobanks
24.9.2015Fighting cancer with mathematics
10.8.2015Saimaa ringed seal aids the study of population genomes
1.8.2015Webmicroscope stores tissue samples in the cloud
15.7.2015Pups and Pooches Behind Genetic Discoveries in Human Diseases Canine Genetic Research Benefits from ELIXIR Databases
5.6.2015Life sciences in European cloud
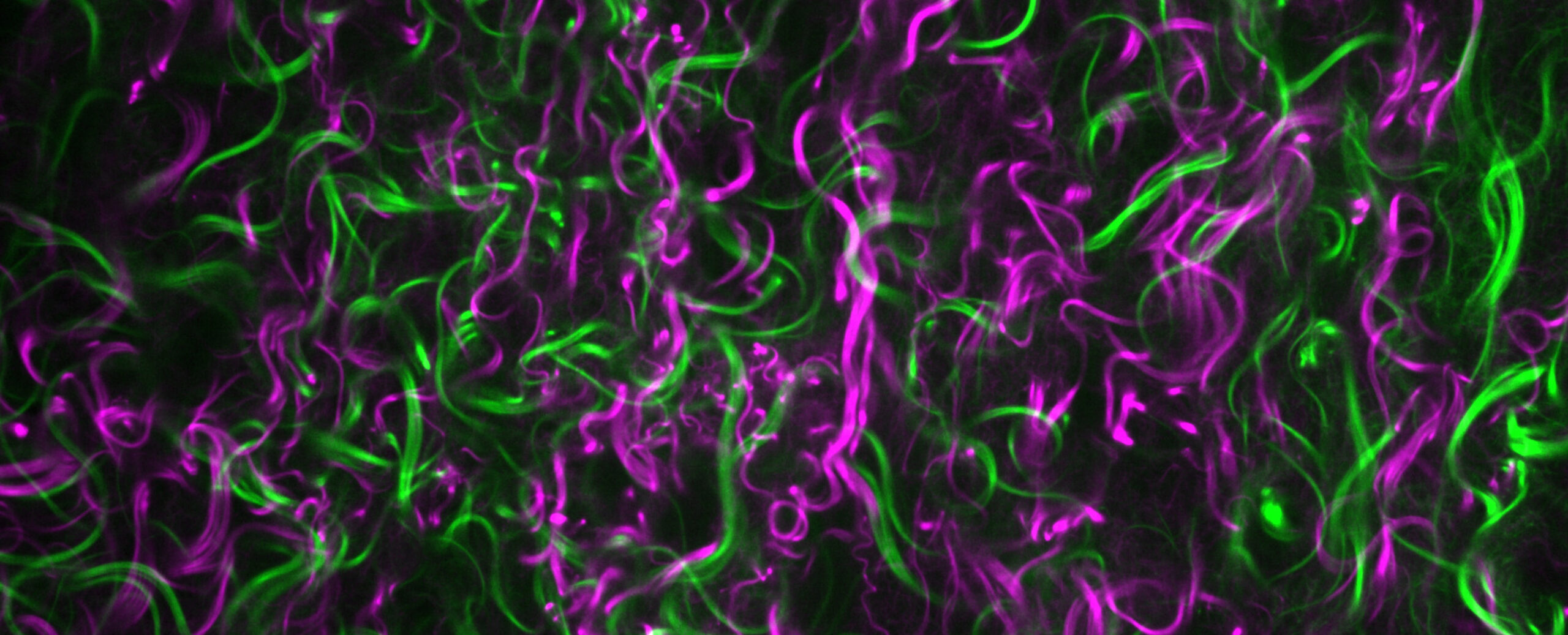
Curly collagen (green and magenta) found in the proximity of a breast tumour.
The movement and adhesion of cells is still poorly understood, even though the phenomenon is of great significance in, for example, the spread of cancers. To study it, Guillaume Jacquemet’s group has even had to develop their own software.
Many who lived their childhood in the 1980s may remember the animated series Il était une fois… la Vie (Once Upon a Time… Life). In it, the human body appeared as a bustling community, where industrious, oxygen-carrying red blood cells and tiny platelets wandered along blood vessels.
The heroes of the story, however, were the white blood cells, which alertly arrived on the scene whenever intruders such as bacteria, viruses or cancer cells entered the body.
But how do white blood cells actually find where they are needed?
In the animation, the matter is passed over as self-evident – after all, we know that our immune system can respond to threats. But that is where our certain knowledge ends, says Guillaume Jacquemet, associate professor of bioimaging at Åbo Akademi.
“We know that white blood cells respond to their environment, seek out where they are needed, and attach themselves in place with proteins. But how they actually do all of this is still in many ways open.”

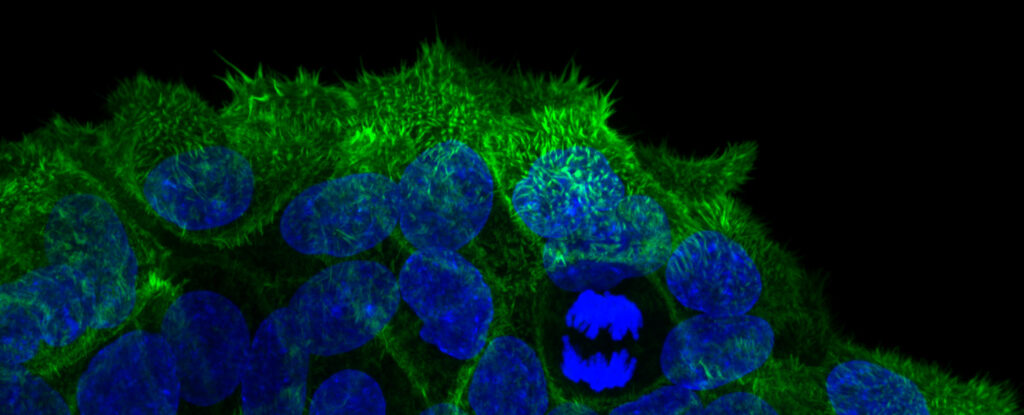
Jacquemet has studied cell movement for a long time, beginning with his doctoral thesis work at the University of Manchester. He arrived in Turku after his doctoral defence to do postdoc research and settled in. Jacquemet founded his own Cell Migration research group in 2019. Currently, the group has 13 researchers.
