

Pekka Ruusuvuori, Associate Professor of the Institute of Biomedicine at the University of Turku, leads the ComPatAI consortium, which is developing new ways to model histopathological tissue samples with generative and predictive AI. In medicine, histological samples are used to assess a patient’s need for treatment. The consortium’s goal is to use big data to create AI models that would produce more accurate diagnostic information in pathology.
In addition, they are developing virtual histological staining models based on generative AI. Besides Ruusuvuori, the consortium consists of research director and Adjunct Professor Leena Latonen of University of Eastern Finland and Teemu Tolonen, Adjunct Professor and chief physician at the department of pathology at the Fimlab laboratories.
The ComPatAI consortium focuses on analysing histological samples related to breast and prostate cancer. Using digitalised images allows the researchers to measure and automatically compute different cell types.
“Our work has focused mainly on prostate and breast cancer. There is ample data available on these types of cancers, as they are the most commonly encountered cancers in men and women. However, we want to create a very general-purpose model that could then be further refined for different and new applications.”
According to Ruusuvuori, the field of pathology is becoming increasingly digitalised. He says that in this sense, the Finnish pathological community is among the pioneers.
“In Tampere and Turku, we have moved completely to using digital pathology in diagnostics. Each time a sample is taken, it is scanned into a high-resolution digital image. There is a lot of routine diagnostics. As the population ages, we encounter more and more patients with cancer. This also means that there is loads of data coming in.”
The consortium receives scanned images of histological slides from Fimlab, the largest healthcare laboratory company in Finland. Fimlab’s clientele includes hospitals, health centres, occupational healthcare service providers and private medical stations. The Finnish Medicines Agency Fimea has currently granted the consortium a licence for using data from 160,050 cases, which translates to a total of approximately 600,000 slide images. Together, the images add up to about 0.8 petabytes of data, meaning that each file accounts for approximately 1.3 gigabytes. The massive amounts of data are currently being anonymised and transferred to the LUMI supercomputer in the CSC – IT Center for Science, an ELIXIR node in Finland. The project is one of the largest data transfers made to LUMI so far.
“It is incredible that we get to use these data for our research. We want to use this data to create AI solutions that function well in pathological work”, Ruusuvuori explains.
The researchers are currently set to have up to 2.5 million digitised whole-slide images at their disposal by the end of the project. This corresponds to a total of three petabytes of data.
“We have been granted permission to use technically all data produced in Fimlab’s routine digital pathology operations.”

Pekka Ruusuvuori has a strong background in signal processing, and he specialises in image analysis. Ruusuvuori is interested in how deep neural networks that are used in AI applications could be developed to be a better fit for diverse use cases.
According to him, machine can generally be taught to recognise the same things that humans would pick up on. It can learn to tell apart different tissue types or to distinguish cancerous tissues from healthy ones. It can be used to measure different factors in images or cells, such as how aggressive a cancer is and how far it has progressed. Artificial intelligence can identify cancerous areas in tissue samples before examination by a pathologist. It may also suggest a score based on the data it has assessed. For example, prostate cancer tumours are given a Gleason score, which indicates how aggressive or advanced the disease is.
“It’s entirely possible to train AI to perform many tasks that human pathologists usually take care of.”
“Previously machine learning models have been built with a certain variable and teaching material that shows a certain object appearing in a specific part of the image and which score this finding corresponds to. It would take countless hours of work for us to mark this information on all the hundreds of thousands of images we are using.”
This annotation data has previously played a key role in teaching artificial intelligence to automatically detect abnormalities such as cancer cells in the samples. Ruusuvuori says that algorithms have been improved and are consequently able to use unannotated raw data.
“I think the most interesting thing about what we are doing with these algorithms is what else we can extract from these images. In other words, to look at the features that machines can detect but humans cannot. These slide images include all visual data that we have. If there is a statistical link to be found there, the machine learning algorithm will find it. However, these links may be extremely complex. Modern neural networks can accurately detect complex links between spatial data and the predicted variable. These things can be very difficult for us humans to grasp.”
Together with his research group, Ruusuvuori has been able to successfully predict gene expression and mutations directly from histological images. Gene expression refers to the process of a cell producing the molecules that is encoded in its DNA. The gene expression varies across different tissues. Based on the images, AI can detect miniscule changes that are invisible to the human eye.
“Based on the images, the machine can identify effects of gene expression in cells and tissues. It can detect even the slightest phenotype variations, including those that we as humans are not trained to see. I want to highlight that so far, we only have indicative results, and that the method will not work for all tissue types or genes. Some gene expressions do not lead to tissue-level changes that could be predicted from a whole slide image.”
ComPatAI consortium is currently developing a so-called foundation model for utilising large datasets. As the name suggests, this model would create a general-purpose foundation for developing further AI solutions. The model is trained in histology based on a large set of samples, without target variables or annotation data.
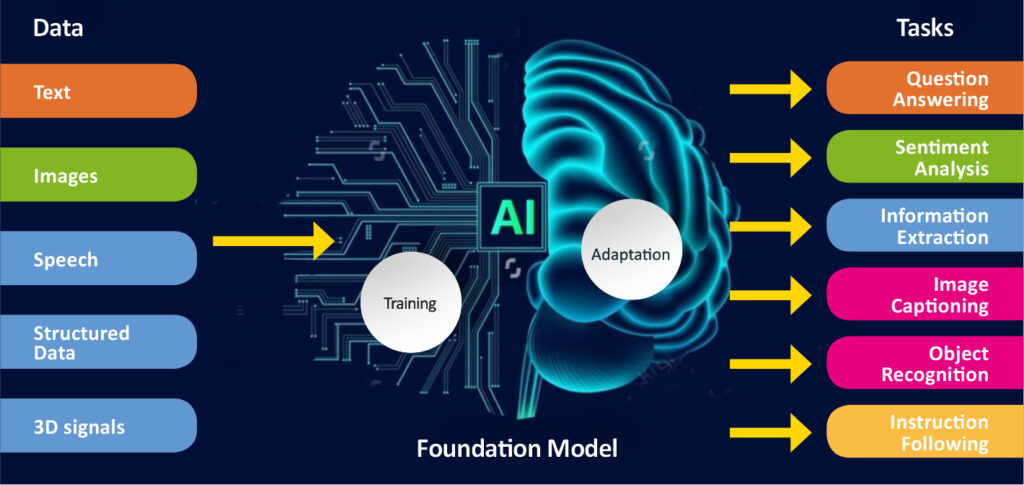
“When you start teaching a model like this to recognise diseases such as breast or prostate cancer, it starts to learn to perform the task it has been given. This will allow us to reach more accurate solutions much faster than before. It allows us to use large, unannotated datasets. This is a big step forward for us.”
