
Immune-mediated diseases such as allergies, asthma, and autoimmune disorders have increased alongside urbanization. A suspected cause is an overly clean environment, where contact with nature and its microbes is lost. Adjunct Professor Olli Laitinen researches the health effects of microbial exposure at Tampere University. He also serves as Chief Research Officer at Uute Scientific Ltd, a company that produces an extract containing inactivated microbes.
Olli Laitinen’s research group at Tampere University and Uute Scientific have utilized the computing and sensitive data services of Finland’s ELIXIR node at CSC – IT Center for Science in their studies. Over the course of more than ten years, samples have been collected from over 500 individuals, including infants, preschoolers, and adults. Part of the data has been stored in CSC’s secure environment.
Laitinen emphasizes that the root cause of today’s immune-mediated diseases is the loss of contact with nature and living in an overly sanitized environment.
For example, atopic dermatitis is a common, partly hereditary disease that affects approximately 20–30% of the Finnish population. Its symptoms include itching, dryness, roughness, redness, and skin lesions. The underlying cause is abnormal immune system function.
“In the Nordic countries, atopic dermatitis is prevalent. It has been observed that many immune-mediated diseases become more common at the population level as one moves northward.”
The PREVALL project, led by the universities of Tampere and Helsinki, has studied the impact of plant- and soil-based materials on the development of allergies in children. The project has also investigated whether the onset of atopic dermatitis in infants could be prevented.
Read more here:
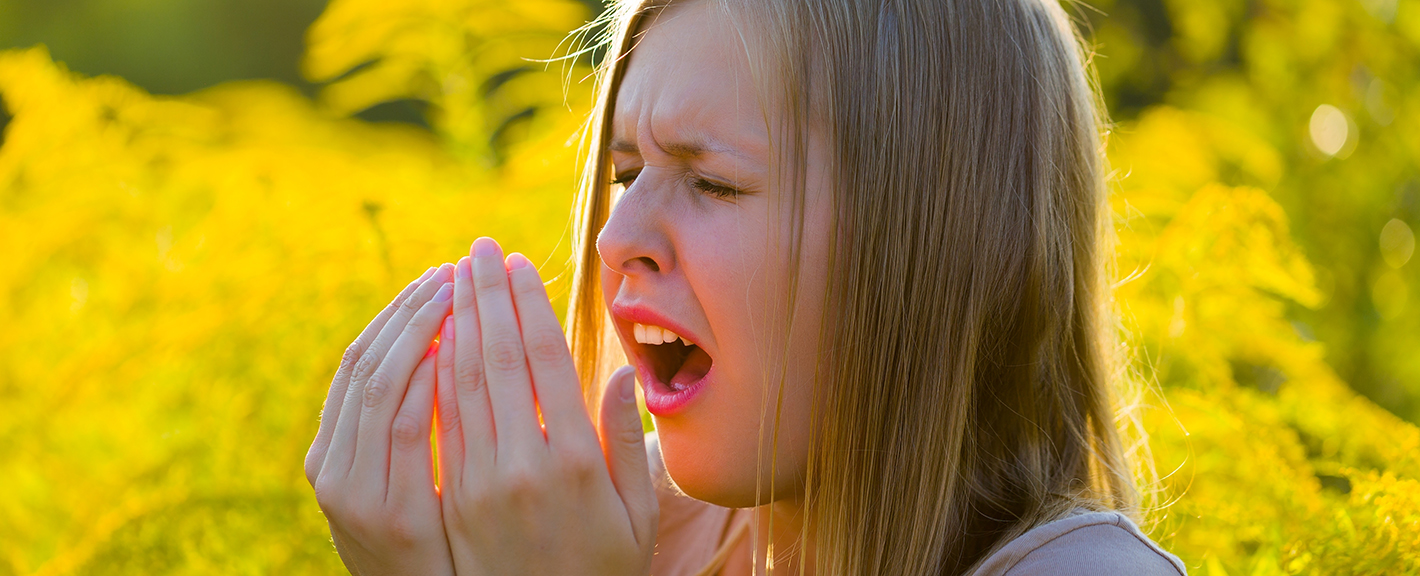

Immune-mediated diseases, such as allergies, asthma and autoimmune diseases, have increased with urbanisation. Researchers suggest that the reason for this is the excessive cleanliness of our environment, as a result of which we have lost touch with nature and its microbes. Adjunct Professor Olli Laitinen studies the health effects of microbial exposure at Tampere University. He also serves as Chief Science Officer at Uute Scientific Ltd. The company produces an extract containing inactivated microbes, which can be used as a raw material in cosmetics, for example.
Microbial exposure can be easily increased by spending time in nature and coming into contact with soil. Handling natural materials with a wide range of microbes changes the microbiota of the body. Laitinen’s projects explore solutions suitable for urban areas to influence the prevalence of immune disorders by modifying the green environment and consumer products.
Laitinen believes that microbial exposure starts at birth.
“When we are born, our bodies are not immunologically complete. At the moment of birth, we encounter practically millions of different life forms. That’s when our immune system starts to learn what is dangerous and what is harmless.”
Laitinen stresses that the root cause of today’s immune-mediated diseases is the fact that we have lost touch with nature and live in an environment that is too clean.
“Humans have lived in natural conditions for hundreds of thousands of years. Mothers have given birth squatting on animal hides, and babies have been wrapped in plant materials. From our first moments, we have been in contact with the soil and nature. We adapted to this exposure.”
“At birth, our bodies get a full blast of microbes and our immune system gets to work. At that point, it is important for the immune system to be able to distinguish between what is dangerous and what is harmless. What is harmless is, of course, our own bodies. However, the immune system also needs to recognise that not all external exposure is dangerous either. Therefore, it is not wise to develop allergies to animal dander, for example. The immune system must learn which microbes are genuinely dangerous and pathogenic.”
Childbirth in a hospital is quite sterile compared to nature.
“If you’re in an environment where the immune system doesn’t get a lot of learning material, the system starts act so that anything external is potentially dangerous. This leads to allergies and asthma, atopic dermatitis, or worse: a situation where the immune system cannot distinguish between the body’s own cells and pathogens, so it starts destroying the former, leading to various autoimmune diseases.”
Promising results have now emerged on how a biodiverse environment can prevent the development of autoimmune diseases, such as type 1 diabetes. Laitinen refers to a part of Noora Nurminen’s doctoral thesis at Tampere University, where she studied the amount of green environment and its impact on the development of type 1 diabetes.
“Type 1 diabetes occurs when inflammatory cells in the immune system are activated in the pancreas and destroy insulin-producing cells. Nurminen examined a cohort of more than 10,000 children to learn how the growing environment during the first year of life influenced the development of diabetes. The results showed that an agrarian environment was healthy for children. Children living in rural areas did not develop diabetes or the autoimmune process leading to it as often as children living in urban areas, or their disease process started much later than that of children living in urban areas.”
The diversity of the microbiota in the human body has declined considerably, especially in the Western world. According to one estimate, urbanised people have 60% of their original microbiota remaining on the skin, and only 50% in the gut. In some areas of the United States, the microbial loss is even higher. As it happens, citizens of the U.S. have more inflammatory diseases than citizens of other countries.
Olli Laitinen believes that the immune system of newborns should start receiving training in the form of exposure to nature when leaving the maternity ward at the latest. Without exposure to nature and its microbes, our bodies’ immune defences cannot function properly. The overreacting immune system may lead to diseases. For example, in the case of allergy, the body misinterprets pollen as a virus.
“We base our research on the function of the immune system and the disruption caused by lack of exposure to nature. The natural role of immunoglobulin E has been to fight parasitic infections, but now that there are far fewer of them, IgE is a free agent looking for new tasks. Such as reacting to pollen.”
Immunoglobulins, or antibodies, are proteins produced by the cells of the body’s defence system. The role of antibodies is to help the defence system destroy invaders, such as bacteria and viruses. IgE-type antibodies have been deprived of their natural activity due to excessive hygiene and sterility, and are therefore inactive. Now, the IgE response is incorrectly activated e.g. against the proteins in pollen, causing allergic hypersensitivity reactions.
Immunoglobulin E is found in allergies and allergic diseases. In allergies, the body produces it to fight off things like pollen or certain foods. The antibody attaches to cells in the skin and mucous membranes and releases histamine. This is what makes us sneeze, our breathing to become obstructed and our eyes swell shut. In developing countries, where parasitic infections are more common, IgE often occurs at high levels without any allergic symptoms.
The “false enemies” of immunoglobulins are a great example of biodiversity loss, which also applies to the microbiota.
“Now that we’re being sold a lot of antibacterial substances, we are actually cleaning away all the bacteria. This is not desirable. It would be better to have a well-established microbiota around us, because that helps us avoid sudden, major changes.”
Changes in the microbiota can cause antibiotic resistance, which is a big problem. Antibiotic-resistant bacteria carry resistance genes and often become dominant in microbial populations.
“Pathogens are usually fast-growing microbes. The abundance of pathogens increases the exchange of genes between them, strengthening their resistance to antibiotics,” says Laitinen, who has also studied antibiotic resistance.
“Hopefully, in the future we will have a safe amount of diverse microbes in our environment so that antibiotic-resistant bacteria cannot thrive.”
Atopic dermatitis is a common, partly hereditary disease affecting about 20–30% of the population in Finland. Its symptoms include itchiness, dryness, roughness, redness and breakouts of the skin. This is due to abnormal immune system function.
“In the Nordic countries, atopic dermatitis is common. It has been observed that many immune-mediated diseases become more prevalent at the population level as we move northwards.”
The PREVALL project, led by the Universities of Tampere and Helsinki, has studied the impact of plant- and soil-based materials on children’s allergies. The project has examined whether it would be possible to prevent the development of atopic dermatitis in babies. Children with both parents diagnosed with atopic dermatitis were included in the study.
“In that case, the child has about a 40% risk of developing the same disease,” Laitinen points out.
In Johanna Kalmari’s and Iida Mäkelä‘s study, a joint project between Uute Scientific and Tampere University, people suffering from atopic dermatitis were given microbial cream containing the extract developed by Uute Scientific. The microbes were not alive, but the cream contained microbial components to which the body and immune defence system can react. In other words, exposure to nature was administered through a cream. The subjects started using the cream in late summer and autumn, as atopic skin gets worse in winter due to dry air and lower temperatures. The lower amount of natural light also has an impact. The subjects used the cream at least three times a week. The researchers took various samples from the subjects, examining the water permeability and redness of the skin, which are indicators of inflammation.
“The biggest difference was seen in the use of medication. The group that used the microbial extract containing cream needed significantly less medication during the trial period of 7 months. The microbial extract containing cream was able to prevent skin deterioration. The cream is a so-called “nature exposure remedy”. It’s a supportive form of treatment that allows patients to use less medication.”
The newest frontier is no more and no less than space itself.
“Astronauts suffer from various skin problems. Not surprisingly, the International Space Station has a very poor microbial environment. Our extract could be taken into space. Discussions have been held with the European Space Agency (ESA) on the use of the cream.”
Olli Laitinen’s research group at Tampere University and Uute Scientific have used the computing and sensitive data services of the CSC, the ELIXIR Node of Finland in their research. Over the course of more than 10 years, the group has been involved in sampling more than 500 individuals, including infants, kindergarten-age children and adults. Some of the data is stored in CSC’s data security environment.
Uute Scientific’s microbial extract is made by combining various plant composts. It contains inactive microbes, which means they are harmless. However, the immune system recognises microbes, microbial particles and also destroyed pathogens. The material was originally developed at the Universities of Helsinki and Tampere. Its biodiversity makes it a unique raw material for cosmetics and other consumer products worldwide. It contains at least 600 different species of microbes.
Ari Turunen
23.6.2025
Read article in PDF
More information:
University of Tampere
https://www.tuni.fi/en
Uute Scientific
https://www.uutescientific.com/fi/
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The ELIXIR Core Data Resources (CDRs) have been selected based on their quality, wide usage, and long-term significance. They are essential to many fields of research, including genomics, proteomics, and drug development. ELIXIR Core Data Resources provide researchers with open and reliable access to biological datasets, promoting new discoveries and accelerating, for example, the development of new drugs, the understanding of diseases, and the identification of biomarkers.
The data analysis services and machine learning models provided by the ELIXIR infrastructure can help identify new drug candidates from large datasets. These resources and databases allow natural compounds to be analysed more quickly and accurately, supporting their development into safe and effective pharmaceuticals.
These are for example ENA, ChEBI and Ensembl. ENA (European Nucleotide Archive) is a database maintained by the European Bioinformatics Institute (EMBL-EBI) that stores and shares sequencing data from various organisms, including microbes, plants, animals, and humans.
ChEBI (Chemical Entities of Biological Interest) is a curated biochemical database that contains information about biologically relevant small-molecule compounds. Ensembl is a genomics and bioinformatics database that provides analysed genomic data from a range of organisms, including humans, animals, plants, and microbes.
The database can be used to look up the biological effects of compound and its target molecules as well as genetic and protein structure data and related genes, aiding research into drug resistance and the effects of mutations.
Read more here:

Tree bark acts as an important chemical defence mechanism against pests. When a plant comes under threat from bacteria or an insect, alkaloids secreted by the plant may, for example, inhibit cell division or the activity of DNA in the insect, preventing reproduction. This is the operating mechanism of paclitaxel and camptothecin, two compounds isolated from the bark of different trees and developed into effective anticancer drugs. Data analyses and databases have now become available to help identify bioactive compounds in trees and other plants.
There are half a million plants in the world, of which an estimated 7% are used for medicinal purposes. Around 25% of prescription medicines in use today are plant-based. This refers to medicines consisting of natural compounds isolated from plants and synthetic derivatives developed from them. Preserving biodiversity is also of paramount importance for pharmaceuticals, as new plant species are constantly being discovered and the chemical composition of even known plants is largely unknown.
Paclitaxel and camptothecin are examples of anticancer drugs that were discovered when samples from potential medicinal plants were systematically screened. The US National Cancer Institute (NCI) screened more than 35,000 plant samples in a research programme that started in 1956 and continued until 1981. The aim of the programme was to identify plant compounds that could be used to prevent or treat cancers.
The ambitious programme also drew on ethnobotany and history. Programme director Jonathan Hartwell compiled an extensive collection of ancient Chinese, Egyptian, Greek and Roman texts on the medicinal uses of plants. To find the samples and obtain accurate botanical information, Hartwell turned to the U.S. Department of Agriculture (USDA). USDA botanists began collecting plants from around the world to be analysed in laboratories.

Research Triangle Institute’s chemists Monroe E. Wall and Mansukh C. Wani received samples of Camptotheca acuminata for study. Known as the Happy Tree in China, Camptotheca acuminata grows naturally on wet banks of the Yangtze River. In traditional Chinese medicine, its leaves and bark have been used to treat various inflammations and infections.
Wall and Wani discovered that the compounds in C. acuminata were highly active in the L1210 mouse leukaemia cell line, meaning that its effects were seen in cancer cells. The L1210 line is commonly used in cancer research and for testing new anticancer therapeutics. It was isolated from a mouse with lymphocytic leukaemia. Wall and Wani isolated an active compound from wood, which was named camptothecin. It was found to be highly effective against leukaemia cells.
Camptothecin binds to an important cellular enzyme, topoisomerase I, and to DNA complexes. This prevents cancer cells from replicating their DNA, resulting in cell death. Despite its effectiveness, camptothecin has serious side effects and poor solubility. A drug being soluble in water is important because it affects the absorption and distribution of the therapeutic agent in the body. Later, derivatives of camptothecin were developed that were water-soluble and better tolerated and retained their efficacy. These include topotecan and irinotecan. Topotecan (Hycamtin) is used for ovarian, lung and cervical cancer, while irinotecan (Camptosar) is used primarily for colon and rectal cancers.
Synthetic derivatives developed from a natural compound can be significantly more effective than the original compound. In the 1980s, the Japanese company Yakult Honsha developed irinotecan, a derivative of camptothecin. It was then discovered that its active form in the body is its metabolic product 7-ethyl-10-hydroxycamptothecin, which is about 100 to 1,000 times more active than irinotecan itself. The compound was given the name SN-38, which stands for the pharmaceutical company code “SmithKline Number 38”. It is not active as such, but acts as a prodrug. SN-38 is a potent anticancer agent that is produced in the body when irinotecan is converted to its active form. Conversion to SN-38 takes place in the liver and other tissues. It is therefore a modified version of naturally occurring camptothecin with added ethyl and hydroxyl groups. These changes resulted in a highly effective therapeutic agent. Some individuals carry the UGT1A1*28 mutation. A mutation in the UGT1A1 gene (such as UGT1A1*28) may reduce the activity of the enzyme and slow down the elimination of SN-38, thereby increasing its toxicity. This may increase the drug’s side effects. The Ensembl database can be used to study the UGT1A1 gene, its mutations and possible effects on SN-38 metabolism, for example.