The majority of the cells in our body do not move anywhere. Exceptions include movement during embryonic development, as well as cells of the immune system – and one of its enemies, cancer cells. It is precisely the migration of cancer cells in the body that is the main research subject of Jacquemet’s group.
“Cancers develop in different ways,” Jacquemet says.
Some form a clear tumour, which can often be surgically removed if it is detected in time. More unpleasant are cancers that quickly begin to send cells elsewhere in the body to form metastases. Cancer that has spread in this way is often difficult to treat.
The particular focus of interest of Jacquemet’s group is pancreatic cancer. Pancreatic cancer is often difficult to detect, as its early-stage symptoms are vague. In addition, it often spreads aggressively. Therefore, the prognosis is often poor: fewer than ten per cent of patients survive.
Jacquemet and his group therefore strive to better understand the way in which cancer cells select where to attach.
An essential role in this is played by filopodia, which are hair-like protrusions projecting from cell walls. With the help of filopodia, cells observe their environment and search for suitable places.
“It is clear that the filopodia of cells use various proteins with which they attach, but we do not yet understand the whole picture.”
The observations are nevertheless useful. For a long time it was thought, for example, that pancreatic cancer cells would attach to the cell wall by the same mechanism as white blood cells. More detailed analysis, however, showed that the mechanisms are similar but different. Cancer uses slightly different proteins than white blood cells.
A small observation can be of great significance in, for example, the development of treatments.
“If one thinks of, say, a drug that would block the function of a protein used by white blood cells but would not after all affect cancer cells, such a drug would cause more harm than benefit.”
Drug treatments are, however, far from Jacquemet’s research. The Cell Migration group conducts above all basic research, in which the aim is to understand the biological functions of cells.
There is no shortage of basic questions. Not all cells, for example, move alone but in clusters. This kind of movement is understood even more poorly.
“An individual cell has to respond to its environment by itself, but moving in a group requires cells to take on different roles. We do not yet understand, however, how the cells in a cluster communicate with each other – or even what role the cells in the centre have in the whole.”
In a healthy human, cells move in groups mainly as part of embryonic development. Cancer cells, however, may also move in clusters.
“This we still understand very poorly.”

When starting out as a researcher, Jacquemet did a great deal of research with microscopes.
Modern microscopes take a large number of images. When following cell movement in microfluidic devices, the typical imaging frequency often resembles film, that is, approximately 24 images per second. This produces smooth movement.
Large numbers of images combined with the flow of cells make the observation of individual cells laborious. Information technology could be of help in screening, but unfortunately suitable free tools were poorly available.
When the coronavirus pandemic closed the laboratories, Jacquemet decided to use his time to build his own analysis tools. He taught himself to code and worked together with his colleagues to develop the ZeroCostDL4Mic tool, which helps identify interesting events in cell movement.
The tool, published as open source code, was a success, and it has since been used in thousands of studies around the world.
Nowadays Jacquemet’s group develops software routinely as part of their research. Three people from the research group code almost full-time.
“All the code we make originates from the needs of our research. This enables very natural development work: we think about what we need, and then we make it.”
The group makes all tools openly available, as well as their datasets, which they deposit in public databases such as Zenodo, PRIDE and BioImage Archive, both of which are ELIXIR Core Data Resources.
“In my view, offering tools for use cannot really be separated from the openness of data. Without the software, others cannot themselves verify how the data has been analysed and how the software affects the end result.”
Open source code is also an ideological solution.
“When software is created as part of publicly funded research, offering it for free is in my view the right solution.”
Besides, open code may find applications that one would not guess in advance.
“We have ourselves also used software developed by astronomers as a guide for our development work, software that was originally designed for tracking the movements of celestial bodies. The same logic works, however, for both planets and cells.”
Open databases help in this too. Openly available data collected by researchers around the world is used as test material when developing new tools.
In recent years, the significance of artificial intelligence has become prominent in software development. Learning artificial intelligence can analyse vast data masses from images and identify interesting events in them.
This makes the researcher’s work easier but on the other hand requires computing power.
“Although the majority of our software runs on ordinary computers, from time to time we also need supercomputers,” Jacquemet says.
In that regard, everything runs smoothly, however. Collaboration with CSC – IT Center for Science has always been flexible, and even the more unusual customer has been served well.
Where there would be room for improvement, however, is the funding of software work – especially in maintenance.
“In practice, all our software is created as by-products of research projects. This model works well during the project, but after the project ends, many software tools are at risk of being left unsupported,” Jacquemet notes.
“In order for software to remain usable, it must also be maintained. In my view, it should be possible to apply for funding for this more easily. Now the risk is that well-functioning and widely used software is left to fend for itself, when no one takes responsibility for continuity.”
Text and video: Juha Merimaa
Photos: Juha Merimaa and CellMig Gallery, cellmig.org/gallery/
20 May 2026
Read the article in PDF format.