ComPatAI consortium is currently building its own foundation model based on a Finnish dataset.
“This is basic research that will allow us to be among the first to further refine these models in Finland. I do not want us to rely solely on big foreign firms or research groups, and instead wish that we would be able to be build a model based on Finnish data. We have high-quality, population-level cohort data that need to put to good use. I hope that this will lead to the establishment of companies in Finland that will develop solutions to benefit patients in routine diagnostics.”
One of the key questions is how quickly data can be transferred and used. Computing and data storage capacity are constantly in high demand. This is where the services provided by CSC, the Finnish ELIXIR node, come in.
“We’re extremely pleased with the support CSC has given us, since this is an exceptionally large project that uses very large datasets. We are in a privileged position because we have the support of an organisation like CSC. This is a clear competitive advantage for us, and we really appreciate it.”

Digital pathological data and other potentially sensitive health data types, such as registry and omics datasets, are going to become more readily available through CSC’s data secure user environment.
“We’re just getting started with the development work”, says Tommi Nyrönen, who leads the ELIXIR Finland Node.
“ELIXIR’s node in Finland has helped transform the biomedical resources required by CompPatAI to a platform service operated by CSC. The CSC Sensitive Data platform emerged from this need. However, it continues to serve various other researcher projects in the field. One of these is the EU’s digital pathology archive initiative bigpicture.eu, which is set to launch in 2026. It is a sustainable solution for managing digital pathology datasets and bringing them to high-performance computing services across Europe.”
Ari Turunen
26.12.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). The ComPatAI consortium uses large datasets to create an AI learning model for pathology. https://doi.org/10.5281/zenodo.14823370
More information:
FIRI
Article was supported by the Research council of Finland under grant number 345591 for ELIXIR European Life-Sciences Infrastructure for Biological Information (FIRI 2021)
Ruusuvuorilab
Fimlab
University of Turku
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Glaucoma is a progressive disease of the optic nerve that causes damage to the optic nerve head and nerve fibre layer. The risk of developing glaucoma increases with age. Some 2% of people over the age of 50 have glaucoma, and more than 5% of those over 75. There are estimated to be more than 60 million patients with glaucoma globally, of which around 6 million are categorised as being visually handicapped.
The challenging thing about glaucoma is that in its early stages it exhibits no or very few symptoms. Early diagnosis is very important because any damage that has already occurred cannot be reversed. The objective with treatment is to prevent any visual handicap caused by glaucoma. With most patients, the condition advances gradually over many years. However, with a small percentage of patients, the disease may lead to damage in a shorter period.
For purposes of glaucoma detection and identification of progressing speed, it would be best if healthcare systems found the high-risk cases as early as possible. Artificial intelligence models are currently being developed for early detection of glaucoma.
Researcher and project manager Ara Taalas specialises in data science, artificial intelligence and machine learning algorithms in medicine. One of his research objectives, in a joint project involving the Institute for Molecular Medicine Finland (FIMM) and Terveystalo health clinic, is to develop effective learning algorithms for glaucoma detection. Previously, Taalas modelled stem cell differentiation processes and worked in drug design.
According to Matti Seppänen, chief physician and Head of Ophthalmology at Terveystalo health clinic, glaucoma diagnosis and classification are based on the examination of the optic nerve head, nerve fibre layer and anterior chamber angle, intraocular pressure measurement, and a visual field test.
“The pathogenesis of glaucoma is not known, but damage to nerve cell structures probably contributes to glaucoma damage.”
Probably some 30–50 per cent of patients have intraocular pressure which is considered normal (10–21 mmHg). Patients have an individual susceptibility to the development of glaucoma damage at different intraocular pressure levels. Some patients develop glaucoma damage at lower pressure levels, while other patients may have minor damage even at higher pressure levels.
“At the moment, a glaucoma diagnosis requires examination by an ophthalmologist and several additional examinations. The optic nerve head can be examined by means of, for example, biomicroscopy and stereo papilla photography. The nerve fibre layer can be examined with colour fundus photography or optical coherence tomography (OCT) of the nerve fibre layer.”
During an examination, glaucoma may be suspected on the basis of the shape of the optic nerve head, for example. The structure of the optic nerve head can be evaluated with a measurement of the cup/disc ratio, meaning that the size of the optic nerve cup is compared to the size of the outer edge of the optic nerve head.
“Damage to the nerve fibre layer may show in an OCT examination as a thinned nerve fibre layer. In colour fundus photography, defects in the nerve fibre layer may also be discovered. A glaucoma diagnosis is often based on several examinations, and currently there is no single method for screening the entire population for glaucoma. Artificial intelligence applications may in the future bring considerable help for screening and diagnostics.”
Esa Pitkänen from the Institute for Molecular Medicine Finland FIMM (University of Helsinki) tells how glaucoma can be studied with the help of algorithms.
When developing the artificial intelligence model, Ara Taalas focused on how the nerve layers of the fundus appear in the photographs. The algorithm will help to detect changes in the fundus pictures that can indicate damage to the nerve fibre layer. The purpose of the model is to find out whether subtle changes in the network in the fundus, as they become darker and more monotonous, can be linked to damage in the nerve fibre layer.
“This is one of the factors the model is designed to focus on. In the future, the model will be taught more nerve fibre patterns in the fundus. The purpose of these algorithms is to find ways that will help doctors to make decisions. And advanced artificial intelligence system may detect changes that not even the most experienced clinician can see.”

Examinations on the eye’s structure and operation involves variations caused by the examination method used, experience of the person assessing the case, and the patient and how serious the disease is. Evaluating the optic nerve head does not always result in sufficient accuracy using the current methods. The result of the visual field examination may be normal even if the optic nerve and nerve fibre layers have been damaged. This is because structural damage usually occurs before any visual field defects. If we are able to develop applications that examine more accurately and more efficiently any structural changes, glaucoma diagnoses may be made earlier.
According to Taalas, one application for the model could be that the artificial intelligence model is always used when performing an eye examination.

“Population surveys have found that up to half of those who have glaucoma have not actually been diagnosed with it. The existing screening methods are not cost-effective enough and a general screening of the population cannot be done for lack of sufficiently good methods. If artificial intelligence applications were able to identify with sufficient accuracy patients that have a higher-than-average susceptibility to develop glaucoma, it would be easier to screen out those among the population that do not yet have the symptoms and offer them early treatment for best results.”
One of the future versions is that during a visit to the opticians or healthcare worker, the examination would include fundus photography, and at the same time artificial intelligence would analyse the patient’s fundus photo. If artificial intelligence indicated that the patient had a higher risk than average to develop glaucoma, the patient would be referred to further examinations at an early stage.