Wall and Wani continued to study the plant samples after the discovery of camptothecin. They were asked to analyse samples of Pacific yew (Taxus brevifolia).
The Pacific yew is one of five genera in the Taxaceae family. It is a slow-growing tree native to North America, where it is found in the shade of giant conifers on the banks of streams, in deep ravines and in wet passes. Its wood is hard but of limited use. The tree has few natural pests because most parts of it are poisonous. In 1971, Wall, Wani and their colleagues published a study in which they presented a compound isolated from the bark of the yew tree. It prevents microtubules from breaking down, stopping cancer cells from dividing. The compound was named paclitaxel (Taxol).
Paclitaxel was an effective cancer drug, but there were environmental concerns. The extraction of the compound from the yew tree killed the rare tree. As the natural source (yew tree bark) was not sufficient for large-scale production of the drug, a semi-synthetic method was developed in the 1990s using 10-deacetylbaccatin from the needle of the yew tree as the raw material. The compound (10-DAB) is a precursor to paclitaxel, and by adding benzylamine to it, pure and ecologically sustainable paclitaxel can be produced. Paclitaxel is one of the most commonly used medicines for breast and ovarian cancers.
The ELIXIR Core Data Resources (CDRs) have been selected based on their quality, wide usage, and long-term significance. They are essential to many fields of research, including genomics, proteomics, and drug development. ELIXIR Core Data Resources provide researchers with open and reliable access to biological datasets, promoting new discoveries and accelerating, for example, the development of new drugs, the understanding of diseases, and the identification of biomarkers.
The data analysis services and machine learning models provided by the ELIXIR infrastructure can help identify new drug candidates from large datasets. These resources and databases allow natural compounds to be analysed more quickly and accurately, supporting their development into safe and effective pharmaceuticals.
ENA (European Nucleotide Archive) is a database maintained by the European Bioinformatics Institute (EMBL-EBI) that stores and shares sequencing data from various organisms, including microbes, plants, animals, and humans.
Since ENA contains genomic and sequencing data from all forms of life, it is a key resource for biodiversity researchers analysing species’ genetic diversity, population genetics, and evolution. It aids in the identification of new species (via DNA barcoding and metagenomics) and the study of relationships between species (through phylogenetic analyses).
The genetic databases included in ENA enable large-scale meta-analyses and comparisons of genetic information across different populations or species. This supports progress in a wide range of research areas such as evolutionary biology, disease research, and medicine. ENA is openly accessible to researchers worldwide.
ChEBI (Chemical Entities of Biological Interest) is a curated biochemical database that contains information about biologically relevant small-molecule compounds. It provides accurate chemical and biological data on compounds such as drugs, metabolites, and natural products.
ChEBI includes precise information on chemical structure, molecular formula, mass, and isomeric details, which helps researchers analyze the chemical properties of pharmaceutical compounds.
Search example: The database can be used to look up the biological effects of paclitaxel and its target molecules.
Ensembl is a genomics and bioinformatics database that provides analysed genomic data from a range of organisms, including humans, animals, plants, and microbes.
Search example: The main molecular target of paclitaxel is the tubulin protein. Ensembl provides genetic and protein structure data on tubulin and related genes, aiding research into drug resistance and the effects of mutations.
Ensembl includes information on genetic variations that may affect the efficacy and side effects of Taxol. For instance, the enzymes CYP3A4 and CYP2C8, which metabolize Taxol, can carry mutations that impact the drug’s effectiveness.

Ari Turunen
8.5.2025
Read article in PDF
More information:
ELIXIR Core Data Resources
https://elixir-europe.org/platforms/data/core-data-resources
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Vinca alkaloids are the first known naturally-derived anti-cancer drugs. Vinblastine and vincristine are on the WHO list of essential medicines. The Madagascar periwinkle (Catharanthus roseus) is one of the few plants that have directly produced approved anticancer drugs. The names vinblastine and vincristine are derived from the genus Vinca, to which the Madagascar periwinkle belongs. This flower that grows on the island of Madagascar. has saved thousands of children with lymphatic leukaemia.
“It is fascinating that molecules created by the process of mutual survival between plants and insects can influence human biological processes. In nature, an active chemical structure is no coincidence, but repurposing these rare molecules for new uses such as medicine requires innovation,” says Tommi Nyrönen, director of ELIXIR Node of Finland. Nyrönen has studied medicinal substances.
“Natural compounds that may be toxic to one species can, if properly dosed, help another species, as in the case of vinca alkaloids. What’s exciting is what we don’t yet know, because we don’t yet know all the microbes or plants on Earth yet. Similar discoveries can be made in the future by collecting and analysing molecular-level data from research in living nature.”
Information on vinca alkaloids can be found in many databases. For example, ChEMBL, BioStudies, UniProt and Reactome provide information on pharmacological properties, target proteins (such as tubulin), mechanisms, and cellular effects.
“ELIXIR is a data infrastructure on living nature. The databases are part of ELIXIR’s data repositories, which are freely available for scientific research, education and industry,” says Nyrönen.
Read more here:

The Madagascar periwinkle (Catharanthus roseus) is a beautiful flower that grows on the island of Madagascar. It is one of the most important medicinal plants in the treatment of cancer and has saved thousands of children with lymphatic leukaemia. The Madagascar periwinkle is a great example of why biodiversity needs to be protected. It has grown in isolation on the island and developed genome mutations, creating secondary metabolic products that help the plant to survive in the Madagascar ecosystem. More than 200 alkaloid compounds are found in the Madagascar periwinkle, of which vincristine and vinblastine are used in medicinal treatments. Although new cancer drugs are constantly being developed, vincristine and vinblastine, or vinca alkaloids, are still important in medicine.
The biosynthesis, which is the enzyme-catalysed process in which simple compounds are converted into new compounds, of the Madagascar periwinkle was studied for years. The plant’s leaves have traditionally been used in Madagascar to lower blood sugar levels and control diabetes, as well as to treat infections and wounds. When Canadian researchers Robert Noble and Charles Beer started to investigate how the Madagascar periwinkle lowers blood sugar in the 1950s, they found something else interesting instead.
Noble and Beer gave rats flower extracts orally, but no effect on serum glucose levels was observed. The researchers tried a different approach, giving the rats the extract intravenously in the hope that it would boost the blood sugar lowering effect. This led to unexpected consequences: the rats died from bacterial infections. However, the researchers found that the extracts of the plant had an immunosuppressive effect, meaning a strong effect on white blood cells and bone marrow. This led to the discovery of anti-cancer properties in further research. Noble and Beer kept analysing Madagascar periwinkle substances until they identified the active compound, which they named vincaleukoblastine (vinblastine). Vinblastine interferes with intracellular metabolism and inhibits cell division, which means it is a chemotherapeutic agent.
Charles D. Carmichael and Harold P. S. Harington isolated vincristine from the Madagascar periwinkle in the 1950s. Carmichael and Harington worked for the Canadian Cancer Research Foundation, and their research focused on discovering anti-cancer agents in wild plants. Vincristine was one of the effective substances they found to prevent cancer cells from dividing.
At the same time, Gordon Svoboda and Irving Johnson at Eli Lilly and Company were studying plant samples from around the world in the hope of finding plant extracts that could be used to develop cancer drugs. They attended a conference where the Canadian researchers presented their research.
The researchers found they shared a common interest in the Madagascar periwinkle and started cooperation.

Svoboda and Irving studied the effect of vincristine on microtubule formation and cell division. Microtubules are important for many cellular functions, such as division, transport of materials and maintenance of cell structure. Cell cultures were treated with vincristine, which allowed the researchers to monitor the effects of vincristine under the microscope and assess its effectiveness in preventing cell division.
Vincristine and vinblastine are toxic to insects and herbivores. They are indole alkaloids that inhibit cell division and can paralyse or kill insects and herbivores if they eat the Madagascar periwinkle. In humans, the compounds have a different effect and have been shown to help the body fight cancer cells.
Most plant-based anti-cancer drugs target cell division in one way or another. This makes them effective in treating cancer. Because cancer cells divide uncontrollably, many drugs aim to prevent this process. Vincristine and vinblastine, and paclitaxel from the Pacific yew (Taxus brevifolia), target microtubules that form a part of the cytoskeleton. The cytoskeleton is made up of proteins called tubulin, which form long strands. Vincristine and vinblastine bind ß-tubulins and block the formation of strands, preventing cells from dividing normally. All three substances affect microtubule function, but in different ways. They inhibit the cell from dividing into the metaphase stage. In other words, affecting microtubules prevents tumour growth, if the drug makes the structure of the cancer cells unstable.
Vincristine is typically more effective in treating blood cancers such as acute lymphoblastic leukaemia. Vinblastine, on the other hand, has a better effect on solid tumours. It is used to treat Hodgkin lymphoma, non-Hodgkin lymphoma, breast cancer and testicular cancer.
“It is fascinating that molecules created by the process of mutual survival between plants and insects can influence human biological processes. In nature, an active chemical structure is no coincidence, but repurposing these rare molecules for new uses such as medicine requires innovation,” says Tommi Nyrönen, director of ELIXIR Node of Finland. Nyrönen has studied medicinal substances.
“Natural compounds that may be toxic to one species can, if properly dosed, help another species, as in the case of vinca alkaloids. What’s exciting is what we don’t yet know, because we don’t yet know all the microbes or plants on Earth yet. Similar discoveries can be made in the future by collecting and analysing molecular-level data from research in living nature.”

Information on vinca alkaloids can be found in many databases. For example, ChEMBL, BioStudies, UniProt and Reactome provide information on pharmacological properties, target proteins (such as tubulin), mechanisms, and cellular effects.
“ELIXIR is a data infrastructure on living nature. The databases are part of ELIXIR’s data repositories, which are freely available for scientific research, education and industry,” says Nyrönen.
ChEMBL (Chemical Database) is a chemical database that focuses specifically on the interaction between drugs and their target proteins. It allows for the examination of drugs’ biological effects and pharmacological profiles. The database contains information on drug efficacy, safety, and other biological responses.
Metabolism enables the body to transform active drug compounds into less active or more easily excretable forms. These chemical changes are often facilitated by cytochrome P450 enzymes. Drug metabolism affects how long a drug remains active in the body, how quickly it is eliminated, and how effective it is. If metabolism is slow, the drug may persist longer in the body, whereas rapid metabolism shortens its duration of action. Metabolic pathways can vary between individuals due to genetic factors, environmental influences, and interactions with other medications. As a result, two individuals may have different responses to the same drug.
A bioassay is an experimental method used to measure the potency or effectiveness of a substance, such as a drug, chemical, or natural product, based on its biological response. This is particularly important in drug development as it provides valuable insights into how a substance interacts within the body.
Search: The database allows users to search for specific compounds and their bioassay results, particularly assessing their effects on cytotoxicity or receptor responses. It also provides information on interactions between the queried substance and various drug compounds (drug matrix).
The BioStudies database serves as a central repository for storing descriptions of biological studies. It contains links to datasets stored in other databases, as well as data that do not fit existing structured archives. This allows for the storage of a wide variety of study types in a simple format.
ArrayExpress functioned as a database for functional genomics for over 20 years. In September 2022, its user interface was discontinued, and all data were transferred to BioStudies. This transition enhances data integration and accessibility for the research community.
Search: For example, when studying the effect of vincristine on cancer cell growth, BioStudies may contain experimental setups, analysis methods, and results that aid in interpretation.
A drug, such as vinblastine, can have multiple target proteins that it activates, inhibits, or modifies to achieve its biological effects. Drug target proteins can be involved in various biological processes and cell membranes across different organ systems, with their number varying depending on the drug’s structure and function.
UniProt (Universal Protein Resource) is the world’s leading high-quality, comprehensive, and freely available database of protein sequences and functions, maintained by the UniProt Consortium. It provides extensive and detailed information on protein structure, function, interactions, genetic backgrounds, and diseases.
The database is particularly useful in drug development and understanding drug mechanisms, as it helps map how drugs affect protein function. UniProt contains amino acid sequences of proteins, detailing their structures. It includes evolutionary information and species-specific variations. The database is linked to the Protein Data Bank (PDB), which provides three-dimensional structural insights into protein mechanisms and molecular interactions.
UniProt also provides data on how drugs bind to proteins and alter their function, which helps in understanding how drugs influence protein activity and vice versa. Additionally, it offers insights into the genes encoding these proteins, how gene regulation occurs, and how genetic mutations (e.g., through mutations) can impact protein function and contribute to diseases.
Search: The database can be used to investigate the interactions between tubulin proteins and vincristine and its effects on cell division.
The Reactome database focuses on cellular processes and signalling pathways. It is a manually curated database providing insights into biochemical reactions within cells and organs, including protein, RNA, and biomolecular interactions such as signalling pathways, metabolic pathways, and gene expression.
It also contains information on how disruptions in specific biological reactions can lead to diseases, making it valuable for drug development and biomarker discovery. Reactome offers visual pathway maps that depict molecular interactions within biological pathways. For example, vincristine’s effects can be linked to pathways involved in cell division regulation and apoptosis (programmed cell death).
Search: The database enables researchers to explore how vincristine affects different signalling pathways and its overall impact at the cellular level.
Ari Turunen
27.3.2025
Read article in PDF
More information:
ELIXIR Core Data Resources
https://elixir-europe.org/platforms/data/core-data-resources
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
A quarter of patients received medications whose efficacy or safety could have been improved by considering the patient’s genome.
Professor Mikko Niemi’s research conducted a nationwide analysis that included all internal medicine and surgical patients in Finnish hospitals, as well as a group of university hospital patients for whom genetic data was available from the THL biobank.
The nationwide cohort included data from 1.4 million people in Finland obtained from THL-managed registries. Two years after hospitalisation, 60 per cent of patients had purchased a prescription medication for which genetic information is relevant.
According to Niemi, genetic information could be highly beneficial in drug treatment.
“Based on current research, many patients could benefit from adjusting their medication based on genetic information.”
If doctors had access to information about patients’ genetics, medication costs and significant adverse effects could often be reduced, and the number of days of sick leave would also decrease.
Niemi’s research group has used the computing services of Finland’s ELIXIR Node at CSC – IT Center for Science to analyse genetic data. Data management has made use of CSC’s sensitive data platform.
In the future, Europeans will have faster and more accurate diagnoses. Collected and analysed genomic data will enable better drug design and preventive treatments.
Niemi sees it as essential that researchers have access to such infrastructure.
“High-quality genomic data storage is crucial for future research. It ensures that new genetic factors influencing drug efficacy and safety can be identified and their impact can be assessed, and that they can ultimately be put into use.”
Read more here:

Professor Mikko Niemi, a pharmacogenetics expert at the University of Helsinki, studies the impact of genes on the effectiveness and safety of medications. In a recently published study, the medications of 1.4 million Finnish patients were analysed, revealing that a quarter of patients received medications whose efficacy or safety could have been improved by considering the patient’s genome. The study used data from registries of the Finnish Institute for Health and Welfare (THL) and biobank data.
People react to medications differently – some experience insufficient effectiveness, while others may have adverse effects. The reason for varying responses may be our physical characteristics, other medication, or genome. If doctors had access to information about patients’ genetics, medication costs and significant adverse effects could often be reduced, and the number of days of sick leave would also decrease.
In the past five years, genetic testing in healthcare has increased.
“There’s now a wealth of research evidence. The key genes influencing drug response have likely been identified. Many of them regulate the amount of a drug in the body. Often, one gene affects many different types of medications,” Niemi says.

In recent years, various gene panels have been developed to analyse multiple genes simultaneously. This can be considered a breakthrough in healthcare. DNA is extracted from the patient’s blood, saliva or tissue. Massive parallel sequencing allows for the targeted study of many genes at once. The panels can be designed to identify genetic variations that may affect, for example, disease risk, drug response or the occurrence of certain hereditary diseases.
Progress in the use of pharmacogenetic laboratory tests occurred in 2020 with the involvement of the European Medicines Agency (EMA).
“At that time, the agency issued a recommendation to test for hereditary DPYD deficiency before initiating fluoropyrimidine-based cancer treatment. This helps prevent serious adverse effects caused by these anticancer drugs. Testing has been a routine since the agency’s recommendation.”
Pharmacogenetic panels typically test between 10 and 20 genes.
“Humans have 20,000 genes. We know well the effects of 10 to 20 genes on drug treatment. These are key to drug response,” says Niemi.
Helsinki University Hospital’s (HUS) pharmacogenetic gene panel covers the 12 most common and clinically significant genes affecting drug treatments. The selection of these genes took into account international guidelines, drug summaries and the prevalence of genetic variations in different populations. Test results are available in Finnish under the title B -PGx-D, in the MyKanta personal health information online service (https://www.kanta.fi/en/mykanta) MyKanta is an online, publicly accessible service where people can access prescriptions, laboratory test results and healthcare records.
“The idea of the panel is that when the suitability of one medication is tested, the patient also has all other relevant genetic factors for many future medications already tested.”
According to Niemi, with the improvement of testing, more drugs are now known to be influenced by genetics. As a result, drug treatment for cancer, for example, has improved. The use of genetic information in psychiatry has also become more common.
”We are starting to have solid research evidence on the benefits of pharmacogenetics in the treatment of depression. Genetic testing has been included in the Current Care Guidelines for the treatment of depression.”
The Current Care Guidelines (Käypä hoito) are expert summaries published by the Finnish Medical Society Duodecim on the diagnosis and effectiveness of treatments for specific diseases.
The required dosage of individual medications can vary dramatically between individuals, sometimes by more than tenfold. This may depend on how quickly or slowly the body eliminates the medication. Cytochrome enzymes (CYP) play a key role in breaking down and eliminating drugs from the body. There is considerable genetic variation in the activity of CYP enzymes, which can lead to vastly different drug concentrations and responses in individuals.