With artificial intelligence applications, the division of work would probably change dramatically in the optical field and the diagnosis of eye diseases. This would also result in significantly higher numbers of patients being treated. As the age structure of the population is changing, the number of glaucoma patients in Finland will double from the current figures by 2030.
Taalas uses the computing services of Finland’s ELIXIR Center CSC. He develops models together with researchers in FIMM’s Machine Learning in Biomedicine team, and the same source code can be used on the computing servers of both CSC and Terveystalo.
“Finland is at a high level in terms of data management, but individual healthcare actors typically do not have a comprehensive picture of their patients – patient data is often scattered between various service providers. When customers go to a different organisation, the data does not follow them, which may make diagnosis and treatment more difficult. From the viewpoint of a researcher, the ideal thing would be to have a site for the entire country where each patient’s medical history could be found in its entirety.”
Data description should also be standardised.
“The structure of patient data systems has a major effect on the usability of any data entered into it. Fields where data can be entered in free form may be convenient for the person typing it in, but cause a lot of trouble to data analysts when trying to utilise it. Analysts often have to do a lot of work to standardise the data and to identify entries that contain errors. Modern patient data systems have in this respect become better in that they are much more structured.”
Ari Turunen
23.11.2021
Read article in PDF
Citation
Ari Turunen, Lila Kallio, Arho Virkki, & Tommi Nyrönen. (2021). Patient data creating better artificial intelligence models. https://doi.org/10.5281/zenodo.8135413
More information:
Institute for Molecular Medicine Finland FIMM, University of Helsinki
Terveystalo
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Medical research will not progress without data and its re-use. Collected data can be used to create artificial intelligence models to make speedier diagnoses and to support care decisions. New data analysis technologies emerge all the time, but how to make data available for all researchers?
One of the strengths of the Genome Center, which is being established in Finland, consists of biobank databases. The Center would be in charge of developing a genome data register, that is, a centralised system for storing and managing genetic data. The aim is to create a high-quality database that describes genetic variation in Finland. Auria Biobank’s Director Lila Kallio believes that good cooperation between the biobanks and the Genome Center can lead to significant results in the screening of gene variants.
“Once the Genome Center has been established and starts operating, genome data created during research can also be stored in the Genome Center. The Genome Center could re-analyse any genome data against all accumulating reference genome data. This would enable, for example, the screening of new identified, clinically significant variants by using previously produced and stored data,” says Lila Kallio.
The Biobank Act was enacted in Finland in 2013, enabling the establishment of biobanks. There are currently 11 biobanks in Finland. In 2020, the biobank network was joined by Arctic Biobank, which stores extensive population data collected by the University of Oulu in the north of Finland. Researchers in Finland can utilise material from all biobanks through the Fingenious online service. Fingenious is a digital tool through which a researcher can send a request for material to be made available to them. This service is provided by the Finnish Biobank Cooperative, FINBB.
“Biobanks store data about samples securely. Data about biobank samples is available to all researchers. A researcher must present a research plan for approval by the biobank steering groups or ethical committee. Biobanks have a process in place for the research use of samples and their related data.”
Finland has exceptionally comprehensive and high-quality health care data resources. The Act on the Secondary Use of Health and Social Data (552/2019) came into effect in 2019 in Finland. Secondary use of data means that customer and register data within social care and healthcare are used for some other than the original purpose. This act on secondary use has also created pressure to amend the Biobank Act from 2013. The significance of data in biomedical research is increasing and the legislation should create the conditions for both research and appropriate data security.
Secondary use obviously requires that data collected of people is managed securely. Identifier data of human samples stored in the biobanks is carefully protected.
“The biobanks remove all personal identifiers and replace them with pseudonym codes. When samples are handed over for research purposes, the pseudonyms are replaced with another code, specific to that particular research. The code key is stored in the biobank. If you need to access the original sample owing to, for example, by some clinically significant detail, this can only be done with the code key,” says Kallio.
The use of a code key enables the data to be re-used in subsequent more research purposes.
“If the sample was anonymised, that is, making impossible to identify, it would be impossible to access it after any findings in biobank research, and no sample-specific data could be added to it later.”
According to Lila Kallio, the real value of samples is in the data created from it.
“Data is created during diagnostics and treatment. Research also results in analysed data, which must be returned to the biobank in possession of the sample, to be appended to it. Biobanks manage not only identifier data but also clinical data and data that has been produced during research.”
The act on the secondary use of health and social data concentrated the permit process management to Findata, a new legal authority. Lengthy processing times of permit applications has become a problem. All applicants are treated equally regardless of the size of material they are making an application for.
Auria’s chief data officer (CDO) and adjunct professor of medical mathematics Arho Virkki points out that material can be used in a number of ways, and that’s why there should be different protection levels based on the purpose of use. According to Virkki, the initially planned data security improvement leap for the secondary use of health data is too great to be taken in one go.
“Extreme protection weakens data availability, which means that data security won’t reach an optimal level. To me, optimal data security means that material is truly available for scientific research, planning of new treatments and controlling treatment processes. In the optimal situation, data is available and at the same time it is adequately protected. The protection level should be set on the basis of the risks involved.”
Because data management is part of the work of doctors and nurses, Virkki says a balance should be found between material availability and its protection. At the moment, the system is lopsided.
“For example, exploring clinical data is part of medical students’ curriculum. One part of their training is the use operational systems to find data in order to learn.”
According to Virkki, the isolated data architecture has been the culprit for a long time. Owing to a defensive approach and regulation in medicine and healthcare, information architecture is more traditional compared to, for example, logistics and the financial sector. This is why various information systems are poorly integrated.
Virkki does admit, however, that hospitals are more complex places than logistics centres, for example. In logistics systems, postal packages follow a pre-defined route and their travel is easily recorded by the system, whereas when a patient arrives at a hospital, the following steps are typically more complex and involve a great number of different options.
The act on secondary use of data, however, makes the assumption that one type of secure data processing environment fits all uses. Virkki says that the legal entity issuing the research permit could provide a range of user environments based on the researchers’ needs.
“There could be a basic environment sufficient for simple data analysis that consists of spreadsheet software and the usual range of statistical programming languages.”
Then again, if researchers need a custom environment, they should be given exact details about data security with assurances that they will comply with them.
“This way, the authorities would set the requirements for data security, but the researchers should be accountable of it, as has been the case up till now. At the end of the day, it is the researchers’ responsibility to ensure that their results are correct, honest, scientific and anonymous.”
According to Virkki, people in the medical field in Finland take great pride in their work and have always strived to process medical material in a proper fashion. Virkki says that data security can be ensured through licensing and training. Data security issues should be part of medical training. Virkki is a regular speaker at the University of Turku on an introductory course on the basics of clinical research about data platforms and data security.
According to Virkki, amendments are in progress for the act on secondary use of data. If the provisions can be made more flexible and the permit processes faster, there will be many opportunities for artificial intelligence research.