Currently, there is limited knowledge about how beneficial and cost-effective pharmacogenetic tests would be if the genetic background of all hospital patients were known. Niemi’s research conducted a nationwide analysis that included all internal medicine and surgical patients in Finnish hospitals, as well as a group of university hospital patients for whom genetic data was available from the THL biobank. The biobank contains the FINRISKI data, which holds an exceptionally large amount of diverse health data about the Finnish population, including laboratory tests and health registry data.
The nationwide cohort included data from 1.4 million people in Finland obtained from THL-managed registries. Two years after hospitalisation, 60 per cent of patients had purchased a prescription medication for which genetic information is relevant.
“We tracked purchases of medications where we knew genetics influences drug suitability. By analysing genetic variations, we now know for sure that 99 per cent of people in Finland have a clinically significant genetic variant affecting the response to at least one medication.”
The university hospital sample included 1,000 patients, whose genetic information was available from the biobank. Forty per cent of these patients received medications during their hospital stay for which genetic testing could be beneficial. A quarter of them had a gene-drug combination that researchers do not recommend: the medication should be used at a different dosage, or it would be better to choose an entirely different medication.
“Genetic variation is common and affects widely used medications.”
According to Niemi, genetic information could be highly beneficial in drug treatment.
“Based on current research, many patients could benefit from adjusting their medication based on genetic information.”
The benefits are also significant for society. Finland has excellent registry and genomic data management, and is a leader in the use of pharmacogenetic panels.
“In the future, the aim is to assess the economic and health benefits of pharmacogenetic panel testing. The goal is to examine the treatment costs of Finnish patients who have undergone pharmacogenetic testing and compare this to a situation where genetic testing has not been used. For example, if it were possible to identify the ten percent of patients who benefit the most from genetic information, it could lead to savings in healthcare costs, medications, and sick leave.”
Niemi’s research group has used the computing services of Finland’s ELIXIR Node at CSC – IT Center for Science to analyse genetic data. Data management has made use of CSC’s sensitive data platform.
The Genomic Data Infrastructure (GDI) launched in 2022 aims to create a federated infrastructure for researchers which enables an access to European genomic and clinical data. In the future, Europeans will have faster and more accurate diagnoses. Collected and analysed genomic data will enable better drug design and preventive treatments.
Niemi sees it as essential that researchers have access to such infrastructure.
“High-quality genomic data storage is crucial for future research. It ensures that new genetic factors influencing drug efficacy and safety can be identified and their impact can be assessed, and that they can ultimately be put into use.”
GDI will enable retrospective research, including cost benefit analysis on European scale cohorts, as described by Niemi.
“By linking genetic information to disease and treatment information, GDI helps researchers to discover cohorts with specific treatments and genetic variants across Europe, increasing the size of these cohorts and hence supporting the discovery novel genetic effects on medication”, says senior coordinator Dylan Spalding from CSC. Spalding is the co-lead of GDI Work Package 5.
“For clinicians who have a patient who is not responding to medication as expected, GDI will also enable them to find other clinicians across Europe who may have similar patients with different and more effective treatment regimes, and hence improve the treatment of these patients.”
Ari Turunen
6.2.2025
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2025). Genetic testing improves medication safety and effectiveness. https://doi.org/10.5281/zenodo.14823385
More information:
Value of Pharmacogenetic Testing Assessed with Real-World Drug Utilization and Genotype Data
Kaisa Litonius, Noora Kulla, Petra Falkenbach, Kati Kristiansson, Katriina Tarkiainen, Liisa Ukkola-Vuoti, Mari Korhonen, Sofia Khan, Johanna Sistonen, Arto Orpana, Mats Lindstedt, Tommi Nyrönen, Markus Perola, Miia Turpeinen, Ville Kytö, Aleksi Tornio, Mikko Niemi
https://ascpt.onlinelibrary.wiley.com/doi/full/10.1002/cpt.3458
DOI: 10.1002/cpt.3458
The research was funded by the Research Council of Finland and the Ministry of Social Affairs and Health. The pharmacogenetics pilot was co-designed and implemented by Kaisa Litonius, Mikko Niemi and Katriina Tarkiainen from University of Helsinki and Helsinki University Hospital (HUS), Noora Kulla, Aleksi Tornio, Kristiina Cajanus and Ville Kytö, from University of Turku and Turku University Hospital, Petra Falkenbach and Miia Turpeinen from University of Oulu, Markus Perola, Kati Kristiansson and Liisa Ukkola-Vuoti from the Finnish Institute for Health and Welfare (THL), Arto Orpana, Mari Korhonen, Johanna Sistonen and Sofia Khan from HUS and Tommi Nyrönen and Mats Lindstedt from CSC.
HUS
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Life science research, such as understanding rare diseases or studying the effects of climate change, requires enormous amounts of data and advanced tools to manage and analyze that data. That’s why ELIXIR Finland was created: to build a unified infrastructure that provides researchers in Finland easy access to these critical resources. Now, ELIXIR Finland celebrates a significant milestone – a decade of advancing life science research in Finland.
Hosted by CSC – IT Center for Science, ELIXIR Finland supports Finnish life science researchers by providing access to essential databases, software tools, training materials, and connecting them to supercomputing resources available across Europe.
ELIXIR Finland’s journey began in 2013, when the state of Finland signed the Consortium Agreement to join ELIXIR, the European infrastructure for life science data. By officially becoming a member in 2014, Finland became the 11th country to participate in the ambitious European initiative and entered the global life sciences and bioinformatics landscape. Today, this membership enables Finnish researchers to collaborate with over 240 research institutes across 22 European countries.
In 10 years, ELIXIR Finland has driven progress and impact in Finnish life sciences e.g. in the following ways:
“The impact of ELIXIR Finland has been intense. In 2014, CSC was working with only tens of computing projects in health and biological sciences. Now, we support over 1,600 research projects with our computing infrastructure. The field is still growing, so in five years’ time we expect to more than double the amount of projects through our European collaborations,” says Tommi Nyrönen, Director of ELIXIR Finland at CSC.
Watch the video of ELIXIR Finland’s ten years:
A game-changer in secure cloud services
From the beginning, ELIXIR Finland has worked closely with the life science community to develop services for their needs. In the early 2000s, cloud services were rapidly evolving. CSC launched its first community cloud service in 2013. It quickly became evident that life science researchers needed a secure cloud service to analyze sensitive human data.
This led to the development of ePouta, a secure cloud service designed with data protection as a top priority. It allows researchers to access resources through a secure network connection, either using dedicated fiber optic links or VPN-like technologies. This added a level of security while making it easier for organizations to use cloud resources, as they were integrated into their own networks.
Looking back, ePouta has been a game-changer, enabling researchers to access cloud resources on their own and ensuring that sensitive data can be securely processed. This development work on security laid the foundation for future customer collaboration, such as development of Findata’s and Statistics Finland’s secure processing environments.
Transforming data access management
Early on, CSC recognized the importance of reliable authorization tools to manage sensitive data. The need for such tools became evident when THL Biobank, a part of Finnish Institute for Health and Welfare, sought a more efficient way to control access to their research data. In response, CSC and ELIXIR Finland developed the Resource Entitlement Management System (REMS), which streamlines data access management and helps make data available for reuse.
REMS is a fully electronic system that handles data access applications, records decisions, and issues machine-readable permissions. Its flexibility and security have made it widely adopted not only in Finland but also internationally, including in Australia. Originally designed to manage research data access, REMS has expanded, now also facilitating tasks like ordering death certificates from Statistics Finland, and integrating software into secure environments.
In line with ELIXIR Finland’s commitment to open standards, REMS supports global data access management standards and is freely available on GitHub under the permissive MIT license. This has enabled quick adaptation to regulations like the Finnish Act on the Secondary Use of Health and Social Data, as well as supporting large-scale international collaborations such as the European 1+ Million Genomes initiative.
Advancing secure data management
CSC’s Sensitive Data Services began as a Nordic project to improve the secure management and exchange of sensitive biomedical data across borders. This collaboration, known as NeIC Tryggve project, led to two important insights: the need for a service to store biomedical data while ensuring it remains within national borders, and the development of a new service, alongside ePouta, to manage sensitive data securely.
The Nordic collaboration quickly demonstrated the potential of securely transferring, storing, and accessing data. The decade’s work was realized in 2019 when video files stored in Finland and Sweden were accessed remotely from Oslo, Norway, using a mobile phone, all within a secure environment on ePouta.
Today, CSC’s Sensitive Data Services have evolved into a trusted platform for secure data management. From early prototypes to the fully operational services available today, ELIXIR Finland’s support and networks have been critical in shaping the vision and capabilities of Sensitive Data Services. One of the most recent developments within this platform is FEGA (Federated EGA), a service developed in collaboration with the ELIXIR network and specifically tailored for sensitive biomedical research data. By leveraging a federated infrastructure, FEGA allows genomic datasets to remain under the control of local institutions while still being accessible for collaborative research.
The ComPatAI consortium focuses on analysing histological samples related to breast and prostate cancer. Using digitalised images allows the researchers to measure and automatically compute different cell types.
One of the key questions is how quickly data can be transferred and used. Computing and data storage capacity are constantly in high demand. This is where the services provided by CSC, the Finnish ELIXIR node, come in.
“We’re extremely pleased with the support CSC has given us, since this is an exceptionally large project that uses very large datasets. We are in a privileged position because we have the support of an organisation like CSC. This is a clear competitive advantage for us, and we really appreciate it, ” says Pekka Ruusuvuori, Associate Professor of the Institute of Biomedicine at the University of Turku.
Read more here:

Pekka Ruusuvuori, Associate Professor of the Institute of Biomedicine at the University of Turku, leads the ComPatAI consortium, which is developing new ways to model histopathological tissue samples with generative and predictive AI. In medicine, histological samples are used to assess a patient’s need for treatment. The consortium’s goal is to use big data to create AI models that would produce more accurate diagnostic information in pathology.
In addition, they are developing virtual histological staining models based on generative AI. Besides Ruusuvuori, the consortium consists of research director and Adjunct Professor Leena Latonen of University of Eastern Finland and Teemu Tolonen, Adjunct Professor and chief physician at the department of pathology at the Fimlab laboratories.
The ComPatAI consortium focuses on analysing histological samples related to breast and prostate cancer. Using digitalised images allows the researchers to measure and automatically compute different cell types.
“Our work has focused mainly on prostate and breast cancer. There is ample data available on these types of cancers, as they are the most commonly encountered cancers in men and women. However, we want to create a very general-purpose model that could then be further refined for different and new applications.”
According to Ruusuvuori, the field of pathology is becoming increasingly digitalised. He says that in this sense, the Finnish pathological community is among the pioneers.
“In Tampere and Turku, we have moved completely to using digital pathology in diagnostics. Each time a sample is taken, it is scanned into a high-resolution digital image. There is a lot of routine diagnostics. As the population ages, we encounter more and more patients with cancer. This also means that there is loads of data coming in.”
The consortium receives scanned images of histological slides from Fimlab, the largest healthcare laboratory company in Finland. Fimlab’s clientele includes hospitals, health centres, occupational healthcare service providers and private medical stations. The Finnish Medicines Agency Fimea has currently granted the consortium a licence for using data from 160,050 cases, which translates to a total of approximately 600,000 slide images. Together, the images add up to about 0.8 petabytes of data, meaning that each file accounts for approximately 1.3 gigabytes. The massive amounts of data are currently being anonymised and transferred to the LUMI supercomputer in the CSC – IT Center for Science, an ELIXIR node in Finland. The project is one of the largest data transfers made to LUMI so far.
“It is incredible that we get to use these data for our research. We want to use this data to create AI solutions that function well in pathological work”, Ruusuvuori explains.
The researchers are currently set to have up to 2.5 million digitised whole-slide images at their disposal by the end of the project. This corresponds to a total of three petabytes of data.
“We have been granted permission to use technically all data produced in Fimlab’s routine digital pathology operations.”

Pekka Ruusuvuori has a strong background in signal processing, and he specialises in image analysis. Ruusuvuori is interested in how deep neural networks that are used in AI applications could be developed to be a better fit for diverse use cases.
According to him, machine can generally be taught to recognise the same things that humans would pick up on. It can learn to tell apart different tissue types or to distinguish cancerous tissues from healthy ones. It can be used to measure different factors in images or cells, such as how aggressive a cancer is and how far it has progressed. Artificial intelligence can identify cancerous areas in tissue samples before examination by a pathologist. It may also suggest a score based on the data it has assessed. For example, prostate cancer tumours are given a Gleason score, which indicates how aggressive or advanced the disease is.
“It’s entirely possible to train AI to perform many tasks that human pathologists usually take care of.”
“Previously machine learning models have been built with a certain variable and teaching material that shows a certain object appearing in a specific part of the image and which score this finding corresponds to. It would take countless hours of work for us to mark this information on all the hundreds of thousands of images we are using.”
This annotation data has previously played a key role in teaching artificial intelligence to automatically detect abnormalities such as cancer cells in the samples. Ruusuvuori says that algorithms have been improved and are consequently able to use unannotated raw data.
“I think the most interesting thing about what we are doing with these algorithms is what else we can extract from these images. In other words, to look at the features that machines can detect but humans cannot. These slide images include all visual data that we have. If there is a statistical link to be found there, the machine learning algorithm will find it. However, these links may be extremely complex. Modern neural networks can accurately detect complex links between spatial data and the predicted variable. These things can be very difficult for us humans to grasp.”
Together with his research group, Ruusuvuori has been able to successfully predict gene expression and mutations directly from histological images. Gene expression refers to the process of a cell producing the molecules that is encoded in its DNA. The gene expression varies across different tissues. Based on the images, AI can detect miniscule changes that are invisible to the human eye.
“Based on the images, the machine can identify effects of gene expression in cells and tissues. It can detect even the slightest phenotype variations, including those that we as humans are not trained to see. I want to highlight that so far, we only have indicative results, and that the method will not work for all tissue types or genes. Some gene expressions do not lead to tissue-level changes that could be predicted from a whole slide image.”
ComPatAI consortium is currently developing a so-called foundation model for utilising large datasets. As the name suggests, this model would create a general-purpose foundation for developing further AI solutions. The model is trained in histology based on a large set of samples, without target variables or annotation data.
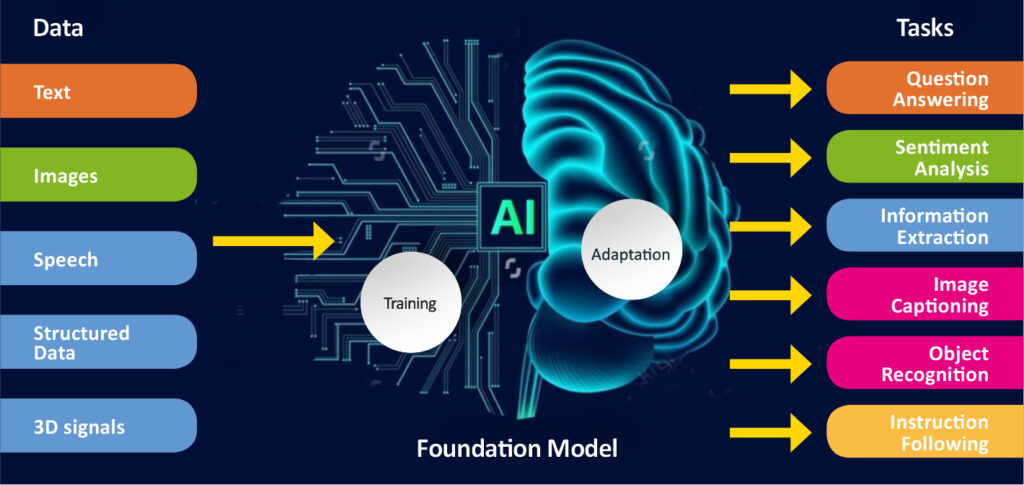
“When you start teaching a model like this to recognise diseases such as breast or prostate cancer, it starts to learn to perform the task it has been given. This will allow us to reach more accurate solutions much faster than before. It allows us to use large, unannotated datasets. This is a big step forward for us.”
ComPatAI consortium is currently building its own foundation model based on a Finnish dataset.
“This is basic research that will allow us to be among the first to further refine these models in Finland. I do not want us to rely solely on big foreign firms or research groups, and instead wish that we would be able to be build a model based on Finnish data. We have high-quality, population-level cohort data that need to put to good use. I hope that this will lead to the establishment of companies in Finland that will develop solutions to benefit patients in routine diagnostics.”
One of the key questions is how quickly data can be transferred and used. Computing and data storage capacity are constantly in high demand. This is where the services provided by CSC, the Finnish ELIXIR node, come in.
“We’re extremely pleased with the support CSC has given us, since this is an exceptionally large project that uses very large datasets. We are in a privileged position because we have the support of an organisation like CSC. This is a clear competitive advantage for us, and we really appreciate it.”

Digital pathological data and other potentially sensitive health data types, such as registry and omics datasets, are going to become more readily available through CSC’s data secure user environment.
“We’re just getting started with the development work”, says Tommi Nyrönen, who leads the ELIXIR Finland Node.
“ELIXIR’s node in Finland has helped transform the biomedical resources required by CompPatAI to a platform service operated by CSC. The CSC Sensitive Data platform emerged from this need. However, it continues to serve various other researcher projects in the field. One of these is the EU’s digital pathology archive initiative bigpicture.eu, which is set to launch in 2026. It is a sustainable solution for managing digital pathology datasets and bringing them to high-performance computing services across Europe.”
Ari Turunen
26.12.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). The ComPatAI consortium uses large datasets to create an AI learning model for pathology. https://doi.org/10.5281/zenodo.14823370
More information:
FIRI
Article was supported by the Research council of Finland under grant number 345591 for ELIXIR European Life-Sciences Infrastructure for Biological Information (FIRI 2021)
Ruusuvuorilab
Fimlab
University of Turku
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Associate Professor Mikael Niku from the University of Helsinki wants to determine the kinds of mechanisms the maternal microbes use to modulate the development of the immune system of the offspring.
His research group carries out data analyses using the equipment of the CSC – IT Center for Science, the Finnish ELIXIR node.
The microbes process the substances our bodies absorb, turning them into metabolites.
“In the past, our group – like many other research groups – focused on investigating whether maternal bacteria are transferred from the mother to the foetus. After all, almost no living bacteria is present in a healthy foetus. However, we now know that microbes produce small-molecule metabolites that are then transferred to the foetus.”
Niku is interested in finding out what sort of metabolites are transferred to foetuses and how these metabolites affect the foetal development.
“The metabolites are absorbed from the gut into the bloodstream, from where they’re transferred to placenta and to the foetus. We found that the concentrations of some metabolites are associated to the functioning of genes in the foetus. These genes are often linked to the immune system and its development.”
The research involves analysing microbiomes using amplicon sequencing, targeting the 16S ribosomal RNA (rRNA) gene region. This makes it possible to study the microbiota composition. The 16S gene regions are sequenced and identified through publicly accessible databases.
According to Niku, before long, it can be determined what kind of bacteria and bacterial products the foetus needs to develop an optimal immune system.
“For example, it could allow us to develop probiotic products that contain necessary microbes or microbial products that are currently unavailable.”
Read article here:

Mikael Niku’s team studies foetuses and how the body’s microbiota develops after birth. Niku is especially interested in how the mother’s microbiota affects the foetal development and immune system in different mammals.
It is a long-established fact that after birth, microbes in the mothers’ body are passed on to the offspring. This prepares the child for life outside the womb.
“The mother’s microbiota also affects immune system development. The immune system learns to accept beneficial gut microbes and to fight pathogens. A diverse microbiota will naturally ward off pathogens,” says Mikael Niku, Associate Professor at the Developmental Interaction Lab at the University of Helsinki.