“Now that the reform on social care and healthcare was passed in Finland, there is a good possibility to combine the patient data of basic and specialised health care, that is, view patient data as a single entity. This in turn will enable the development of new AI applications for the clinical side.”
The algorithms of AI models can perform text-based analyses, make medical records or learn to identify details in images that can be used in diagnoses.
“In fact, artificial intelligence is just modern statistics, a refined branch of mathematical statistics. AI models make use of complex statistical methods. When you talk of machine learning, you actually mean statistical learning. Today you can calculate predictions in such a precision that is almost feels like magic.”
Virkki has been intrigued by AI models for a long time. In his doctoral thesis, he created an AI model for human respiratory system during sleep. Recently he has been developing a prediction model for pulmonary embolism. The model is used as a tools for decision-making. Pulmonary embolism occurs when a blood clot gets wedged into an artery in the lungs. The most common symptom is sudden shortness of breath. In serious cases of pulmonary embolism, the clot is diluted by injecting an anticoagulant into a vein.
“If there is reason to suspect that a patient in an emergency room has pulmonary embolism, you have to act fast. A machine can quickly go through a set of scanned images and inform the radiologist where they should focus on in any image. After that, the decision is made whether to start diluting or not. If not, another treatment is chosen. You should be able to do all the following in less than 10 minutes: lung imaging, diagnosis and starting the treatment.”
According to Virkki, the pulmonary embolism model was the first scientific test trying to solve a difficult problem with very little data. However, a more comprehensive and more accurate AI model is under development. Scientific publications and dissertations will be published on the subject.
“If realised, the model will speed up decision-making in case situations, but also assist in quality control. For example, we can screen afterwards whether we detected any smaller cases of pulmonary embolism.”
The development of artificial intelligence models requires a lot of data with which algorithms are taught, and plenty of computing power.
The hospital district of Finland Proper uses the ePouta cloud service of Finland’s ELIXIR centre’s CSC, with a dedicated 10 GiB connection. Virkki hopes researchers could have better access to the ELIXIR network.
“It would be great if researchers were given capacity from the ELIXIR infrastructure for their work. The data resource would be made available directly in the ELIXIR environment, and ELIXIR would ensure there was enough computing capacity.”
The ELIXIR Node in Finland (ELIXIR-FI) is hosted at the CSC – IT Center for Science Ltd. CSC operates resources and services that are part of ELIXIR, like pan-European ELIXIR identity and access infrastructure. In ELIXIR the data needs to be managed as a federation, where data providers work as a single infrastructure providing mechanisms where researchers can bring their analysis to where the data is located. The ELIXIR Compute Platform infrastructure will allow life scientists to easily access, share and analyse data from different sources across Europe. The objective is to combine all components of the ELIXIR Compute services into a seamless workflow. A researcher may use the ELIXIR Authorisation and Authentication services to securely create a scientific software analysis environment, and use the environment to access large biological data resources stored in a cloud.
Text written or dictated by a doctor can be utilised in artificial intelligent models used in aid of current care guidelines and diagnoses. Statements and sentences can be constructed into data and teach the algorithm to make deductions. In the project participated in by Auria Biobank, Turku University Hospital and the University of Turku, artificial intelligence was taught to extract data about smoking from about 30,000 patient records. In the project, headed by researcher Antti Karlsson, a language model called ULMFiT was used. The model was trained using the analysis computers of Finland Proper Hospital District, making use of texts from Wikipedia. After this, the model was trained to become a classifier by means of some 5,000 manually annotated sentences. These days there are also more sophisticated, pre-trained models in Finnish, most notably perhaps FinBERT, based on the Google BERT model. This was created by a University of Turku research group led by Filip Ginter, using computing power from Finland’s ELIXIR centre CSC.
Based on the data collected by the artificial intelligence model, the study showed that quitting smoking, even once a cancer diagnosis has been made, may extend the patient’s life expectancy considerably.
“I’m sure that future patient record systems will not be formal, with items picked from drop-down menus, but written more in prose, with systems that automatically structure them,” says Karlsson.
“This also improves work efficiency. I don’t even want to imagine how difficult it must be for a busy doctor to enter complex matters into the systems.”
When you mine a large data mass, you save an awful lot of time and money. The artificial intelligence model trained by Antti Karlsson analysed patient records, searching for smoking-related issues. In the study, the model analysed text date obtained from the records of 30,000 patients. According to Karlsson, models like this can produce analyses that are more than 90% accurate, in hours or even minutes. It’s quite different from manually reading through the texts of 30,000 patients, entering variables into a table.
“In the best-case scenario, such models could be readily available in a data lake, structuring this tobacco data automatically for research purposes,” says Karlsson.
The model does not produce treatment instructions for an individual patient, but creates a good overall picture.
“A believe that at least initially, the automated systems of the future will collect data relevant to reporting and research, while the really important things, such as dosages and allergies must still be checked and filled in manually by experts.”
Ari Turunen
26.10.2021
Read article in PDF
Citation
Ari Turunen, Pasi Kankaanpää, Sirpa Soini, & Tommi Nyrönen. (2021). Sensitive data infrastructure. https://doi.org/10.5281/zenodo.8135532
More information:
Karlsson et al. (2021): Impact of deep learning-determined smoking status on mortality of cancer patients: never too late to quit. Esmo Open Cancer Horizons. Vol 3. Issue 3.
https://www.esmoopen.com/article/S2059-7029(21)00135-6/fulltext
Auria Biobank
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org

Deep learning has revolutionised cancer research. Deep neural networks can automatically detect features within a patient’s sample data that can be used to identify cancers. In the future, learning algorithms will be able to identify potential early-stage cancers from a blood sample. Esa Pitkänen and his research group at the Institute for Molecular Medicine Finland (FIMM) are developing a new generation of deep-learning algorithms.
Algorithms have been used to identify cells in sectional images of tissue samples. For instance, if tissue cells appear atypical, the algorithm will spot this and determine if the cells are cancerous. DNA sequence data from tumours is now being used along with imaging data to identify cancers.
“Until recently, it was difficult to tell from a DNA sequence what kind of tumour an identified sequence came from. Now new technologies and deep learning algorithms have been created,” Pitkänen says.
Pitkänen and his team are developing algorithms that identify short, repetitive snippets of DNA sequences. These algorithms can be used to find DNA sequences that mutate frequently in a particular type of cancer or to which certain proteins involved in gene regulation bind. Analysis of these sequences can be used for various purposes, such as charting the causes of cancer and developing medicines.
“The replication of DNA in cell division is not perfect; mutations can occur during the process. The division of a single cell involves the copying of about six billion nucleotide pairs in DNA, so errors will inevitably occur. Even the slightest probability of errors is enough to guarantee mutations,” says Pitkänen.
“If enough mutations occur in genes that prevent tumour growth, for example, cancer may start to develop.”
An example of this is a point mutation in which one base within the DNA strand is replaced with another. The enzymes involved in copying DNA may make a mistake when a cell is dividing, for instance by incorrectly repairing the part of DNA that was damaged by ultraviolet radiation from sunlight. A typical mutation caused by ultraviolet radiation that can result in skin cancer is that two consecutive cytosine (C) base molecules in the base pairs of human DNA are converted to two thymine (T) base molecules. When skin cancer-specific mutations of this type are detected in sufficient numbers, the algorithms can learn to associate them with a particular type of cancer.
“We try to predict the type of cancer and tumour from the mutations. This also provides information on how treatment can be developed.”
Pitkänen and his group analyse blocks of sequences and train algorithms to pinpoint deviations in them. From these abnormalities, the algorithm can detect tumours and classify them into different cancer types.
“Before joining the Institute for Molecular Medicine Finland, I worked at the European Molecular Biology Laboratory in Heidelberg, where I participated in the Pan-Cancer Analysis of Whole Genomes (PCAWG) project. This project involved the analysis of more than 2,600 entire cancer genomes. My group is using data from the PCAWG project in several of our cancer genomics projects.”
An algorithm developed by the group has been taught the mutations found in the tumour samples of 2,600 cancer patients. This sample set contained about 47 million mutations. Approximately 50 million somatic mutations were found in the sequence data.
“We trained the algorithm to try to deduce the type of cancer from these sequence changes. Once the algorithm is given all the mutations of different tumours and their sequences, in the future it will be able to determine the kind of tumour that has been detected. This deduction process is based on the algorithm learning these connections.”
Through deviations in the sequence data in tumours, the algorithm learns to identify when a given tumour corresponds to a particular type of cancer. It can group tumours based on sequence data alone.
“A researcher in my group, Prima Sanjaya, has developed neural network models for analysing sequence data. Every now and then, researchers come across cases in which a cancer has metastasised without being able to tell where it has spread from. Such cases could in the future be dealt with by means of a liquid biopsy – that is, it will hopefully be possible to determine from a blood sample if the patient has cancer, and if so, what kind.”

Liquid biopsies are based on the fact that cells release into the bloodstream and other body fluids a type of DNA called cell-free DNA (cfDNA). Cancer cells also release DNA, which makes it possible to test for cancer mutations in the blood plasma.
“If a liquid biopsy shows traces of cancer, we don’t know exactly what kind of cancer it is, as it could have entered the bloodstream from anywhere in the body. If we have the means to examine these cases more closely, for example with deep learning algorithms, we could obtain valuable information on where in the patient’s body the tests should be focused. The algorithm may indicate that the source of the cancer DNA could be the large intestine, for example. I believe that such algorithms will become extremely important. Liquid biopsy and algorithms can make it possible to diagnose cancers without surgery.”
In addition to hereditary factors, the development of cancer also depends on the person’s lifestyle. Plentiful research has been conducted at the University of Helsinki on various cancers, such as intestinal cancers.
“It is known that eating red meat contributes to the incidence of colorectal cancer. The mechanisms by which the disease is caused require further research, but in recent years a lot of progress has been made in understanding the significance of DNA alkylation reactions, which are caused by red meat, for example.”
Colorectal cancer is one of the most dangerous cancers in Western countries. In countries such as Finland, it leads to death in 30 per cent of cases. About 15 per cent of colorectal cancers belong to the cancer group that exhibits microsatellite instability (MSI). Microsatellites are sequences of DNA that can vary in length from person to person, and thus function as individual identifiers in much the same way as fingerprints. Microsatellite instability occurs when the post-replication repair mechanism of cellular DNA does not function properly. This causes mutations to begin to accumulate, especially in microsatellites.
“In an MSI tumour, microsatellites are easily vulnerable to single-base additions or deletions. For example, out of eight consecutive adenine microsatellites, one adenine may be lost. When it occurs in a gene, such a change causes a complete transformation in the content of the amino acid chain of the protein that is encoded by the gene. If there are enough changes in genes that are important for preventing uncontrolled cell growth, cancer may begin to develop.”
MSI is often associated with other cancers as well as colorectal cancer, such as stomach, uterine, ovarian and brain cancer. MSI analysis can be used in the prognosis of cancer. The treatment choice may be influenced by the analysis.
“An interesting thing is that the deep neural network is also learning to classify different subtypes of cancers. For instance, it identified the MSI subtype of colorectal cancers,” says Pitkänen.
The ELIXIR Node in Finland, hosted by CSC – IT Center for Science, is one of the main partners in the Personalised Medicine in Europe (PerMedCoE) project. For example, the three-year the HPC/Exascale Centre of Excellence for Personalised Medicine in Europe project is aimed at making effective use of cancer-related data in healthcare and speeding up the process of diagnosis
“Individualised treatments of the future, among them cancer treatments, will be based on a precise understanding of the patient and their illness. This will result from gathering a large volume of data of different types, such as tumour-related data and imaging data during cancer treatment. Many data collection methods produce a mass of data, and the new computational methods developed for analysing it require a very large amount of computational resources,” says Pitkänen.
“Developing a new computational method from the idea stage into a functional healthcare technology is a huge challenge in an operating environment like this. With cancer treatments in particular, it is important that information relevant to patient care be made available to the doctors as rapidly as possible. I’m confident that the results of the PerMedCoE project will provide a basis for deriving relevant information from a colossal volume of health data to help doctors in their work, and thus significantly improve treatment outcomes.”
Ari Turunen
16.9.2021
Read article in PDF
Citation
Ari Turunen, Esa Pitkänen, & Tommi Nyrönen. (2023). Teaching an algorithm to identify cancer from sequence data. https://doi.org/10.5281/zenodo.8135303
1.External factors (e.g. UV radiation from sunlight), 2.Internal factors (e.g. a spontaneous deamination reaction, in which the amine group of a base changes, for example from adenine to uracil) 3. DNA copying errors.
A mutation is a change in the nucleotide sequence of DNA or RNA. A nucleotide consists of a base, a sugar molecule and a phosphate group. The sugar in DNA is deoxyribose and the sugar in RNA is ribose. The four nitrogenous bases that DNA contains are guanine (G), adenine (A), cytosine (C) and thymine (T). RNA has three of the same bases as DNA, but instead of thymine its fourth base is uracil (U).
A mutation may be a change of a single nucleotide – that is, a point mutation – or the change may involve multiple nucleotides. In a point mutation, one base is replaced by another in the RNA or DNA strand. Large mutations can involve thousands of nucleotides, and are called structural changes. A structural change can affect multiple genes at the same time. Cancers are usually caused by several somatic mutations. Mutations of this kind are not inherited, and can occur at any time of life from embryonic development onwards. Mutations may bring about a change in the functioning of a normal cell, causing it to begin to divide uncontrollably.