Niku’s aim is to determine the kinds of mechanisms the maternal microbes use to modulate the development of the immune system of the offspring. His research group carries out data analyses using the equipment of the CSC – IT Center for Science, the Finnish ELIXIR node.
The research involves analysing microbiomes using amplicon sequencing, targeting the 16S ribosomal RNA (rRNA) gene region. This makes it possible to study the microbiota composition. The 16S gene regions are sequenced and identified through publicly accessible databases.
A large proportion of the immune system’s cells are in the intestines. They develop from blood stem cells, taking the form of different types of white blood cells. Diet, lifestyle and the medicines a person takes and the chemicals in their environment all affect the intestinal microbiota. The microbes process the substances our bodies absorb, turning them into metabolites.
“In the past, our group – like many other research groups – focused on investigating whether maternal bacteria are transferred from the mother to the foetus. After all, almost no living bacteria is present in a healthy foetus. However, we now know that microbes produce small-molecule metabolites that are then transferred to the foetus.”
Niku is interested in finding out what sort of metabolites are transferred to foetuses and how these metabolites affect the foetal development.
“The metabolites are absorbed from the gut into the bloodstream, from where they’re transferred to placenta and to the foetus. We found that the concentrations of some metabolites are associated to the functioning of genes in the foetus. These genes are often linked to the immune system and its development.”
One thing that Niku’s research group is currently studying is extracellular vesicles that are produced by bacteria. Vesicles are small, liquid-filled sacs of membrane that are produced by both animal and bacterial cells. They are found in all bodily fluids. Although vesicles were discovered as early as 1946, research on them took off in earnest only in the 2000s. Vesicles contain various cellular products.
“Vesicles may play an important role in things such as recycling materials in the body, cell-to-cell communication, immune regulation and various diseases, among other things.”
Researchers at the University of Oulu were the first team in the world to publish a study that showed that vesicles are transferred from the maternal microbiota to the foetus. They discovered a previously unknown interaction mechanism between the maternal microbiota and the developing foetus.
Niku and his team is further investigating how the foetal immune system develops before birth. Is recognising the good bacteria that the body should not attack something that the foetus’ immune system learns already at this stage?
“It’s possible that vesicles carry bacterial macromolecules, such as proteins, into the foetus, thus training its immune system to discern good bacteria from pathogens. This would allow the offspring to recognise the gut microbes of their mother or their species prior to birth.”
The team’s next focus is on how vesicles present in foetal tissues, how they are able to pass through the placenta and how they affect the foetus.
According to Niku, before long, it can be determined what kind of bacteria and bacterial products the foetus needs to develop an optimal immune system.
“For example, it could allow us to develop probiotic products that contain necessary microbes or microbial products that are currently unavailable.”
Ari Turunen
14.11.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Microbiota affects the immune system. https://doi.org/10.5281/zenodo.14823362
More information:
Developmental Interaction Lab at the University of Helsinki.
https://www.helsinki.fi/en/researchgroups/developmental-interactions
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Without exposure to nature and its microbes, our immune system dos not function as it should. Mira Grönroos, community ecologist at the University of Helsinki, is interested in how spending time in the forest and connecting with nature affect the skin microbiota. One study increased the interaction between daycare children and natural microbiota. The studies showed, for the first time in the world, that the children’s immune system regulation changed while they were in contact with the diverse microbiota of natural materials.
The microbiota collected from sand, skin and gut was sequenced. The study examined how the microbiota changed between the test group and control group. In the study, the gene region of 16S ribosomal RNA (16S rRNA) was sequenced and the bioinformatics was carried out with the resources of Finnish ELIXIR Node, CSC – IT Center for Science. The 16S gene regions have remained unchanged for millions of years of bacterial evolution, which is why they can be used to identify different species.
The samples taken from the children’s skin helped identify the composition of the bacterial community, i.e. the metagenome. The relative amount of more than 30 bacterial genera increased on the children’s skin. The increase in the amount of immune-boosting gammaproteobacteria was connected to a change in interleukin-17A, which is associated with the development of allergies and immune-mediated diseases.
“Efficient sequencing methods and the data they generate are vital for studying microbial diversity and its impacts. Cultivation methods alone aren’t enough for studying things like this,” Grönroos says.
Next-generation sequencing technology enables the simultaneous sequencing of millions or even billions of DNA segments in a sample.
Read more here:

Just as those in the gut, the microorganisms in the skin play an important role in improving the body’s immune system. Mira Grönroos, community ecologist at the University of Helsinki, is studying the connections between the environment, microbiota and human health. She is interested in how spending time in the forest and connecting with nature affect the skin microbiota. Her aim is to find ways to improve the human immune system. This topic has not been studied much before.
Allergologist Tari Haahtela has made a special biodiversity hypothesis about human health: without exposure to nature and its microbes, our immune system dos not function as it should. If there is little or no interaction with nature, the immune system can’t learn to distinguish between what is dangerous and what is not. The body goes into a state of stress, which results in low-grade inflammation. The immune system overreacting may lead to diseases.
Mira Grönroos is working as a postdoctoral researcher in a multidisciplinary NATUREWELL project (2019–2025) funded by the Research Council of Finland. The project, led by Docent Riikka Puhakka, aims to study the impact of outdoor recreation on the health and well-being of Finnish youth. In the project, Grönroos focuses on studying how outdoor recreation and activities affect the human microbiota.

“The youth participated in a variety of outdoor activities in nature. Microbial samples were taken from their skin before and after the activities. We are studying whether hiking in a forest or spending time in urban nature changes the microbiota of the youth. We are also looking for ways to encourage young people to go out in nature,” Grönroos explains.
Grönroos is part of a research group led by Aki Sinkkonen, who works as a senior researcher at the Natural Resources Institute Finland (Luke). For the other studies in the group, researchers have been measuring interleukin and T-cell levels, for example. Cytokines, which are small proteins, function as messengers in the system controlling cellular functions in the body. These include interleukins, which increase or decrease inflammation. T cells help destroy pathogens living inside cells. B cells, on the other hand, are responsible for antibody-mediated immunity. The studies found that the levels of anti-inflammatory interleukin-10 proteins increased after microbial exposure.
According to Grönroos, the immune system and microbes are in constant interaction.
“The results so far are very encouraging, and now we’re studying the intensity of the natural exposure required. Spending time in nature also has many other wellbeing benefits that make the forest a good place to be. One way to increase microbial contact while enjoying nature is to eat snacks without washing or sanitising hands first,” says Grönroos.
Sinkkonen’s research group has been carrying out intervention studies, where the researchers’ intervention in the phenomenon under study is an integral part of the method. One study increased the interaction between daycare children and natural microbiota. The study followed daycare children between the ages of three and five in ten daycare centres in Lahti and Tampere for a month.
“The yard of the daycare centre was made greener to increase the children’s contact with natural materials. In another study, material containing microbiota was added to the sand in the yard,” Sinkkonen says.

The studies showed, for the first time in the world, that the children’s immune system regulation changed while they were in contact with the diverse microbiota of natural materials.
The microbiota collected from sand, skin and gut was sequenced. The study examined how the microbiota changed between the test group and control group. In the study, the gene region of 16S ribosomal RNA (16S rRNA) was sequenced and the bioinformatics was carried out with the resources of Finnish ELIXIR Node, CSC – IT Center for Science. The 16S gene regions have remained unchanged for millions of years of bacterial evolution, which is why they can be used to identify different species.
The samples taken from the children’s skin helped identify the composition of the bacterial community, i.e. the metagenome. The relative amount of more than 30 bacterial genera increased on the children’s skin. The increase in the amount of immune-boosting gammaproteobacteria was connected to a change in interleukin-17A, which is associated with the development of allergies and immune-mediated diseases.
“Efficient sequencing methods and the data they generate are vital for studying microbial diversity and its impacts. Cultivation methods alone aren’t enough for studying things like this,” Grönroos says.
Next-generation sequencing technology enables the simultaneous sequencing of millions or even billions of DNA segments in a sample. Sinkkonen’s research group has also started random sequencing, or shotgun sequencing as it’s called.
“This method gives us more detailed information about the taxonomic profile of the entire microbiome and its functionalities, such as genes and metabolic pathways,” Sinkkonen says.

Mira Grönroos’s research is multidisciplinary, and now also includes a social and pedagogical perspective. The aim is to promote interaction with nature. The previous studies conducted with daycare children already showed that children love playing with natural materials. A recently started project at Tampere University studies children’s view on microbes.
“Microbes are made visible through both science and art. I’m in charge of the part about science. The children can decide what they want to take a sample of. They also get to follow the progression of the samples in the laboratory via video messages. Then I’ll present the results of the sequencing to the children.”
Ari Turunen
21.10.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). The skin’s wide range of microbiota improves the immune system. https://doi.org/10.5281/zenodo.14823352
More information:
University of Helsinki
https://www.helsinki.fi/en/researchgroups/nature-based-solutions
Natural Resources Institute Finland
https://www.luke.fi/en/projects/biwe
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Piia Bartos from University of Eastern Finland has been studying RNA-binding proteins (RBPs), which may play a role in cancer treatment. RBPs have been found to play a role in cancer cells, particularly in drug responses and the development of drug resistance. More than 1,500 RBPs have been discovered so far. Changes in the function of these proteins can affect the level of cancer gene expression.
“We’re particularly interested in argonaute proteins which play an important role in RNA-mediated gene silencing. The most important of these is Ago2,” says Bartos.
When RNA is bound to Ago2 protein, this combination is called the RNA-Ago2 complex. Argonaute 2 protein binds microRNA molecules in cells.
“As argonaute-2 is a protein that’s vital for cell function, it’s likely to affect all types of cancer. If it is removed from the cells, the cells will not survive. If its activity could be eliminated in cancer cells, those cells would not survive. This would prevent the growth and spread of cancer cells.”
Molecular dynamics simulations provide insights into how biomolecules interact with each other at the atomic level. Because atoms are in constant motion, the forces between them are calculated and used to determine factors such as the new positions, velocities and energies of the protein atoms. This will provide new information for drug design. Simulations of molecular dynamics can be used to make a kind of video of the movements of Ago2-RNA complexes and to compare the differences between activating and silencing complexes.
The RNA sequence data used in the simulation was obtained from the A.I. Virtanen Institute for Molecular Sciences. Six RNA molecules were used in the simulations, three of which increased protein production and three of which decreased it. For all of these, molecular dynamics simulations were run for about 50 microseconds, or a millionth of a second per system. The simulations placed high demands on the computing resources of the Finnish ELIXIR Node, CSC – IT Center for Science.
Read more here:

Piia Bartos, a senior pharmaceutical researcher at the University of Eastern Finland’s School of Pharmacy, is interested in RNA, the proteins that bind RNA, and how this system can be influenced to prevent cancer growth. She studies RNA and the function of the argonaute protein that binds to it using massive simulations.
Molecular dynamics simulations provide insights into how biomolecules interact with each other at the atomic level. Because atoms are in constant motion, the forces between them are calculated and used to determine factors such as the new positions, velocities and energies of the protein atoms. This will provide new information for drug design.
Bartos has been studying RNA-binding proteins (RBPs), which may play a role in cancer treatment. RBPs have been found to play a role in cancer cells, particularly in drug responses and the development of drug resistance. More than 1,500 RBPs have been discovered so far. Changes in the function of these proteins can affect the level of cancer gene expression.
RNA interference (RNAi) is a biochemical mechanism whereby RNA causes the cleavage of messenger RNA in the cell, disrupting gene expression. The researchers who discovered RNAi, Andrew Fire and Craig Mello, were awarded the Nobel Prize for medicine in 2006. RNAi can be used to switch off the expression of proteins that promote cancer growth.
“We’re particularly interested in argonaute proteins which play an important role in RNA-mediated gene silencing. The most important of these is Ago2,” says Bartos.
When RNA is bound to Ago2 protein, this combination is called the RNA-Ago2 complex. Argonaute 2 protein binds microRNA molecules in cells.
“As argonaute-2 is a protein that’s vital for cell function, it’s likely to affect all types of cancer. If it is removed from the cells, the cells will not survive. If its activity could be eliminated in cancer cells, those cells would not survive. This would prevent the growth and spread of cancer cells.”

The challenge is that two types of RNA molecules can be bound in the RNA-Ago2 complex. The first inhibits protein production, whereas the second increases it. In the latter case, the production of cancer cells may increase.
“I simulated the function of RNA separately, and also with the Ago-2 protein. I have tried to clarify how Ago-2 complexes differ structurally – that is, when they contain RNA that increases protein production, and when they contain RNA that decreases protein production. We’ve just finished running the simulations and we’re now analysing the results.”
Simulations of molecular dynamics can be used to make a kind of video of the movements of Ago2-RNA complexes and to compare the differences between activating and silencing complexes.
The RNA sequence data used in the simulation was obtained from the A.I. Virtanen Institute for Molecular Sciences. Six RNA molecules were used in the simulations, three of which increased protein production and three of which decreased it. For all of these, molecular dynamics simulations were run for about 50 microseconds, or a millionth of a second per system. The simulations placed high demands on the computing resources of the Finnish ELIXIR Node, CSC – IT Center for Science.
“It’s a fairly big protein. Along with the RNA and the surrounding water, there are about 300,000 atoms, and we had to calculate the speed and position of all of them every four femtoseconds.”
A femtosecond is a millionth of a billionth of a second. Bartos is aiming to find out whether the shape of the complex changes, and whether a part of the protein moves differently when it has an increasing or decreasing RNA bound to it.
“It’s likely that the change in the shape of the complex can indicate that the complex binds to different proteins.”
There must therefore be a difference in the structure or movement of the complexes that causes different effects that either increase or decrease gene expression.
By understanding the structural differences between RNA-protein complexes that reduce and increase gene expression, it is possible to design and screen drugs that bind only to the desired complex. According to Bartos, such drugs would be a medical breakthrough, and would offer a new way to treat cancers where protein production is impaired.
“RNA interference-based drugs are a good alternative. These drugs could be more specific and better targeted to the cancer cell than a standard small-molecule cancer drug. With RNA interference, we could, if necessary, block the expression of any protein in a cancer that we wanted to block. So this would give highly selective drugs.”
According to Bartos, however, modelling the function of RNA is still a challenge. In simulations, force-field models work well for proteins, but not for RNA.
“The reason for this is that RNA is chemically and physically quite different from proteins.”
An example of a problem is the phosphate that forms the RNA strand with deoxyribose.
“The phosphate in RNA is electrically charged and is not very well modelled by these current force-field equations. So there’s clearly a lot of work to be done in developing the tools.”
Drug design has been making great strides on many levels. DeepMind’s artificial intelligence AlphaFold can already solve how a sequence becomes a protein structure. It uses known protein structures, and predicts the structure for all known proteins.
Sequencing can be used to identify mutations in cancer, and models can be used to study how mutations affect the action of anticancer drugs.
“For example, the mutation may prevent the cancer drug from binding to the target protein at the drug target, in which case the patient will rarely benefit from the drug.”
As computing capacity increases, it will also become possible to simulate larger entities.
“It would be great to simulate a single protein at a larger unit, for instance at the cellular level. We could simulate how the protein interacts with other proteins, cell membranes and cell organelles.”
Ari Turunen
30.9.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). New drug targets from RNA-binding proteins. https://doi.org/10.5281/zenodo.14810576
More information:
Hanna Baltrukevich & Piia Bartos: RNA-protein complexes and force field polarizability. Front. Chem., 22 June 2023
Sec. Theoretical and Computational Chemistry
Volume 11 – 2023 | https://doi.org/10.3389/fchem.2023.1217506
Milla Kurki et all: Structure of POPC Lipid Bilayers in OPLS3e Force Field. Journal of Chemical Information and Modeling. Vol 62/Issue 24
https://pubs.acs.org/doi/full/10.1021/acs.jcim.2c00395
University of Eastern Finland
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The University of Eastern Finland performed a virtual search of 1.56 billion molecules to test two drug candidates. This was the world’s most extensive screening of its kind.
Professor Antti Poso’s research team were looking for molecules that would react with SurA chaperone and cyclin-G-associated kinase (GAK), two candidates with medicinal effect. The project tested the HASTEN algorithm developed for the screening, and created a new machine learning model.
“The machine that had been trained completed the screening much more quickly than would have been possible with the traditional docking method. While the calculation of docking took a couple of months even using powerful computers, with machine learning the learning process and prediction only took a few days.”
“Obviously it follows that with supercomputers we can take even bigger databases and screen thousands of billions of molecules with this method.”
The research made use of the supercomputing resources, data storage and tool containerisation of the Finnish ELIXIR Node, CSC – IT Center for Science.
Read more here:

The University of Eastern Finland performed a virtual search of 1.56 billion molecules to test two drug candidates. This was the world’s most extensive screening of its kind.
Most drugs available today have been designed so that the target molecules are the body’s own proteins. Once the structure of one member of a protein family has been determined, the structure of other proteins in the same family can be predicted through modelling. A successful drug can be developed, for example, by screening a large library to find a molecule with a three-dimensional structure enabling interaction with the target protein.
Professor Antti Poso’s research team were looking for molecules that would react with SurA chaperone and cyclin-G-associated kinase (GAK), two candidates with medicinal effect. The project tested the HASTEN algorithm developed for the screening, and created a new machine learning model.
“These target proteins, SurA and GAK, were already known to us from existing academic research projects. The results of the massive screenings can be used in other research. We not only just validated a method but are also able to help various academic research projects,” says Poso.
Chaperones contribute to protein folding and regulate protein interaction. Kinases have a role in cellular signalling, among other things.
“The SurA chaperone is related to a collaborative project with the University of Tübingen, with the aim of developing new antibiotics. Kinases, on the other hand, are a large family of proteins. Most cancer drugs are kinase inhibitors. There are some 500 types of kinase, with cyclin-G-associated kinase, or GAK, being one of them. GAK’s potential lies in cancer drugs and the treatment of viral infections.”
Poso’s team is studying the interaction of drugs and proteins, and creating target protein models. The point at which a drug binds to a protein can usually be identified in the target protein structure, thereby making the drug work. The model can be used specifically in virtual screening. This involves searching large molecular databases for new ideas for drug development.
“Chaperone’s protein structure is very different from that of kinase. So we are talking about two very different target proteins that were worth testing together.”

The structural difference of two drug candidates was a key factor, because the algorithm must work in all protein families.
“Two drug candidates were used to test how the HASTEN algorithm developed by Tuomo Kalliokoski at Orion works in the CSC supercomputing environment. The scalability was successful.”
The target protein screening was performed, for purposes of comparison, with the HASTEN algorithm and the traditional docking method. In docking, the search algorithm calculates the interactions between the protein and the drug candidate in the database. The value given by the algorithm shows how well the drug binds to the protein.
Poso’s team screened 1.56 billion molecules containing the drug candidate. The molecules were screened from the REAL database of Enamine, a large Ukrainian chemical company.
“First we calculated every two-dimensional molecule drawn in the database and converted them into three-dimensional format. After that the software tried to fit each molecule inside GAK or SurA. An individual fitting can have hundreds of thousands of alternatives.”
Then the researchers tested how machine learning fared compared to docking. The HASTEN algorithm was used for machine learning.
“We first chose a million molecules at random to see how the docking worked. We then fed the results to AI. So what the machine did was learn to predict the result on the basis of a million molecules, meaning that when a molecule has a specific shape, it docks into a specific location.”
After this, all 1.56 billion molecules were fed in to the AI to predict results using the results of the initial million molecules. The ones that had the highest prediction were docked again, followed by another round of machine learning. After a few rounds the AI was able to predict docking to the accuracy of 90 per cent.
“The machine that had been trained completed the screening much more quickly than would have been possible with the traditional docking method. While the calculation of docking took a couple of months even using powerful computers, with machine learning the learning process and prediction only took a few days.”
According to Poso, researchers can new routinely screen billions of molecules in the same time that previously only managed a million. And thanks to the machine learning model, billions of molecules can now be screened without a supercomputer.
“Obviously it follows that with supercomputers we can take even bigger databases and screen thousands of billions of molecules with this method.”
The next thing Poso’s team will be looking at is what is known as the vivid screening method.
“Instead of just predicting a single activity or docking, we can simultaneously predict a number of different properties, such as predicting a docking that can cause side effects, while maintaining solid docking to a good location.”
The research made use of the supercomputing resources, data storage and tool containerisation of the Finnish ELIXIR Node, CSC – IT Center for Science.
Ari Turunen
31.8.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). New machine learning method speeds up drug screening hundred-fold. https://doi.org/10.5281/zenodo.13691983
More information:
Toni Sivula, Laxman Yetukuri, Tuomo Kalliokoski, Heikki Käsnänen, Antti Poso & Ina Pöhner (2023): Machine Learning-Boosted Docking Enables the Efficient Structure-Based Virtual Screening of Giga-Scale Enumerated Chemical Libraries. J. Chem. Inf. Model. DOI: 10.1021/acs.jcim.3c01239. Available at: https://pubs.acs.org/doi/full/10.1021/acs.jcim.3c01239
HASTEN algorithm
https://github.com/TuomoKalliokoski/HASTEN
University of Eastern Finland
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
An international research consortium including researchers from the University of Helsinki and Nanyang Technological University in Singapore has compiled the genome of three coffee species from DNA sequences of coffee plant leaf cells. The genome of Arabica, or Coffea arabica, was assembled in Singapore and Helsinki, and those of Robusta (C. canephora) and C. eugenioides were assemled at Cornell University in the United States.
“The aim is to find traits that improve yield and quality. Cultivated Arabica is less genetically diverse, and therefore more susceptible to disease,” says Salojärvi, an assistant professor at Nanyang Technological University.
Salojärvi holds positions both at Nanyang Technological University and the University of Helsinki. The very broad and international research team makes extensive use of computational resources and databases in both countries.
Establishing a chromosome-level assembly required also the determination of the three-dimensional structure of the chromosomes. The computing resources of the CSC– IT Center for Science, the Finnish ELIXIR Node, were used for this task. In the process the contiguous coffee genome sequences assembled in earlier steps were combined into chromosome-length scaffolds.
“This process means that a chromosome is made up of fully sequenced fragments and empty spaces in between. Among other things, the structural analysis will reveal the connection with genes and the regions of the genome that regulate them.”
All three coffee genomes have been shared via EBI/NCBI. Additionally, these annotations have been made accessible via ORCAE, a database with tools to further work on the gene structures, and containing annotations of diverse eukaryote genomes. It is operated by the Belgian ELIXIR Node.
Read more here:
The study led research professor Markus Perola of the Finnish Institute for Health and Welfare (THL)involves collecting data on more than 3,000 people who were hospitalised with the virus, or, in milder cases, sought coronavirus testing. The survey uses registry data. Sample collection is carried out in collaboration with biobanks. Blood samples are tested for other co-infectious diseases, the severity of the inflammation, and other values that indicate the biological balance of the body.
“There is always talk of different risk groups, but we forget that a large proportion of people at risk of the coronavirus either do not end up in intensive care or do not die of the disease. For example, the mortality rate for people over the age of 80 is around 10 per cent, but several times that number do not die. So what is the difference between these groups? And why do some very overweight people end up in intensive care with the coronavirus, but not others? Our aim is to find the groups at risk to benefit from vaccination the most.”
Another thing that was studied was respiratory syncytial virus (RSV) infection in children under the age of 1. Respiratory syncytial virus (RSV) is a ribonucleic acid (RNA) virus that causes millions of respiratory infections worldwide every year. It is a major cause of infections in young children.
“The registry data was used to follow families whose child had been hospitalised after contracting RSV. The study used data related to socioeconomic status, use of intoxicants by the child’s parents, and the child’s birth characteristics.”
According to Perola, this was highly valuable information that was obtained using artificial intelligence. The computer was fed registry data and taught to identify certain features in the dataset.
“This could not be done with anything other than CSC’s sensitive data services and supercomputing environment.”
Read more:

Sequencing of the Arabica coffee genome was published in April 2024. It was one of the latest major crop plants for which genome sequencing had not yet been published. According to Jarkko Salojärvi, who led the research, genes can now be found that will improve coffee’s yield and resistance to disease.
An international research consortium including researchers from the University of Helsinki and Nanyang Technological University in Singapore has compiled the genome of three coffee species from DNA sequences of coffee plant leaf cells. The genome of Arabica, or Coffea arabica, was assembled in Singapore and Helsinki, and those of Robusta (C. canephora) and C. eugenioides were assemled at Cornell University in the United States.
“The aim is to find traits that improve yield and quality. Cultivated Arabica is less genetically diverse, and therefore more susceptible to disease,” says Salojärvi, an assistant professor at Nanyang Technological University.
The economic importance of coffee is huge: it is grown in 70 countries and provides a livelihood for more than 100 million people. This makes coffee one of the world’s most important commercial products. Coffee breeding has its risks, however.
“In general, the genetic diversity of crops decreases with breeding. In cultivated coffee varieties, the genes that provide disease resistance are therefore not very diverse. That is why Arabica is vulnerable to pathogens.”
Altogether 60 per cent of the world’s coffee is produced from Coffea arabica. Besides Arabica, another species used for commercial production is Coffea canephora, or Robusta, which contains more caffeine than Arabica and is more bitter. It is used in instant coffee in particular. Vietnam is the world’s largest producer of Robusta. The rare Coffea eugenioides, or Eugenioides coffee, is sweet due to its low caffeine content. It has a lower yield than Arabica and Robusta.

Salojärvi specialises in plant genomics, and has been involved in research on the genomes of avocado, birch, lychee and Darrow’s blueberry. Salojärvi holds positions both at Nanyang Technological University and the University of Helsinki. The very broad and international research team makes extensive use of computational resources and databases in both countries.
Whole genome sequencing allows both common and rare mutations to be detected across the genome. Due to the complex structure, the genome of Arabica coffee was not sequenced until 2024.
Arabica is a hybrid of Coffea eugenioides and Robusta. Since both parental species are diploid – that is, they have two sets of homologous chromosomes – Arabica is a tetraploid, with four sets. Tetraploid plants often grow faster and larger than diploids. Their genome structure is often very complex, making assembly of the genome more difficult. The assembly of the Arabica genome was further complicated by the fact that the two subgenomes, Robusta and C. eugenioides, are very similar due to their close evolutionary history. Speciation – the process of biological species formation – occurred only about 4.5 to 7.2 million years ago.
In contrast, the Arabica hybrid, native to Ethiopia, is relatively young, around 350, 000 years old. Arabica has undergone many genetic bottlenecks, whereby a significant proportion of the population is prevented from reproducing and the population is substantially reduced. Due to these bottlenecks and the recent establishment of the species the genetic diversity of Arabica is not very high. The cultivated variants have even less genetic variation than the wild Arabica. This is due to a human-made bottleneck: most of the Arabica grown in the world is descended from just two coffee plants that lived ca. 300 years ago.
Arabica seeds were smuggled out of Yemen in the early 1600s, and since then Arabica began to be cultivated in Southeast Asia and later in the Caribbean. This variety of Arabica is called Typica, and its cultivation was managed by the Dutch. In the 1700s, the French started growing Arabica on the island of Réunion in the Indian Ocean. This variant is called Bourbon, in homage to the old name of Réunion. Roughly 95% of the current Arabica cultivars are descended from either Typica or Bourbon lineages.
Climate change is already affecting coffee yields. Drought has reduced yields in Brazil, Colombia and other countries. Arabica is grown at an altitude of over 1,500 metres in the tropics. As the climate warms, it will have to be grown at even higher altitudes, which will result in a reduction in the suitable area under cultivation.
Global warming will also increase the incidence of diseases. Hemileia vastatrix, or coffee leaf rust, causes the coffee bush to shed its leaves. The disease does not survive in temperatures below 10 ºC, so warmer nights at high altitudes contribute to its spread.
According to Salojärvi, however, the resilience of coffee can be improved through genome-based breeding.
“It’s possible to develop predictive models by sequencing the parents of a population and learning to predict the phenotypes of the offspring based on their parents’ genomes. This will help in identifying markers that can be used to select the next generation of individuals that are likely to produce better yields or be more resistant to pathogens. This is particularly important for coffee, for which the area under cultivation could be halved in the next 30 years or so due to climate change.”
Sequencing can be used to search the genomes of coffee species for genomic regions that are resistant to heat and disease. Robusta is known to be more resistant to hot weather than Arabica. It is also resistant to diseases such as coffee leaf rust. A hybrid between Robusta and Typica Arabica (Hibrido de Timor), found on the island of Timor in the 1930s, is particularly hardy.
According to Salojärvi, the regions found in its genome may enable genome-based breeding of Arabica.
“The genes from the Timor hybrid are only candidate genes, though. The next step is to examine whether the link is actually causal. It will probably take between five to ten years to test it, at which point that information can be used for breeding.”
The research will focus on the function of these candidate genes during onset of the disease.
“For example, it could be that those genes do get activated with the onset of coffee rust, but they may be such a late response that they are of no further use in preventing it,” Salojärvi points out.
“The next step would be to silence those genes and see if the resistance goes away. Or they could be transferred to a variety that is susceptible to coffee rust and see if it can build up resistance. Neither version can be used for coffee production because they would be transgenic individuals, but they can be used to ensure that the genes are the correct ones. Breeding can then focus on offspring with that resistance region.”
Establishing a chromosome-level assembly required also the determination of the three-dimensional structure of the chromosomes. The computing resources of the CSC– IT Center for Science, the Finnish ELIXIR Node, were used for this task. In the process the contiguous coffee genome sequences assembled in earlier steps were combined into chromosome-length scaffolds.
“This process means that a chromosome is made up of fully sequenced fragments and empty spaces in between. Among other things, the structural analysis will reveal the connection with genes and the regions of the genome that regulate them.”

After studying the coffee genome, Salojärvi will next study the genomes of rainforest plants. The Bukit Timah Nature Reserve in Singapore, which spans 163 hectares, is home to over 800 flowering plant species. In a project underway at Nanyang Technological University in his group is studying the rainforest biodiversity through sequencing the genomes of all the flowering plant species in the area. The focus now is on charting the gene content in a rainforest and to look for novel biosynthetic pathways, where plants use enzymes to make complex compounds from simple compounds.
“It’s particularly interesting to study what variations to the main biosynthetic pathways the different plant species have.”
Plant metabolites are important targets for research, for example to find new pharmaceuticals. According to Salojärvi, machine learning will revolutionise the study of drugs and metabolites.
“For instance, Alphafold 3, Google’s new AI software, can predict protein structures and different modifications to metabolites from the plant genome. Once the genome has been sequenced, this research will move forward rapidly thanks to artificial intelligence.”
All three coffee genomes have been shared via EBI/NCBI. Additionally, these annotations have been made accessible via ORCAE, a database with tools to further work on the gene structures, and containing annotations of diverse eukaryote genomes. It is operated by the Belgian ELIXIR Node.
The Belgian Node supports Plant and Biodiversity research. It provides resources for genomics and management of phenotyping data. Bioinformatics groups from VIB/UGent, who also participated in the coffee research, have developed tools for curation of annotations of genomes (ORCAE) and comparative genomics (PLAZA). ORCAE is an online genome curation and browsing portal for eukaryotic species, whereas PLAZA is an access point for plant comparative genomics, centralizing genomic data.
“Any publicly funded project has to release generated raw data to the scientific communities. Each system offers interfaces and procedures to help uploading the raw read data with their associated metadata”, says Principal Scientist Stephane Rombauts from VIB-UGent Center for Plant Systems Biology.
“We are developing better, newer interfaces to make the whole submission process easier”
Belgium ELIXIR node has also been developing tools to facilitate submission to ENA. The European Nucleotide Archive (ENA) is a fully open repository for storing raw sequencing data, assemblies, and annotation data.
The ENA Data Submission Toolbox simplifies the submission of sequence data by offering a single-step submission process, a graphical user interface, tabular-formatted metadata and client-side validation.
“The interfaces only offer a way to upload the data, while if serving as a backup, it would be an incentive to upload data sooner”, says Rombauts.
“Once uploaded the data has to be validated by experts before added to the system and it obtains its unique accession number only at the end. The process can sometimes be slow as sequencing gets cheaper and easier, while human experts still need to validate the ever increasing uploads.”
“Also, genomic data increasingly comes as long reads, or as raw, more rich, data in much larger quantities than before, making these interfaces sometimes inappropriate for the very latest types of data, or for the newest applications.”
Ari Turunen
22.7.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Mapping the coffee genome to improve disease resistance. https://doi.org/10.5281/zenodo.13691962
More information:
The genome and population genomics of allopolyploid Coffea arabica reveal the diversification history of modern coffee cultivars.Nature Genetics, 56, 721-731 (2024).
https://doi.org/10.1038/s41588-024-01695-w
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The pandemic may be over, but the COVID-19 virus, or coronavirus, has certainly not disappeared, and could mutate into a dangerous form again. Research professor Markus Perola of the Finnish Institute for Health and Welfare (THL) and his team are using registry and genomic data to determine which factors contribute to the risk of some people developing severe coronavirus requiring hospitalisation. The research requires a large amount of computing and processing of sensitive data.
The emergence of the COVID-19 virus in late 2019 caused a pandemic that shocked the world. The disease was life-threatening for elderly people. By March 2021 – in a period of about 15 months – around 2.5 million people worldwide had died from the disease. The global crisis affected both the global economy and the health sector.
“Pandemics come about because of population growth and because we are living closer to farm animals. At the same time, biodiversity is declining and we have a narrower diet. The globe is essentially a petri dish in which pandemics are bred,” Perola says.
Because new forms of the COVID-19 virus can emerge, it’s crucial to understand how the virus works and how to fight it. The COVID-19 Host Genetics Initiative project, among others, brought together researchers from around the world to gather information on the characteristics of coronavirus infection.
The aim is to identify individuals with a high risk of developing the serious form of the disease. As a result of the project, more than 50 genomic regions were identified that may contain genes that predispose a person to COVID-19. Some of these also predispose a person to a particularly severe form of the disease.
“This information will be used in THL’s own study to find out why some people who contract coronavirus end up in hospital. One reason may be found in genes.”
The study led by Perola involves collecting data on more than 3,000 people who were hospitalised with the virus, or, in milder cases, sought coronavirus testing. The survey uses registry data. Sample collection is carried out in collaboration with biobanks. Blood samples are tested for other co-infectious diseases, the severity of the inflammation, and other values that indicate the biological balance of the body.
“There is always talk of different risk groups, but we forget that a large proportion of people at risk of the coronavirus either do not end up in intensive care or do not die of the disease. For example, the mortality rate for people over the age of 80 is around 10 per cent, but several times that number do not die. So what is the difference between these groups? And why do some very overweight people end up in intensive care with the coronavirus, but not others? Our aim is to find the groups at risk to benefit from vaccination the most.”
According to Perola, combining genetics and registry data will shed further light on these questions. Perola’s team has amassed some exceptionally interesting research results from analysing large amounts of data. Working with Tero Hiekkalinna and Joseph Terwilliger, Perola ran a simulation to test the use of data from a million human genomes. The data also included clinical phenotype data. The anonymised data was analysed by a supercomputer at the Finnish ELIXIR Node, CSC – IT Center for Science, including not only genomic data but also information on health, family relationships, age and gender. This test yielded valuable new insights into how big data could be used in public health in the future.
Why is understanding national health variations important for the country’s healthcare?
“If we don’t know what’s specific to Finns, no one else will research this. A good example is the diseases of the Finnish disease heritage, of which there are about forty rare diseases that are concentrated in Finland. There is intensive international collaboration in gene research to identify these genes and understand how they work. But it is researchers in Finland who are making it a clinical reality.”
Finland is a genetic isolate – a population with minimal genetic mixing due to geographical isolation or similar mechanisms – because it has historically developed in a somewhat isolated way from other European countries. The Finnish population has its own form of inheritance, which – from a research point of view – is easy to approach differently from others. Some of the biological characteristics found in Finland are not found anywhere else in the world. Several hundred disease variants that are not seen in other populations have been found here – in other words, they are uniquely Finnish disease variants.
According to Perola, the Finnish population is, in a way, the largest genetic isolate in the world.
“We have the statistical power to find more of these variants compared to other isolated populations, such as the population of Iceland. Rare gene variants offer new insights into disease biology that are not available from other populations. They can open up whole new ways of understanding diseases. Is there already a cure for a given disease, or do we need to develop one?”
According to Perola, Finland is the country that others look to when it comes to the use of registry data. This was the case, for instance, with the creation of the European Health Data Space (EHDS). Registry data has been collected in Finland for decades. For example, a cancer registry was set up as early as the 1950s.
“We have extensive data in registries, for example the Kanta Services, which stores people’s health data and prescriptions. Not many countries have a similar situation – for example, there are not many countries where all laboratory data is available as it is in Finland today. We have access to data from the entire population, regardless of the different data systems or governance structures.”
Perola gives the example of one of his previous studies, which involved using registry data to find out what distinguished people in Finland who took the first coronavirus vaccine from those who refused it.
“We wanted to determine the factors that describe the proportion of the Finnish population – almost 20 per cent – who did not take the first vaccine. We looked at family relationships and socioeconomic variables: whether the person was in paid employment, their area of residence and native language. The data allowed us to scientifically justify that the message about vaccines did not reach migrants in time, and that there were people who did not have the resources to find out about vaccination themselves.”
Another thing that was studied was respiratory syncytial virus (RSV) infection in children under the age of 1. Respiratory syncytial virus (RSV) is a ribonucleic acid (RNA) virus that causes millions of respiratory infections worldwide every year. It is a major cause of infections in young children.
“The registry data was used to follow families whose child had been hospitalised after contracting RSV. The study used data related to socioeconomic status, use of intoxicants by the child’s parents, and the child’s birth characteristics.”
According to Perola, this was highly valuable information that was obtained using artificial intelligence. The computer was fed registry data and taught to identify certain features in the dataset.
“This could not be done with anything other than CSC’s sensitive data services and supercomputing environment.”
Perola uses genetic and registry data in his research.
“Research infrastructure is important. Research needs organisations such as the CSC to enable analysis. It doesn’t matter whether the scientist is an astronomer or a geneticist – they both use the same infrastructure. It’s always difficult to get money for infrastructure when foundations don’t fund it but instead just assume that the state will pay. But the state says, ‘Get outside finding,’ so we’re in a catch-22 situation. Supporting research infrastructure is essential to ensuring research excellence in science in Finland.”
Ari Turunen
25.6.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Why do some get the severe form of COVID-19?. https://doi.org/10.5281/zenodo.14810467
More information:
CSC SD-connect
https://docs.csc.fi/data/sensitive-data/sd_connect/
Finnish Institute for Health and Welfare THL
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Associate Professor Andrea Ganna from the Institute for Molecular Medicine Finland (FIMM) at the University of Helsinki uses large datasets to identify the demographic and genetic traits that underlie common and complex diseases. AI can make a risk calculation for each individual by modelling data from longitudinal tracking of diseases and medications along with genetic, family and demographic data. Once the data has been collected from different registers, the individual data is encrypted and stored in the sensitive data services of the Finnish ELIXIR node of the CSC - IT Center for Science.
The dataset contains data from 7.2 million individuals, i.e. the entire population of Finland and many relatives who have already died. It contains a lot of different, wide-ranging information, including health information, information on family relationships, socio-economic information, and laboratory results and prescriptions.
Read more here