At the middle of the picture are presented different types of mutations, distribution of mutations on chromosomes and epigenetic information. Epigenetic inheritance is influenced by many external factors, such as nutrition. An example is the development of identical twins so as to become distinct from each other in appearance.
Modelling mutations:
Linear models
Deep neural networks
Transformer models. Transformers are a family of deep learning models that work particularly well with certain types of data, such as textual data. This makes them well suited to machine translation, for instance. In cancer research, transformer models can draw attention to mutation types that are important for identifying a particular type of cancer. For example, in skin cancers that contain many sunlight-induced mutations (C> T, CC> TT), the transformer will focus on these particular mutations.
Ari Turunen
For more information:
HPC/Exascale Centre of Excellence in Personalised Medicine (PerMedCoE)
Institute for Molecular Medicine Finland (FIMM)
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Turku University Hospital and Auria Biobank aim to have all tissue specimens in digital format. The samples would be scanned from glass slides, with diagnostics in pathology performed on computers. They will also develop artificial intelligence models, or classifiers, to identify e.g. cancer from digitalised samples.
Turku University Hospital alone takes 200,000 patient samples every year. Tissue samples are placed in formalin and cast into a paraffin block that can be sliced and subsequently examined under a microscope. The paraffin blocks are stored at the end of the process. Managing the samples is laborious and time-consuming. Systematic digitalisation of the samples makes the job easier.
“Since the samples are so numerous, metadata will enable us to quickly find the samples that we want,” says Antti Karlsson, data analyst at Auria Biobank.
You can, for example, search the database for all samples indicating breast cancer tumours. By using metadata, searches can be narrowed down to pinpoint, say, samples with a certain receptor status from 60-year-old breast cancer patients.
In the digital pathology project, samples on microscope slides are scanned. Then a pathologist can view the samples on a computer screen and describe and classify them. All this annotation data is relevant to teaching artificial intelligence to automatically detect abnormalities such as cancer cells in the samples. This would considerably speed up the work of pathologists. Auria Biobank has invested in data analytics, development of algorithms, and machine learning models.
Turku University Hospital has huge numbers of tissue specimens stored on microscope slides. The problem is that no metadata can be stored on the slides and transferred into databases automatically. The idea now is that pathologists add metadata to new samples by means of graphics software.
Karlsson says that the work is mechanical to begin with. Pathologists use graphics software to indicate the points where e.g. cancer is found on the scanned samples.
Description data is also required. This is where neural-network language models might come in. A pathologist would add information about the sample directly into a computer. This has been studied in cooperation with Filip Ginter’s research team at the Department of Future Technologies of the University of Turku. The research team has focused on how computer software can be used to analyse natural text and speech. From a large amount of unclassified text, the language model learns how a spoken language seems to work statistically. Auria Biobank and Turku University Hospital are interested in how medical statement texts could be formed into classified and structural information by means of language models.
“One application of digital pathology could be to mine various types of information from statements, such as which part of the sample contains interesting tissue, making sample selection for research purposes easier. We could also develop a model that automatically structures regular medical statement texts. Pathologists could speak in ‘prose’, which artificial intelligence would collect and compile into a structured table.”
According to Karlsson, such tables are already used relatively frequently, for example when pathologists have agreed on what aspects of each tumour should be reported.
“At the moment we are experimenting with these models, for example to detect and classify smoking data from among hundreds of thousands of statements, and to detect cancer metastases, symptoms related to hospital infections and various diagnoses.”
The challenge is that data comes in a variety of forms. For example, scanners by different equipment manufacturers produce different kinds of data that should be presented in a systematic way.

Metadata and digitalised sample material are used to develop artificial intelligence applications, for example, which are taught to automatically classify the locations with cancer cells in images. To teach the artificial intelligence system, we require some material classified by pathologists. According to Antti Karlsson, you do not in fact need very many images for the algorithm to start learning.
“A few dozen images is enough to get started. A single, whole slice image may yield a thousand small images that can be used to train the models.”
This means that up to 10,000 small images can be obtained from 20 patients.
“A large image cannot be used with an algorithm, because no computer has the kind of graphics processor memory required to deal with it.”
Karlsson stresses that artificial intelligence models which examine images are different to models examining texts.
“Clearly, they are all manifestations of artificial intelligence, and even of neural networks, but their structures and operating principles are quite different. Artificial intelligence is actually more like a collection of tools, each one of which is useful for a specific application.”

Auria Biobank’s Director Lila Kallio says that, in addition to genome data being used for research purposes, digital pathology making use of data analytics is a focus area at Auria.
“There is growing interest in how digitalised cancer tissue samples can be used to identify various issues. We are involved in studies where we try to use an algorithm to examine an image of a primary cancer tumour and predict how it will respond to treatment, or whether the primary cancer tumour will metastasise. There are indications that the algorithm may be able to predict something that is not otherwise visible from a histological image.”
According to Lila Kallio, Finland has been a pioneering country in data management and sharing. The Finnish Biobank Act has enabled research and the combination of data from various registers. It is critically important that clinical information can be connected to samples.
“Services for researchers have been provided on a one-stop basis. Biobanks take care of the permits, collect the samples and combine all clinical data related to the research. All this can then be combined with other data, such as genetic data.”
Researchers get all the samples through a biobank.
“Biobanks cooperate in Finland. Researchers can request samples from all Finnish biobanks with a single request made to the Finnish Biobank Cooperative.
According to Lila Kallio, the challenge now and in the future is data storage and management.
“Data is stored inside firewalls in hospital districts. If diagnostic samples will be digitalised on a larger scale within pathology, the storage capacity problem must be solved as well. In addition, the image sizes are so huge that they cannot be transferred on ordinary data networks.
The Finnish ELIXIR Center CSC plays an important role in terms of computing power, and safe storage and usage environments.
Ari Turunen
28.2.2020
Read article in PDF
Citation
Ari Turunen, Antti Karlsson, Lila Kallio, & Tommi Nyrönen. (2020). Tissue samples into digital images, interpreted by artificial intelligence. https://doi.org/10.5281/zenodo.8134949
More information:
Auria Biobank
https://www.auria.fi/biopankki/en/
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Breast cancer is the most common type of cancer in women. One quarter of all cancers in women are breast cancers. Until now, genetic risk factors for breast cancer have usually been studied as single factors. Professor Arto Mannermaa intends to explore the big picture and look for more factors which significantly increase the risk of illness when they interact. This is where artificial intelligence comes in.
Mannermaa’s team is developing algorithms that can learn on the basis of genomic and clinical data, and identify and predict risk factors. Learning algorithms are also used in the interpretation of mammography images. Genomic and clinical data are integrated to an AI model that not only helps to determine the risk of illness, but also in drawing up individual treatment plans.
“I’m a biologist and geneticist by training. I engaged in the close study of human genetics and specialised as a hospital geneticist. I have been working on clinical studies in genetics laboratories, but have been interested in cancer throughout my researcher career,” says Arto Mannermaa.
Mannermaa is a Professor of personalised medicine and biobanking at the University of Eastern Finland. In his research, he has focused on the genetics of breast and ovarian cancer. Since its inception, Mannermaa’s team has been involved in the work of the world’s largest genetic epidemiology consortium, the Breast Cancer Association Consortium (BCAC). The consortium has the world’s largest centralised collection of breast cancer tissue samples, collected from over 200,000 patients and controls. The collection includes well-annotated data on factors and clinical results related to breast cancer.
“My research team has been engaged in long-term work to determine the genetic risk variants of breast cancer. We can now identify some 200 variants within normal genomic variation which increase the risk of getting breast cancer. Together with the BCAC, we have also learned about the genetic mutations that are major contributing factors to breast cancer. These include among others mutations of the BRCA and PALB-2 gene, which we were involved in finding to be a contributing factor in breast cancer.”
If a woman has a BRCA1 or BRCA2 gene mutation, she has a 60–80% risk of developing breast cancer. The risk impact of PALB-2 is almost the same.
It is estimated that genome accounts for about 30 per cent of susceptibility to breast cancer, while 70 per cent is determined by environmental factors. Risk factors for breast cancer include the total amount of oestrogen during a lifetime, which depends on the number of pregnancies and children, and weight. Other factors include smoking, alcohol consumption and minor exercise. According to Mannermaa, risk factors tend to have been studied one at a time. Now the BCAC has been able to study the prevalence of breast cancer in the close relatives of patients. This includes plenty of international material that can be used for making comparisons.
“For example, studies have been done on whether there are more common factors in Finnish cancer data than in international data.”
Research material has been obtained from the Biobank of Eastern Finland and Kuopio University Hospital.
“The research team is grateful to all volunteers who participated in the study. Without their consent, this kind of work would not be possible. ”
Finnish data has been compared with data obtained through BCAC, collected from more than 100 research teams around the world. Although data has been obtained from around the world, for Mannermaa the challenge lies in the fact that the material was collected for different purposes, and is not always in the same format.
“In order to use the material, the data collected from different sources must be unified, which often takes up a large amount of the total time spent on research.”

The question Mannermaa wants to answer is which factors have contributed to the onset of breast cancer in a patient. Mannermaa’s team has created an AI model for breast cancer risk factors that is being tested with Finnish and international material.
“We also have material obtained from the Biobank. We are comparing the data of breast cancer patients and healthy individuals and trying to find the interactive combination of all variables that has the greatest influence on the onset of breast cancer.”
One of the study’s targets concerns normal genomic variation, or SNPs. The rapid development of DNA sequencing techniques has made it possible to determine single nucleotide polymorphisms (SNPs), providing a very accurate estimate of the differences between individuals. SNP is the difference in the DNA chain caused by a mutation within a population. According to some estimates, the human genome has 4–5 million SNPs, located in the DNA chain, either in the inter-gene or gene region. They can act as biomarkers, helping researchers to locate genes related to diseases. Certain SNPs can affect the operation of the gene and thus directly affect the onset of the disease.
“In practice, we focus on the differences between cancer patients and control groups. We want to learn how many SNPs are in common with these groups, and what the common SNP network is like among cancer patients compared to healthy individuals.”
Mannermaa’s team is working to identify SNPs related to breast cancer, by means of AI and learning algorithms.
“We teach our algorithm to detect SNP networks. With the help of artificial intelligence, we can identify the interactive group of SNPs with the greatest impact on disease risk.”
The results have been promising. The algorithm helped to identify genes close to SNPs, and these SNPs are probably affecting the operation of the genes. We found a gene network related to oestrogen metabolism.
“Oestrogen metabolism is a key component in the development of breast cancer, while another group that we found was related to apoptosis, or programmed cell death. Apoptosis is crucial in cancer development, because cancer cells must be able to prevent programmed cell death. That’s why we believe that the AI models helped us find the correct breast cancer factors. ”
The amount of data in Mannermaa’s team’s study is so huge that CSC’s (ELIXIR Finnish centre) supercomputing capacity is required.
“About 200,000 SNPs can be identified from one laboratory sample. Each SNP is compared with all the others. In addition, we simulate genetic variation, in other words what SNPs they have in common but remain unidentified. This means that up to another 10 million SNPs can be added to the equation. Add to this variables from imaging and the biobank, and computing capacity is definitely called for.”
The basic model of the Mannermaa team’s AI is based on genetic data. Clinical variables, i.e. breast cancer risk factors, have now been added to this model. Mannermaa believes that the models will significantly improve diagnostics.
“Artificial intelligence enhances screening and diagnostics. In the future, we can avoid overdiagnosis and use the data to differentiate those who need more accurate screening from those who don’t. This means that certain women do not need frequent mammography, due to their low risk of developing breast cancer.
Once genetic data is combined with not only known risk factors but also breast cancer diagnosis and treatment, the predictability of the disease will improve and personalised treatment plans can be drawn up.
Biobanks play a crucial role in research of this type. It is essential that all data is available.
“If the person giving a sample has consented to their data being used for biobank purposes, this data is combined with other data. The Biobank Act is the basis for secure data storage, and enables people who have given their consent to cancel it if they wish. Biobank consent is general consent based on the law. Through biobanks, everyone has the opportunity to participate in research aimed at developing health care.”
Mannermaa leads the SOTE AI Hub project, funded by the Regional Council of Pohjois-Savo. The project is seeking to improve the use of various data sources and AI in aid of decision-making. The project involves utilising and developing health data in a data lake. The Pohjois-Savo data lake consists of social and health data from the Biobank of Eastern Finland, Kuopio University Hospital and the City of Kuopio.
According to Mannermaa, the health data can be used to evaluate the impact of the research results. In addition to receiving plenty of data on the actual patient, we can see the impact of cancer patients’ treatment alternatives and solutions based on new research.
“The model and its prediction can help determine how it affects a patient’s life, and how resources should be allocated. This can make treatment more effective in the future. Patient-specific profiling and individualised treatments help to provide the right treatments for the right patients, and thereby make health care more efficient. This requires a multidisciplinary network.”