By combining genomic data with data in national health registries, an artificial intelligence model can be developed that can be asked questions about potential future disease treatment. Such statistical and machine learning models are able to predict the occurrence of a disease.
Associate Professor Andrea Ganna from the Institute for Molecular Medicine Finland (FIMM) at the University of Helsinki is interested in combining genetic and statistical data.
“Healthcare can benefit from machine learning, which is constantly learning from the huge amount of data available to it. Questions can be posed to AI about potential hospital treatments in the future. AI can tell what a person’s life expectancy is, or how much prescription drugs will cost next year with certain life choices.”
Ganna uses large datasets to identify the demographic and genetic traits that underlie common and complex diseases. AI can make a risk calculation for each individual by modelling data from longitudinal tracking of diseases and medications along with genetic, family and demographic data.
In particular, Ganna uses FinRegistry data in his research. FinRegistry is a joint research project of the Finnish Institute for Health and Welfare (THL) and FIMM, led by Research Professor Markus Perola from THL. It is one of the world’s largest studies that makes secondary use of registry data.
“The dataset contains data from 7.2 million individuals, i.e. the entire population of Finland and many relatives who have already died. It contains a lot of different, wide-ranging information, including health information, information on family relationships, socio-economic information, and laboratory results and prescriptions. This is an enormous data set.”
The database includes data from 19 national registers, such as the Finnish Cancer Registry, the Drug Purchase Register and Kanta. Kanta is a register that brings together customer and patient data from healthcare and pharmacies. More than one billion pharmaceutical purchases alone have been registered in the collection so far. These are data points, with each individual fact being one data point. In total, there are more than 6.5 billion data points in the dataset.
“I consider the project to be unique. The data is rich and varied,” Ganna says.
“Combining health information with social and economic information is extremely important to me. These are often considered to be separate from each other, but combining the data is vital for health. We need to consider socio-economic information to understand how “fair” AI models are. We don’t want AI model to work worst in the most fragile sectors of our population. ”
Once the data has been collected from different registers, the individual data is encrypted and stored in the sensitive data services of the Finnish ELIXIR node of the CSC - IT Center for Science. Ganna and his research team analyse the data in this secure environment.
“We have worked with the CSC to make services more useful for researchers. We started with simple analyses and moved towards more complex models.”
There is a colossal amount of sensitive data in Ganna’s research.
“We are creating a data matrix for AI and machine learning models, but we are also very aware of the sensitive nature of the data. We cannot re-identify individuals and we use very advanced security measures to avoid unauthorized access. ”
This information may be used for various purposes.
“We are gaining a better understanding of the different clusters of disease, and are able to make better predictions. We can even create a digital clock that describes ageing. It uses data from the whole population to give each person in Finland a kind of digital age, based on an indicative trajectory derived from health data.”
The plan is to integrate the registry data by Ganna and his research team into the genomic data in the biobanks. This is an ambitious project, aiming to identify emerging diseases in individuals that could be prevented from developing. In the future, the data could be used to identify at-risk individuals who could benefit from preventive drug treatment.
There is already enough data to make this possible, according to Ganna. Ganna cites the FinnGen research project, which has already produced genome data on half a million people in Finland deposited in the biobanks. The project involves investigating the genetic background of various diseases in the Finnish population. The next step is to determine how genes influence the progression of diseases.
“It would be possible to contact people at risk, as their information is in the biobanks. Of course, this assumes that the people in the biobanks have given their consent to be contacted.”
In Ganna’s view, the CSC’s sensitive data services should be further developed to support machine learning models in particular. So far, AI models have only been tested in research. This is because, under current Finnish legislation, it is not possible to automatically use registry data to re-contact people at risk.
“We can make these beautiful models, but we can’t warn people at risk,” Ganna says. However, he adds that if the models are simplified enough, they can be used in clinical care.
One example he cites is the respiratory syncytial virus (RSV) that Pekka Vartiainen from FIMM and Markus Perola from THL studied in the FinRegistry project. RSV is the commonest virus causing respiratory infections in young children worldwide. The researchers created a simplified model that can be used in the clinical management of RSV. In Finland, doctors could now use registry data to identify who is at risk of contracting the virus and who could be treated in time.
Ganna believes that in the future, healthcare will benefits from AI models that understands health data.
“AI will support clinical decision making, by helping doctors to better summarize health trajectories of their patients. The future is bright.”
Ari Turunen
30.5.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). An AI model that understands health data warns of future diseases. https://doi.org/10.5281/zenodo.13691998
More information:
Finnish Institute for Life Science (FIMM)
FIMM is part of HiLIFE Helsinki Institute of Life Science -research center.
https://www.helsinki.fi/en/hilife-helsinki-institute-life-science/units/fimm
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Finnish Use Cases 2 publications present a range of Finnish biomedical research that has benefited from the resources of the European biomedical infrastructure ELIXIR. Almost 200 research organisations have joined the ELIXIR infrastructure, contributing to the work of more than half a million researchers across Europe. Its Finnish operations are run by the CSC IT – Center for Science.
The publication presents a variety of research that has made use of new bioinformatics methods. The subjects range from drug development, disease mechanisms, cell modelling, microbiomes, utilisation of AI models in diagnostics and personalised drug treatments.
Biomedical research has become data- and computation-intensive. Data analysis calls for more and more sophisticated software and combinations of it. Researchers also need services and resources for data storage and description for further processing.
For example, translational medicine uses basic research in clinical trials, patient samples and disease models to identify disease mechanisms and drug effect targets. The starting point is interdisciplinarity, which helps patients as well as serving research well. The idea behind combining several different data sources is to make more information available. The integration is very much computational, and requires CSC – IT Center for Science’s resources and infrastructures like ELIXIR.
CSC has created services in the infrastructure that are particularly suited for sensitive data.
“The management of research data of humans must be carried out professionally, and this is one of the key objectives of ELIXIR Finland. A combination of high-quality management of sensitive data and high-performance computing creates unprecedented opportunities for Finnish scientists to create models that predict human health, ultimately benefiting all of us – we just need to get the different parts of the ecosystem to work together,” says Tommi Nyrönen, head of the Finnish ELIXIR node.
The construction of the bioscience infrastructure ELIXIR began in Europe in 2013. ELIXIR enables biomedical researchers access to biomedical databases and the computational resources and software to process it, and training in using the system. By April 2024, CSC had 2,386 biomedical and healthcare researchers as its customers.
Download the report here

The CSC – IT Center for Science co-led the European Beyond One Million Genomes (B1MG) project, which focused on creating a secure cross-border federated infrastructure for the use of genomic data. The project is now being followed by the European Genomic Data Infrastructure (GDI) to allow researchers to access European genomic and clinical data.
The aim is to improve diagnostics and pharmacogenomics – in other words, to improve the impact of individual differences in hereditary factors on drug response. Another aim is to support the secondary use of these data for research. Valuable data will been collected from patients with cancers and rare and polygenic diseases. Data has also been collected on disease-causing pathogens as well as infectious diseases such as COVID-19.
This data can provide the basis for personalised drug treatments using multi-gene risk assessment. The risk is calculated using a personal polygenic risk score (PRS), which covers millions of genetic variations.
The three-year B1MG project ended in October 2023. The Finnish ELIXIR Node CSC led the technical infrastructure work in the project. The Genomic Data Infrastructure project, launched in 2022, is coordinated by ELIXIR, a cross-European life-science infrastructure for biological information. The aim of the GDI is to create the final infrastructure to provide access to genomic and clinical data collected from Europeans. GDI is a consortium of partners from 20 European countries. The B1MG project made recommendations to GDI regarding data and metadata management.
”B1MG was a coordination and support action grant, that was tasked with determining the roadmap and best practices for the deployment of the required infrastructure to support the 1+ Million Genomes ambition. As work package co-lead for the technical infrastructure CSC was able to drive forward the decisions on the roadmap ensuring that these aligned with existing and future CSC requirements, such as Sensitive Data Services,” says senior coordinator Dr Dylan Spalding from CSC.
“CSC has supported deploying federated sensitive data nodes. In this role CSC has used its experience in federated sensitive data services.”
Spalding worked for B1MG-project as a work package co-leader, which focused on the personalised medicine.
”The real benefit of B1MG is how it has set the direction for the GDI project, which will deploy a federated infrastructure across Europe to support cross-border access to over 1 million genomes. This has the potential to help democratise research, and drive personalised medicine across the EU. “
CSC as co-leads of the infrastructure pillar has a leading role in this work. Also the Life Science AAI (Authentication and Authorization Infrastructure) & REMS (Resource Entitlement Management System) are applications already in use to support access management to data. According to Spalding this should align well with the existing Federated EGA node and Sensitive Data Services. The Federated EGA (European Genome-phenome Archive) is a distributed solution for sharing and exchange of human -omics data across national borders.
”GDI is very important for rare disease and personalised medicine, but also cancer, infectious disease, and common and complex diseases. However, the infrastructure isn’t specialised for any particular disease, but should support research into all disease types. The development is driven by the 1+ Million Genomes use cases, as well as Genome of Europe which is aiming to build reference cohorts of 500,000 citizens across Europe.”
According to Spalding B1MG demonstrated a proof of concept version of the Starter Kit for both rare disease and cancer use cases. The Starter Kit is a set of software applications and components co-developed by the 20 GDI nodes,
The Starter Kit has been created for the basis of GDI. There are five functionalities which were defined in B1MG that need to be supported – data reception, data discovery, data access management, storage and interfaces, and processing.
The Starter Kit includes more than 2,500 synthetic genomics and phenotypic data on cancer and rare diseases. It is a first step towards a production infrastructure.
”Starter kit contains all necessary functionality to deploy a demonstration system that allows the discovery, access, and analysis of sensitive genomic and phenotypic data. A set of synthetic data is included that can demonstrate these functionalities without risk of leaking real genomic or phenotypic data.”
An evolved version of Starter Kit will be integrated to the GDI portal.
Spalding believes that the huge amount of data in the GDI project will enable a better-personalised treatment.
”GDI has the potential to support new machine learning and AI methods, speeding up the transition to personalised medicine across Europe.”
Professor Arto Mannermaa’s group from the University of Eastern Finland is developing learning algorithms based on genomic and clinical data to identify and predict risk factors for breast cancer. Genomic and clinical data are combined to form an artificial intelligence model that not only helps to determine the risk of illness, but also in drawing up individual treatment plans.
Mannermaa’s group creates AI models from image data. What other data should be combined with image data to improve healthcare?
“We have now incorporated genomic data into our imaging data. The more data modalities we can combine, the better we can identify factors related to successful cancer treatment, and the more likely we are to identify factors influencing disease risk.”
Factors influencing disease risk include data on treatment response or other clinically relevant information related to treatment.
“The more data there is, the more demanding the computing environment becomes. Ancillary data can be obtained from various sources, such as electronic health record systems through biobanks.”
Ari Turunen
29.4.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). An infrastructure for genomic data. https://doi.org/10.5281/zenodo.13691595
More information:
Genomic Data Infrastructure
https://gdi.onemilliongenomes.eu
Beyond One Million Genomes
https://b1mg-project.eu/1mg/genome-europe
University of Eastern Finland
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The European Genomic Data Infrastructure (GDI) allows researchers to access European genomic and clinical data. The aim is to improve diagnostics and pharmacogenomics – in other words, to improve the impact of individual differences in hereditary factors on drug response. Another aim is to support the secondary use of these data for research. Valuable data will been collected from patients with cancers and rare and polygenic diseases. Data has also been collected on disease-causing pathogens as well as infectious diseases such as COVID-19.
CSC as co-leads of the infrastructure pillar has a leading role in this work. Also the Life Science AAI (Authentication and Authorization Infrastructure) & REMS (Resource Entitlement Management System) are applications already in use to support access management to data. The Starter Kit has been created for the basis of GDI. The Starter Kit includes more than 2,500 synthetic genomics and phenotypic data on cancer and rare diseases. It is a first step towards a production infrastructure.
Read more here:
The aim of the BeYond-COVID project (By-COVID) is to make COVID-19 data collected in different European countries available to researchers, hospitals and public administrations.
A portal with COVID-19 data is available on the By-COVID project website.
The project is coordinated by the ELIXIR infrastructure, whose member organisation EMBL-EBI has compiled the main coronavirus datasets on the portal. Researchers can use the portal to analyse the COVID-19 reference data. It contains more than 8 million COVID sequences.
The Research Council of Finland funded the Finnish ELIXIR Node’s CSC experiment, where data from the COVID 19-portal was analysed and tested on the Finnish LUMI supercomputer. The work supports the By-COVID project: an important role of the CSC is to promote the use of supercomputers in data-intensive computing.
According to Tommi Nyrönen, Director of the ELIXIR Finland node, the project overcame many of the technical challenges of data management.
“Computational work at the EMBL-EBI European Bioinformatics Institute enabled the analysis of the COVID-19 viral data, and this work was done in collaboration with experts from the CSC and EMBL-EBI. With European supercomputing, we can now transfer hundreds of thousands of virus data points daily between computing centres with the help of European research networks.”
As a result, the capacity of a supercomputer will be needed in the future to analyse all the data.
“This is essential for a rapid response in the event of a new pandemic, and also to update the data available on the COVID-19 portal.”
Read more here:

The aim of the BeYond-COVID project (By-COVID) is to make COVID-19 data collected in different European countries available to researchers, hospitals and public administrations. Identifying data, combining it from different sources and integrating it for analysis is a major undertaking, one that has been taken up by no fewer than 53 organisations from 19 countries. The Finnish organisations involved are Tampere University and the Finnish Institute for Health and Welfare (THL). Data collected in Finland at THL has been processed in the CSC – IT Center for Science.
According to THL research professor Markus Perola, the By-COVID project is simply preparing for the next pandemic by analysing the COVID-19 data.
“We are now piloting how this kind of collaboration can be done when the next pandemic comes. This seems to be necessary.”
There is a genuine need for data harmonisation between European countries, Perola says.
“For instance, different countries may have very different ideas about what is considered essential in COVID infection chains.”
Perola uses CSC’s computing and sensitive data storage and analysis services for almost all his research. In addition to genetic data, he makes extensive use of register data. In the By-COVID project, his research team has used Finnish communicable disease registers and mortality data from Statistics Finland. The data will be used for joint analyses in the By-COVID project. THL’s raw data is available in CSC’s sensitive data services, but it is kept within Finland. The By-COVID project is also collecting information about the virus itself. This is open research data.
“THL is participating in one work package which involves federated analysis of register data from different countries. The project will extract specific issues from different registers and combine them to move towards a common analysis across Europe.”
The register data collected from Finland includes all Finnish residents who have a personal identity code.
According to Perola, it is necessary to collect and analyse this kind of data. In his view, it would be unethical not to use the important data that is collected on European citizens.
“Why collect data if you’re not going to use it? Statistics are important, but they’re enough to translate the information into clinical work or social policy-making. This requires peer-reviewed scientific research, and that is what By-COVID provides.”
The project will end in autumn 2024.
A portal with COVID-19 data is available on the By-COVID project website. The project is coordinated by the ELIXIR infrastructure, whose member organisation EMBL-EBI has compiled the main coronavirus datasets on the portal. Researchers can use the portal to analyse the COVID-19 reference data. It contains more than 8 million COVID sequences.
The Research council of Finland funded the Finnish ELIXIR Node’s CSC experiment, where data from the portal was analysed and tested on the Finnish LUMI supercomputer. The work supports the By-COVID project: an important role of the CSC is to promote the use of supercomputers in data-intensive computing.
According to Tommi Nyrönen, Director of the ELIXIR Finland node, the project overcame many of the technical challenges of data management.
“Computational work at the EMBL-EBI European Bioinformatics Institute enabled the analysis of the COVID-19 viral data, and this work was done in collaboration with experts from the CSC and EMBL-EBI. With European supercomputing, we can now transfer hundreds of thousands of virus data points daily between computing centres with the help of European research networks.”
As a result, the capacity of a supercomputer will be needed in the future to analyse all the data.
“This is essential for a rapid response in the event of a new pandemic, and also to update the data available on the COVID-19 portal.”
Ari Turunen
1.4.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). European research community preparing for next pandemic. https://doi.org/10.5281/zenodo.13691578
FIRI
Article was supported by the Research council of Finland grant No: 345591 for ELIXIR European Life-Sciences Infrastructure for Biological Information (FIRI 2021)
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
HPV vaccines provide protection against cervical cancer in particular, but are also effective against HPV-related cancers in other parts of the genital area and the mouth and throat. According to Ville Pimenoff, associate professor in evolutionary medicine at the University of Oulu, the immune protection including herd immunity provided by the HPV vaccine is an effective way to protect the population from HPV-related cancers, especially when both girls and boys are vaccinated. In addition, adequate vaccination of the whole population changes the ecological dynamics of the remaining papillomaviruses.
“This leads to the conclusion that in the near future, cervical cancer screening for oncogenic HPV infections should be eased or completely stopped for those who have been vaccinated.”
Pimenoff’s study of genetic variation in viruses and other microbes in general requires a large amount of CSC’s computation and data processing.
In studying cancer-causing papillomaviruses at population scale, Pimenoff used a large cervical HPV infections cohort data collected from 33 Finnish cities and towns, where a total of 22,000 young individuals were monitored for 16 years since most of them had received the HPV vaccine. The research dataset is the world’s largest community randomized vaccination cohort, providing an excellent setup to examine the evolutionary dynamics of cancer-causing papillomaviruses in the population that has been HPV vaccinated using different vaccination strategies and compared to the non-vaccinated fraction of the population.
“From this cohort data, I used computer-assisted methods to simulate HPV infections prevalence dataset among half a million young Finnish women. I used the computing power of CSC and the sensitive data virtual cloud for the original data derived simulations, and for editing and reviewing the resulting synthetic and original data.”
Read more here:

Ville Pimenoff, associate professor in evolutionary medicine at the University of Oulu, makes extensive use of the supercomputing environment and sensitive data services of the CSC – IT Center for Science to study the population genetics of viruses and other biological exposures to humans. Pimenoff specialises in environmental exposures and the human exposome based on biobank data. The concept of an exposome refers to the totality of exposures an individual is subjected to throughout their lifetime in their living environment. The aim of exposome research is to understand the impact of different exposome components in human health.
One of Pimenoff’s recent studies showed that human papillomavirus (HPV) vaccination provides effective immune protection against the most cancer-causing papillomaviruses, while significantly altering the ecological dynamics of the remaining less oncogenic papillomaviruses.
Pimenoff conducted his doctoral dissertation on the genetic history of Finns and Finno-Ugric peoples. Currently he has ten research projects underway, focusing on the short and long-term evolution of microbes that infect humans.
“My work stems from computational metagenomics, using DNA sequence data to analyse genomes of different organisms and microbes in a biological sample. This means that research datasets often amount to tens, sometimes even hundreds of terabytes of sequence data.”
Pimenoff’s study of genetic variation in viruses and other microbes in general requires a large amount of computation and data processing.
Researchers have identified over 200 types of human papillomaviruses (HPVs) that can infect humans. Some dozen of these are among the most common sexually transmitted viruses, and can cause cervical cancer. Basically, most of us will be infected with oncogenic HPV at some point in our lives, usually at a young age. If the oncogenic HPV infection becomes persistent, it can progress to a precancerous lesion and eventually may develop into cancer. Cervical cancer is almost always associated with a long-term HPV infection.
This has led to the development of the HPV vaccines. They all provide protection against cervical cancer in particular, but are also effective against HPV-related cancers in other parts of the genital area and the mouth and throat. According to Pimenoff, the evolutionary ecology study of HPVs indicated that the immune protection including herd immunity provided by the HPV vaccine is an effective way to protect the population from HPV-related cancers, especially when both girls and boys are vaccinated. In addition, adequate vaccination of the whole population changes the ecological dynamics of the remaining papillomaviruses.
“This leads to the conclusion that in the near future, cervical cancer screening for oncogenic HPV infections should be eased or completely stopped for those who have been vaccinated.”
Pimenoff’s research interests range from population genetics of humans to evolution of pathogens affecting human health.
“Clinician looks at the papillomavirus as a viral infection that may indicate the patient will develop cancer. An epidemiologist can investigate whether there are factors in the lifestyle of the population that may increase the risk of HPV infection leading to cancer.”
Pimenoff looks at infections from the pathogen population perspective.
“My point of view is that there is always a large number of viral populations circulating among humans. What I am trying to understand is the transmission dynamics by which these millions of viruses spread in the human population in time and space and affect our health, and how these dynamics may change if, for example, a reasonable number of individuals are vaccinated against some of these viruses. I explore these ecological and evolutionary dynamics at the short (ie. human lifetime) and long (ie. millenia) evolutionary timescales.”
In studying cancer-causing papillomaviruses at population scale, Pimenoff used a large cervical HPV infections cohort data collected from 33 Finnish cities and towns, where a total of 22,000 young individuals were monitored for 16 years since most of them had received the HPV vaccine. The research dataset is the world’s largest community randomized vaccination cohort, providing an excellent setup to examine the evolutionary dynamics of cancer-causing papillomaviruses in the population that has been HPV vaccinated using different vaccination strategies and compared to the non-vaccinated fraction of the population.
“From this cohort data, I used computer-assisted methods to simulate HPV infections prevalence dataset among half a million young Finnish women. I used the computing power of CSC and the sensitive data virtual cloud for the original data derived simulations, and for editing and reviewing the resulting synthetic and original data.”
Synthetic data can be analysed across all of CSC’s services. CSC has a dedicated service for sensitive data
In addition to projects related to viral genomics, Pimenoff makes extensive use of CSC’s services, especially for research on environmental contaminants.
“For a pilot project, we have recruited 100 women in Finland for an exposome study on environmental exposures. Of the women in the pilot, some are also pregnant. For four months the women carry a small air pump, which we call an exposometer. It has a filter that collects small particles such as bacteria, fungi and chemical particles in the air. The filters are changed every two weeks, providing a long sample series of environmental contaminants. DNA is isolated from the microbes in the filters and sequenced. This provides what is known as the metagenomic exposome data.”
The filter extracts also enable a mass spectrometry analysis to detect the different chemical compounds to which the women have been exposed during the monitoring period. This is important particularly because the participants were recruited from different locations, allowing us to assess the similarities and differences in environmental exposures in Finland during different seasons, both in urban and rural areas.
“The processing of such data can only be done in a secure supercomputing environment, and CSC’s sensitive data services enable this.”
In collaboration with CSC, Pimenoff has built a system where access of the sensitive data can be shared internationally as well. The data is used within the same project in collaboration with partners, and if needed it can be shared with specific research groups through the SD Connect service. Another service, SD Apply, is designed to grant licences for data.
“We can grant permission for access to anonymous data to be shared with our international collaborators, but the data sets are processed only within the CSC environment and cannot be physically transferred abroad. CSC’s tools for analysing sensitive genomic data are now better than before. CSC has made a good improvement,” Pimenoff says.
However, some obstacles remain in terms of data processing and the user-friendliness of CSC’s system. How is possible to handle large datasets as smoothly as possible and distribute access rights to different data sets? And how can access rights be managed agilely if a CSC customer is involved in several projects? According to Pimenoff, the user interface of sensitive data services should be developed so as to allow more flexible analysis of large data sets with research collaboration outside Finland.
Ari Turunen
8.3.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Evolutionary dynamics of viruses and other microbes affect human health. https://doi.org/10.5281/zenodo.13691466
More information:
FIRI
Article was supported by the Research council of Finland grant No: 345591 for ELIXIR European Life-Sciences Infrastructure for Biological Information (FIRI 2021)
University of Oulu
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The genomes of one million europeans were simulated with CSC’s LUMI supercomputer. The dataset was anonymized, so there is no unique and identifiable data. The goal is to create federated management that transcends national boundaries and gives access to national genomic archives. The simulation was done for the EU’s 1+MG Initiative. In 2018, the EU published the 1+Million Genomes (1+MG) Initiative, with an ambitious goal of collecting data covering the genomes of one million Europeans. The project was one of the biggest of its type in the world, with 27 countries participating.
“1+MG synthetic data project, from my viewpoint, the unique challenge was how we can create an effective simulation of a population which in its final generation includes a million people and corresponds in all its properties – in terms of genome, data formats and size – to genuine genomic data which, when simulated, is freely shareable without data security issues.,” says associate professor Tero Hiekkalinna of THL.
He was part of the simulation project group together with Markus Perola from THL and Joseph Terwilliger from the Univeristy of Columbia.
Read more here:

The EU-funded EOSC-ENTRUST project launches today, 1 March 2024, with the aim of enhancing European interoperability for sensitive data access and analysis. Led by ELIXIR and the EUDAT Collaborative Data Infrastructure (CDI), the project brings together partners from 15 European countries and will run for three years, with an EU contribution of €4.2M.
EOSC-ENTRUST will build a European network of Trusted Research Environments (TREs) for sensitive data and develop a common blueprint, or reference architecture, for federated data access and analysis. TREs provide secure platforms for handling sensitive datasets, enabling data privacy and legal compliance in research involving confidential information.
Europe’s landscape of TREs is currently fragmented, presenting challenges in access and management for researchers and providers. The variety of systems and procedures complicates research efforts, and there is a pressing need for a unified approach to managing federated access across differing technologies and governance frameworks. EOSC-ENTRUST brings together providers of operational TREs from 15 European countries with a shared goal to implement, validate and promote their capabilities through a common European framework using shared standards and common legal, operational and technical language.
The project will create a reference architecture for interoperability, based on the European Open Science Cloud (EOSC) Interoperability Framework, to address legal, organisational, technical and semantic interoperability aspects. It includes driver projects in genomics, clinical trials, social science, and public-private partnerships to test and refine this blueprint and facilitate secure data analysis through federated workflows. Targeted outreach activities will expand the provider network and develop policy papers and guidelines to create a long-term operational TRE framework within EOSC.EOSC-ENTRUST will work closely with the SIESTA and TITAN projects, funded in the same call, to provide trusted environments for sensitive data management in EOSC.
Peter Maccallum, ELIXIR’s Chief Technical Officer and Coordinator of EOSC-ENTRUST, said “We are delighted to launch the EOSC-ENTRUST project, and look forward to working with Europe’s TRE providers and wider sensitive data community to produce an interoperability blueprint to enable biomolecular and biomedical research involving sensitive data”.
Yann Le Franc, head of the EUDAT secretariat, said “EOSC-ENTRUST brings together two major data centric infrastructures, EUDAT and ELIXIR. With our experience and international networks, we will address the growing multidisciplinary demand for secure digital environments by orchestrating European-wide coordination and interfaces with other EU organisations and initiatives like EOSC and the European Data Spaces”.
Contact: grants@elixir-europe.org
Find out more

The Finnish Institute for Health and Welfare (THL), together with CSC, has simulated the genomes of one million Europeans. The data used for the simulation was publicly available full-genome sequences, but in the simulations they were formed into synthetic genomes, no longer representing real people. The simulation was done with CSC’s LUMI supercomputer. This is one of the most comprehensive human genome simulations in the world. The simulation was done for the EU’s 1+MG Initiative.
In 2018, the EU published the 1+Million Genomes (1+MG) Initiative, with an ambitious goal of collecting data covering the genomes of one million Europeans. The project was one of the biggest of its type in the world, with 27 countries participating. The secure use of European genomic data enables personalized healthcare and better diagnostics. This improves the treatment prognosis of especially cancers and neurological disorders.
The dataset is anonymized, so there is no unique and identifiable data. The goal is to create federated management that transcends national boundaries and gives access to national genomic archives.
“1+MG synthetic data project, from my viewpoint, the unique challenge was how we can create an effective simulation of a population which in its final generation includes a million people and corresponds in all its properties – in terms of genome, data formats and size – to genuine genomic data which, when simulated, is freely shareable without data security issues. In the end, we simulated a population of some 25 million people, of which just over a million was assigned synthetic genomes. This type of dataset enables a wide range of research, training and development project, such as 1+MG, without having to consider any ethical or legal issues or data security problems,” says associate professor Tero Hiekkalinna of THL.
At this point, a simulation of the synthetic material of a million people was conducted, with dozens of phenotypes. This means that environmental effects on the phenotypes of individuals are included in the dataset.
The simulation of a million genomes was funded in Finland by the Ministry of Social Affairs and Health as well as the Ministry of Education and Culture. Hiekkalinna says that there were huge challenges in creating and managing the material.
“Owing to the size of the material, we needed dozens of terabytes of disk space.”
The 1+MG Initiative was followed by B1MG (Beyond 1 Million Genomes) that ran from 2020 to January 2024. The B1MG project specified the guidelines and recommendations for the federated management of genomic data obtained from various European countries. CSC, the ELIXIR node in Finland, was one of the project’s managers and coordinators. The aim is to make the operation of biobanks compatible with cross-boundary data infrastructure. In the B1MG project, CSC was in charge of the technical infrastructure work.
The million-genome data simulated by THL and CSC will be made available in the Federated European Genome-phenome Archive (FEGA). The FEGA is designed for the storage and publishing of biomedical data for research purposes, but not to be available for the general public. The Finnish database is maintained by CSC. The FEGA is connected to the European Genome-phenome Archive (EGA). The EGA is one of the world’s most extensive data storage facilities.
The same simulated data will in future be available also for the Genomic Data Infrastructure (GDI) project. Started in 2022, the Genomic Data Infrastructure is coordinated by ELIXIR. The goal of the GDI is to create the final infrastructure, giving access to genomic data and clinical data collected of Europeans.
In future, Europeans can expect to have faster and more accurate diagnoses. Collected and analyzed genomic data enables better drug design and preventive drug treatment. All this will lead to better health and a higher life expectancy. This requires data preprocessing and harmonization, and secure, scalable and flexible technical solutions.
These three related projects make use of five use cases. These use cases are relevant for the construction of the final GDI infrastructure. The Genome of Europe will create a collection of reference data for health programs utilizing genomics in European countries: each country submits genomic data in proportion to its population. A data model will be developed from clinical cancer information and metadata obtained from genomics. A polygenic risk score (PRS) is created for the purpose of deciding on the patient’s treatment: the individual risk score will take into account millions of genetic variations. In rare diseases, the key aspect lies in the presence of gene variants in different populations, and discovering the link between gene mutation and illness. The sharing of COVID-19 data collected by each country in Europe will also be tested.
Ari Turunen
2.3.2024
Read article in PDF
Citation
Nyrönen, T., & Turunen, A. (2024). A million European genomes. https://doi.org/10.5281/zenodo.13691032
More information:
Hiekkalinna, Tero; Heikkinen, Vilho; Perola, Markus; Terwilliger, Joseph (2023):
Simulated European Genome-phenome Dataset of 1,000,000 Individuals for 1+Million Genomes Initiative.
1+MG Framework
https://framework.onemilliongenomes.eu
Beyond 1 Million Genomes (B1MG)
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Researchers can transfer their computing from Google’s Colab to CSC’s environment using a new application. A similar approach can be adapted if one wants to move from one supercomputing environment to another.
Application Specialist Laxmana Yetukuri at CSC – IT Centre for Science and Specialist Researcher Michael Courtney at Turku Bioscience Centre have customised GPU-empowered notebooks originally developed by the Centre’s team, that can apply deep learning models for biological image data in the CSC supercomputing environment. There is also an open-source toolkit, ImageJ/Fiji, for the deep learning models in microscopy and the usage of the toolkit is under exploration at CSC, the Finnish node of the ELIXIR infrastructure.
Researchers can transfer their computing from Google’s Colab to CSC’s environment using an in-house developed container wrapper. The container wrapper allows researchers to define a standardised environment in which they can run scientific software. The program’s code, along with its libraries and settings, is placed within the container. Once the software and its dependent packages are installed as part of the wrapper tool, other users can start using the application without any pre-installations. A similar approach can be adapted if one wants to move from one supercomputing environment to another.
“We make the work of researchers easier by providing easy-to-follow instructions for the installation of user’s custom notebooks. Once a project member installs an application in their project area, other researchers don’t need to install any software to use custom notebooks – they can start working immediately. Accessing CSC’s notebooks in supercomputing environment requires just a few clicks on the web interface (www.puhti.csc.fi),” says Yetukuri.
“Biological image analysis typically needs larger disk space for storing image data. CSC object storage ALLAS provides a good storage environment. The computing environment can be accessed only with a user account obtained from CSC,” Yetukuri says.
Read more here:

Researchers can transfer their computing from Google’s Colab to CSC’s environment using a new application. A similar approach can be adapted if one wants to move from one supercomputing environment to another.
Application Specialist Laxmana Yetukuri at CSC – IT Center for Science and Specialist Researcher Michael Courtney at Turku Bioscience Centre have customised GPU-empowered notebooks originally developed by the Centre’s team, that can apply deep learning models for biological image data in the CSC supercomputing environment. There is also an open-source toolkit, ImageJ/Fiji, for the deep learning models in microscopy and the usage of the toolkit is under exploration at CSC, the Finnish node of the ELIXIR infrastructure.
Turku Bioscience Centre has been using Google’s Colab Notebook cloud service to analyse and visualise data. Large scale data-intensive research activities, however, have their limitations in using Google services in free-of-cost models. Researchers often require significantly more storage and computing capacity for data processing than basic users. CSC’s supercomputing environment provides vastly superior storage and computing capacity free-of charge for academic users, and now it’s possible to make the switch from Google Colab environment to CSC environment. A personal laptop/PC can easily access CSC’s supercomputing environment through a web browser, thanks to recently developed user-friendly web interfaces to supercomputers at CSC.
Researchers can transfer their computing from Google’s Colab to CSC’s environment using an in-house developed container wrapper. The container wrapper allows researchers to define a standardised environment in which they can run scientific software. The program’s code, along with its libraries and settings, is placed within the container. Once the software and its dependent packages are installed as part of the wrapper tool, other users can start using the application without any pre-installations. A similar approach can be adapted if one wants to move from one supercomputing environment to another.
“We make the work of researchers easier by providing easy-to-follow instructions for the installation of user’s custom notebooks. Once a project member installs an application in their project area, other researchers don’t need to install any software to use custom notebooks – they can start working immediately. Accessing CSC’s notebooks in supercomputing environment requires just a few clicks on the web interface (www.puhti.csc.fi),” says Yetukuri.
“Biological image analysis typically needs larger disk space for storing image data. CSC object storage ALLAS provides a good storage environment. The computing environment can be accessed only with a user account obtained from CSC,” Yetukuri says.
Yetukuri and Courtney were able to utilise the existing CSC infrastructure by customising a Google Colab notebook to CSC’s computing environment to analyse microscopic image data. The custom notebooks were used to build machine learning models using microscopy image data. The notebooks were accessed through the Puhti web interface. The researchers are now exploring containerised deployment of imaging-related software, Fiji/ImageJ in CSC supercomputing environment to perform downstream analysis.
Biological imaging and image data analysis use algorithms to extract a significant amount of quantitative information from the images. This information can be used for pattern recognition and classification of image data, providing biologically significant insights. Using the system, Yetukuri and Courtney aim to develop and apply machine learning models for identifying SYNGAP1 gene variants causing brain disorders, and, in the future, for drug screening.
Courtney, Co-principal Investigator Li-Li Li and their colleagues at Turku Bioscience Centre are investigating disease-causing variants of SynGAP1 proteins. The SynGAP1 gene is located on the sixth chromosome and produces the SynGAP protein. The protein regulates synapses, the junctions through which nerve cells communicate with one another. A variant of the SynGAP1 gene causes the production of SynGAP protein to drop below a sufficient level. This leads to abnormal communication between nerve cells, leading in turn to various neurological disorders. In order to develop in a normal way, the brain requires two correct genes that encode the SynGAP1 protein. Mutations can lead to one of these not being expressed, resulting in developmental delays.