Ari Turunen
14.2.2020
Read article in PDF
Citation
Ari Turunen, Arto Mannermaa, & Tommi Nyrönen. (2020). All breast cancer risk factors evaluated with AI. https://doi.org/10.5281/zenodo.8131216
More information:
School of Medicine, University of Eastern Finland
https://www.uef.fi/en/web/laake
Institute of Clinical Medicine, University of Eastern Finland
https://www.uef.fi/en/web/kliinisenlaaketieteenyksikko
Cancer Center of Eastern Finland, CCEF
The Breast Cancer Association Consortium
http://bcac.ccge.medschl.cam.ac.uk
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Cardiovascular diseases are the most common cause of death in the world. More than a third of deaths in Finland are caused by cardiovascular diseases. The current objective is to create an assessment, based on health data, of each person’s risk of illness before they consult a doctor.
Andrea Ganna, Group Leader from Institute for Molecular Medicine Finland FIMM at the University of Helsinki and instructor from Harvard Medical School, wants to establish a nationwide, personalised risk assessment as foundation for planning public health interventions. The assessment is based on the health, demographic and genetic information of the citizens. The assessment, which uses artificial intelligence, improves the allocation of preventive treatments with a lower cost than today.
”Nordic countries and specifically Finland have a unique opportunity and setting, since they have been collecting health and demographic data for years. But the way they have used this data in the past is somewhat outdated. Only very specific correlations and associations in the data have been looked at. However, new methods, such as AI, are emerging, which now allow us to push for a much bigger and ambitious vision.”
Andrea Ganna and his group is developing artificial intelligence (AI) approaches to model health trajectories.
”You have a certain health trajectory and have taken certain medication. We ask if there are other people who have followed a similar path. There may be thousands out there. We leverage those people and ask: What happened to those? Let’s take that experience and bring it back to you to help you to reduce your disease risk. We can use all this data in a more comprehensive way to help public health and give more information to patients and doctors for decision-making.”
Andrea Ganna is interested in epidemiology, genetics and statistics. He has been focusing on leveraging large-scale epidemiological data sets to identify socio-demographic, metabolic and genetic markers of common, complex diseases. In Boston he worked with large-scale exome and genome sequencing data.
According to Ganna, cardiovascular diseases are ideally suited for analysis by artificial intelligence, since their treatment is preventive.
”Accurate identification of individuals at high risk is one of the cornerstones of primary prevention of cardiometabolic diseases,” he says. ”However, at the moment, risk factor assessment for cardiometabolic diseases requires patients to go to the doctor for lipid measurement. ”
Lipid is the umbrella term used for all fatty acids or their derivatives that circulate in the blood. The body stores fats from food for later use. A diet rich in fats will cause them to attach to the walls of the arteries, leading to cardiovascular and arterial diseases. Lipid measurement is effective, but some members of the population are unaware that they belong to a risk group.
Ganna wants to revolutionize primary prevention by providing risk assessment before an individual even steps into the doctor’s office.
”Some simply don’t go to the doctor’s and so lots of people are missed. However, since all the data on medication and diagnoses has already collected, we can identify high-risk patients without them going to doctor. We can make a risk map of cardiovascular diseases of the whole country by including every individual.”
Calculating the risk is done by modelling longitudinal histories of diseases and medications with the gene data, family and demographic data.
”We are trying to understand how genetics interact with data regarding medications, diagnoses, demographics, and familial risk. This can provide an unprecedented holistic view of an individual’s health status.”
Ganna gives an example.
”When you break your leg, you go to the doctor. However, today the doctor is just looking at the leg, although during the same visit other information could also be obtained. We can inform the doctor about other risks the patient has based on the collected data. We can precompute the other risks of the patient, for example if this patient has also high risk for cardiovascular diseases. Thus, during the visit, the doctor can also give advice or refer the patient to a specialist.”

Ganna chose to come to Finland because of the large genetic project, FinnGen. The FinnGen project will record the genomes of half a million Finns. The project, launched in August 2017, utilises samples collected by all Finnish biobanks. The data from genomes is combined with the information in national health care registers. FinnGen is one of the very first personalised medicine projects of this scale and the public-private collaborative nature of the project is exceptional.
“Finland also has a favourable legislation, giving access to nationwide population data. For me, this is a unique setting,” says Ganna.
Ganna and his research group integrate registry-based information with genetic information from large biobank-based studies (e.g. FinnGen) to help identify groups of individuals that can most benefit from existing pharmacological interventions.
”Probably the most important population is younger individuals who do not see a doctor very often. Current risk factors do not work well in this group. Genetics is particularly valuable because it can capture disease risk at an earlier age than other risk factors,” says Ganna.
”The first step is to understand how people perceive this information. We have to ensure that doctors use the data in a right way and what can be done with it.”

Ganna aims to integrate national and regional registries with deep and machine learning.
”Traditional methods have an advantage since they are relative simple and easy to interpret, but they simple do not scale. In the past 20 years, more than 500 million medical diagnoses have been made of Finns. We are talking about huge data sets. Every year there are millions and millions of new medication purchases and diagnoses. To scale and to leverage this massive data, deep learning methods are needed.”
Artificial neural networks are efficient machine learning algorithms, which can be used in pattern recognition. Recurrent neural networks can use their internal memory to process sequences of inputs. This makes them applicable to tasks such as unsegmented recognition. Ganna wants to expand long short-term memory recurrent neural networks to data.
”You can imagine the sequence of health events that we are trying to model as “text” in which each word is a different disease, medication, sociodemographic event etc. across the lifetime. These are naturally suited to model the sequential happening of events, for example they are used to predict the next most likely word in a text message.”
Deep learning methods need large supercomputing infrastructure.
“CSC has created a secure environment for this computation. Without a secure supercomputing environment, we could not carry out this project. To be successful, we need, on the one hand, research and development, and, on the other hand, a powerful computing environment.”
Patient data is important for research, but personal data is also protected. For example, VEIL.AI application created by FIMM anonymises patient data better and faster than traditional methods, and retains information more effectively. If necessary, the application can also produce synthetic, fully anonymous statistical data, which cannot be traced back to any individual.
“We need to guarantee individuals’ privacy but, at the same time, we need to integrate a lot of personal data to really leverage the power of artificial intelligence/deep learning approaches to better target public health interventions. Generating synthetic health trajectories will help to respect privacy and, at the same time, to combine a lot of personal information within Finland, but also across Nordic countries.”
”My hope is that personal data that is routinely collected in healthcare can help and benefit everyone. My hope is that that this information can help doctors to make better decisions, but also help patients in motivating life style changes. Thus everyone is helping everyone.”
Ari Turunen
30.9.2019
Read article in PDF
Citation
Ari Turunen, Andrea Ganna, & Tommi Nyrönen. (2019). Risk assessment of cardiovascular diseases for all citizens. https://doi.org/10.5281/zenodo.8131074
More information:
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org