SynGAP1 encephalopathy is an early-onset intellectual disability. The developmental delay characteristic of the condition is typically observed during the first or second year of life. Additionally, about eight out of ten encephalopathy patients are diagnosed with epilepsy. The symptoms of epilepsy vary individually, and it can be difficult to treat. Behavioural disorders and autism occur in half of the patients.
Turku Bioscience Centre’s microscopy screening unit analyses normal SynGAP1 genes and point mutations that may impair protein function, sometimes to nearly non-existent levels. Point mutations only change one amino acid in the protein, but the consequences of this require further clarification.
The SynGAP1 protein is present only in nerve cells. Using high-throughput microscopes, it is possible to simultaneously examine 384 living nuron circuits over time. In the neurons SynGAP1 is labelled with a fluorescent protein tag that can be detected with frequency-tuned light. In each circuit normal SynGAP1 or a different pathogenic form can be studied. Aberrations in protein function can be observed based on the images.

The microscopes perform automatic image capture and can sample changes in circuits every 20 seconds. When studying different variants of the protein, it is possible to compare whether its function is normal, or whether is enhanced or has completely ceased, both of which could lead to pathology.
“We have been able to create experimental and analytic setups that investigate the functions of damaged SynGAP1 that deviate from the normal. This may potentially provide a pathway for drug screening in the future. There are also gene variants that are found in patients with the disease, but it’s unknown how or even if they are actually causing it,” Michael Courtney says.
“With our method, we can determine if these gene variants have a similar impaired function as known pathogenic variants.”
Once the sample preparation requirements were satisfied, a key step was to develop a deep learning model that automates the identification of SynGAP1 puncta (a bunch of dots) usually located at the synapses, the sites of communication between the neurons. Puncta are discrete regions of images where the fluorescent tag is visible.
”Once these are identified, their number and some 25 properties of each puncta can be extracted. Once demonstrated, this approach will be extremely valuable for future arrayed drug screens where each missense variant will be exposed to each of the up to 4,000 separate drugs in the screening library.”
According to Courtney only by testing drugs there is a hope to rapidly find a compound that already has known clinical safety and tolerability data. This information is crucial to clinicians and a potential short-cut to achieving a benefit for patients.
”As we are studying a rare disease with so many different variants of an essential protein, it is very difficult to carry out any kind of clinical trial to find effective drugs. Even generating animal models for this diversity of disease-causing variants is highly challenging. ”
Ari Turunen
20.2.2024
Read article in PDF
Citation
Nyrönen, T., & Turunen, A. (2023). Efficient transfer and analysis of biological image data through web interfaces. https://doi.org/10.5281/zenodo.13691023
More information:
Research is funded by the patient advocacy groups SynGAP Research Fund US and EU, and Leon and friends e.V.
FIRI
Article was supported by the Research council of Finland grant No: 345591 for ELIXIR European Life-Sciences Infrastructure for Biological Information (FIRI 2021)
A free and open-source notebook for Deep-Learning in microscopy at CSC. Possibility to run Google Colab notebooks at CSC HPC environment via the web interface
GitHub: https://github.com/yetulaxman/ZeroCostDL4Mic
Democratising deep learning for microscopy with ZeroCostDL4Mic
https://www.nature.com/articles/s41467-021-22518-0
Turku Bioscience Centre
https://www.utu.fi/fi/yliopisto/turku-bioscience
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
ELIXIR Finland is pleased to announce participation in ELIXIR-STEERS, a three-year EU funded project launched today with a total budget of €4M.The project is a collaboration between all ELIXIR Nodes, encompassing 36 institutes across 23 countries, plus EMBL-EBI. It aims to enhance large-scale, cross-border federated analysis in the life sciences throughout the European Research Area.
ELIXIR-STEERS addresses the need for good software and workflow management, sometimes overlooked by funders and policymakers, which is essential for reproducible and efficient analysis of life science data. By adopting common analysis tools and good workflow practices, the project aims to minimise duplication, reduce energy consumption and lower the carbon footprint in computational life science, particularly in resource-intensive applications like AI.
The ELIXIR-STEERS project builds on the achievements of two previous EU-funded projects: ELIXIR-EXCELERATE (2015-19), which helped establish ELIXIR as a coordinated European life science infrastructure, and ELIXIR-CONVERGE (2020-23), which enhanced data management and stewardship across ELIXIR Nodes.
Andy Smith, ELIXIR Interim Director and Coordinator of ELIXIR-STEERS said “We are excited to initiate the ELIXIR-STEERS project, focusing on expanding ELIXIR’s expertise in research software and workflows. This project is an important step towards strengthening the emerging European federated data landscape and optimising environmental sustainability in computational practices”.
By improving standards and best practices in research software and workflows across the ELIXIR Nodes, ELIXIR-STEERS strengthens Europe’s position in global research while supporting environmental sustainability and enhancing international collaboration.
For further information about ELIXIR-STEERS, please contact steers-coordination@elixir-europe.org.
FInd out more
Liquid biopsy is becoming an increasingly important diagnostic technique for cancers.
Liquid biopsy is based on the fact that cells in the body release DNA into the bloodstream and bodily fluids, a form known as cell-free DNA (cfDNA). In other words, DNA from cancer cells is released into the patients’ bloodstream, containing mutations specific to cancer. The cfDNA is sequenced, revealing the genetic alterations present in the tumour.
The more intact – less fragmented – cell-free DNA is, the more it is associated with a poor prognosis for breast cancer.
“We have access to CSC’s resources specifically for machine learning purposes. So far, we have developed cancer risk analytics, but the same models are utilised in the further work with these liquid biopsy results. This data has not been further processed yet,” says professor Arto Mannermaa from the University of Eastern Finland.
Read more here

Cancer cell DNA can be isolated from a breast cancer patient’s blood sample, and the degree of fragmentation of the DNA can be assessed to determine whether the patient has a poor or good prognosis for treatment.
Breast cancer is the most common cancer in women, with over two million women being diagnosed with it in 2020. Fortunately, the prognosis for breast cancer has improved, because it can be detected in the early stages. In addition, treatment methods have advanced. One of these is liquid biopsy, which is becoming an increasingly important diagnostic technique for cancers.
Professor Arto Mannermaa’s research group, specialising in personalised medicine and biobanking, has been studying liquid biopsy since 2015. Liquid biopsy is based on the fact that cells in the body release DNA into the bloodstream and bodily fluids, a form known as cell-free DNA (cfDNA). In other words, DNA from cancer cells is released into the patients’ bloodstream, containing mutations specific to cancer. The cfDNA is sequenced, revealing the genetic alterations present in the tumour.
“We have investigated the concentration, fragmentation level and mutations of cell-free DNA that are associated with the prognosis of breast cancer patients. Similar connections can be found in several other forms of cancer,” says researcher Jouni Kujala. Kujala works in Mannermaa’s research group at the University of Eastern Finland.
“This is a research topic that involves many computational aspects, especially processing sequencing data,” Kujala says. In the future, Kujala plans to focus on cell-free microRNA.
“It is an entirely different type of nucleic acid that can be isolated from the blood samples of cancer patients. Cell-free microRNA regulates gene function, but its predictive value is not yet fully understood.”
Mannermaa’s research group has studied the fragmentation of cell-free DNA. Based on this research, the prognosis of breast cancer patients can now be assessed. The result is important, because the method helps identify breast cancer patients with a poor prognosis earlier and more accurately than before. Early detection is one of the key ways to reduce breast cancer mortality.
The researchers have analysed the causal association of cell-free DNA integrity with breast cancer treatment prognoses.
“When cancer cells release cell-free DNA into the bloodstream, it gradually begins to fragment into smaller pieces until it breaks down completely.”
Integrity reflects the degree of DNA fragmentation – that is, how much DNA has fragmented in the blood.
“The more intact – less fragmented – cell-free DNA is, the more it is associated with a poor prognosis for breast cancer.”
The results of the study by Mannermaa’s research group were made possible by extensive patient data, gathered by the Kuopio Breast Cancer Project (KBCP).
“The KBCP includes over 500 breast cancer patients, and comprehensive data has been collected from them. We know their lifestyles and the cancer treatments they have received. We have follow-up data for up to 25 years, which is exceptionally long even on an international scale.”
In this study, the sample included breast cancer patients who had not yet started any form of cancer treatment.
“The sample consisted of early-stage breast cancer patients who were initially considered to have a good prognosis.”
There was a clear rationale for selecting such a sample, as breast cancer recurs in up to a third of patients and is the most common cancer-related cause of death in women. The goal of Mannermaa’s group is that in the future, patients with aggressive breast cancer could be identified even earlier through integrity measurement, and directed to enhanced treatment and monitoring if necessary.
The measurement of DNA integrity is a simple one. The isolated sample is placed in a measuring device that determines the relative proportion of DNA fragments in the sample. Then it is possible to calculate the degree of integrity of the cell-free DNA in the sample.
According to Kujala, even such a simple value describing the quality of a DNA sample can be useful in predicting cancer outcomes. In the future, this method could be put to further use in training artificial intelligence.
“When measuring the concentration and integrity of cell-free DNA, these are purely quality indicators. They are not currently used in assessing the patient’s prognosis. The actual diagnostic aspect has largely focused on mutations and other features of DNA. Machine learning could enable more effective use of this data – it is collected from all samples that are examined, but it is hardly utilised at all.”
Mannermaa’s team is developing algorithms that learn from genomic data and clinical information to identify and predict risk factors for breast cancer. Genomic and clinical data are combined to form an AI model that not only helps to determine the risk of illness, but also in drawing up individual treatment plans.
The amount of data in Mannermaa’s team’s study is so huge that it requires the supercomputing capacity of the Finnish ELIXIR node of the CSC – IT Center for Science.
“We have access to CSC’s resources specifically for machine learning purposes. So far, we have developed cancer risk analytics, but the same models are utilised in the further work with these liquid biopsy results. This data has not been further processed yet,” says Mannermaa.
Ari Turunen
23.1.2024
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Improving breast cancer treatment prognoses with liquid biopsy. https://doi.org/10.5281/zenodo.13691344
More information:
Maria Lamminaho, Jouni Kujala, Hanna Peltonen, Maria Tengström, Veli-Matti Kosma ja Arto Mannermaa. High Cell-Free DNA Integrity Is Associated with Poor Breast Cancer Survival. Cancers. 2021.
https://doi.org/10.3390/cancers13184679.
University of Eastern Finland (UEF)
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
THL and Finland’s ELIXIR Node CSC – IT Center for Science were involved in a project called Towards the European Health Data Space (TEHDAS), which included 25 countries and concluded in July 2023. TEHDAS was a collaborative project that conducted preparatory work and provided recommendations to promote the implementation of the EHDS. According to Doupi, the wider utilization of health data imposes new requirements, particularly when it comes to cross-border secondary use.
“The goal of the TEHDAS project was to determine how cross-border/EU-level secondary use of data could be organised and what kind of legislation is needed”, says ays Persephone Doupi, Senior Medical Officer at the Finnish Institute for Health and Welfare (THL).
She works in THL’s Data and Analytics Unit as the coordinator for international secondary use data management projects. The unit promotes the diverse use and interoperability of data resources. One of its tasks is to develop open data interfaces and services.
”What would be the governance model and tasks of data permit authorities, and how could access to data be organised? At the same time, the project involved investigating what kind of information-system architecture and technical solutions would be most suitable, how to ensure data quality and the interoperability of datasets, and what standards are available.”
TEHDAS continues with the TEHDAS-2 project. It includes a work package that addresses secure environments, with the goal of creating sustainable solutions for the secure management of data.
TEHDAS continues with the TEHDAS-2 project. It includes a work package that addresses secure environments, with the goal of creating sustainable solutions for the secure management of data.
“CSC is involved in the development of secure environments. The biggest question concerns the definitions, administrative models and solutions for secure environments. CSC has had a central role from the beginning, especially when considering datasets related to genomics. In Finland, there are no other actors besides CSC that could provide similar expertise in this regard.”
Read more here

Cross-border health data has not been appropriately utilised in research and decision-making in Europe. There is a strong desire in the EU to create infrastructure for the secondary use of health data and sensitive genomic data. The European Health Data Space (EHDS) and specifically its HealthData@EU environment is being established for this purpose.
The pandemic caused by the COVID-19 virus revealed significant shortcomings in the sharing and coordination of health data in Europe. It was recognised how crucial it is to ensure secure access to health data across Member States, especially when people were moving freely within the EU during the pandemic. Decision-makers faced difficulties during the pandemic in obtaining the necessary electronic health information. Individualised drug treatments are also possible only if patient data is available and stored, pre-processed and classified in a consistent manner in all countries.
Sharing health data across borders has been surprisingly difficult. The availability of personal health information and genetic data in digital form varies between Member States. Legislation also varies. The EHDS ensures the coordination and consistency of primary and secondary use of health data. Permanent structures are now being established for collaboration.
The EHDS will markedly transform healthcare in the coming decades. The EHDS creates a common space for managing and transferring electronic health data, such as patient records, patient registers and genomic data. It also gives researchers the opportunity to access health data reliably. Privacy protection is also maintained.
“While discussing the legislative framework in Europe, infrastructure is being built to move forward. What’s positive is that the views of experts, such as researchers, have been taken into account in legislation. Large pilot projects were initiated even before the legislative work began,” says Persephone Doupi, Senior Medical Officer at the Finnish Institute for Health and Welfare (THL).
She works in THL’s Data and Analytics Unit as the coordinator for international secondary use data management projects. The unit promotes the diverse use and interoperability of data resources. One of its tasks is to develop open data interfaces and services.
“We need to consider the entire lifespan of data, the model and the approach as a whole. This is especially evident when talking about data quality and standardisation – particularly, how information in different systems is semantically interoperable. We should understand this early enough. What’s absolutely central is how healthcare professionals document health data in different countries,” says Doupi.

THL and Finland’s ELIXIR Node CSC – IT Center for Science were involved in a project called Towards the European Health Data Space (TEHDAS), which included 25 countries and concluded in July 2023. TEHDAS was a collaborative project that conducted preparatory work and provided recommendations to promote the implementation of the EHDS. According to Doupi, the wider utilization of health data imposes new requirements, particularly when it comes to cross-border secondary use.
“The goal of the TEHDAS project was to determine how cross-border/EU-level secondary use of data could be organised and what kind of legislation is needed. What would be the governance model and tasks of data permit authorities, and how could access to data be organised? At the same time, the project involved investigating what kind of information-system architecture and technical solutions would be most suitable, how to ensure data quality and the interoperability of datasets, and what standards are available.”
TEHDAS continues with the TEHDAS-2 project. It includes a work package that addresses secure environments, with the goal of creating sustainable solutions for the secure management of data.
“CSC is involved in the development of secure environments. The biggest question concerns the definitions, administrative models and solutions for secure environments. CSC has had a central role from the beginning, especially when considering datasets related to genomics. In Finland, there are no other actors besides CSC that could provide similar expertise in this regard.”
Doupi points out that even if the legislative work makes progress, it will still take years to see the true impact of secondary data use on research. Even at this stage, however, collaboration between different authorities has increased and has even become mandatory, which Doupi sees as a positive development.
“The EHDS enables the study of complex and significant diseases in a more reliable way in the future. In the Nordic countries, for instance, it has been acknowledged that there is not sufficient data for every research subject individually. When we combine all the datasets from the Nordic countries, we get more reliable data for research. This is crucial when studying rare diseases, for example, or the safety and efficacy of medications. The dataset of just one country is not sufficient for studying such topics.”

One good example Doupi mentions is the European Union’s 1+ Million Genomes initiative (1+MG). The initiative aims to enable secure access to genomic data and related clinical datasets to support better research and decision-making. National collections, combined through the 1+MG initiative, together form the Genome of Europe, a vast European database. Over a million genomes had been sequenced by the end of 2022. 1+MG and its follow-up project Beyond 1 Million Genomes (B1MG) are among the world’s largest projects in their category. Operative infrastructure will be in place in 15 countries by 2026. 1+MG and B1MG collaborate closely with the EHDS. The future looks promising, Doupi says.
“I assume that awareness of data quality will increase. At the same time, interdisciplinary collaboration will break down unnecessary silos. Hopefully this will also initiate a societal discussion, for example about the use of artificial intelligence. The processing of large datasets requires artificial intelligence, and there must be a flexible approach to it. We must be constantly vigilant in adapting to new information and changing environments. That will happen through the EHDS.”
Ari Turunen
15.12.2023
Read article in PDF
Citation
Nyrönen, T., & Turunen, A. (2023). The European Health Data Space: health data moves across borders for research purposes. https://doi.org/10.5281/zenodo.13691001
Finnish Institute for Health and Welfare THL
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Gut microbiota have an effect on human health, but how they work is not known well enough.Previous studies have mainly focused on discovering the composition of microbiota, while there is still very little information about the behaviour of gut microbiota at protein level. Metaproteomics is a good method for studying human gut microbiota, because it can identify and categorise the proteins involved.
“Metaproteomics requires massive computing power, because identification is so difficult. As reference, we have millions of possible proteins from thousands of bacteria, which we are trying to identify in the samples. However, we get really accurate and comparable measurements as a result,” says reseacher Tomi Suomi from the University of Turku.
Suomi is a member of the reserach team which is using the computing power of Finland’s ELIXIR node CSC – IT Center for Science. The virtual computers of CSC have been connected as an extension of the local computing cluster of the University of Turku.
According to Suomi, this measurement method seems to be working really well for clinical samples. Future applications may include comparable measurements of stool samples in biobanks.
“We may have hundreds of samples from different individuals. At the moment, using this method alone, we are able to perform sufficiently comparable measurements between individuals, for example as part of studying how diseases develop. With the new method, it will be possible to study large cohorts.”
Read more here
Industrial processes such as mining have led to increased concentrations of nitrogen and heavy metals in soil and water. Post-doctoral researcher Kaisa Lehosmaa works in Anna Maria Pirttilä’s research group at the University of Oulu, studying endophytic microbes living inside moss and other plants and their suitability for water purification.
In Lehosmaa’s research, floating hook-moss proved to be an effective accumulator for metal-rich waters even at low temperatures. When combined with a woodchip bioreactor, the combined unit removed nitrogen particularly well. Lehosmaa and her colleagues used sequencing methods to identify the microbial symbionts of floating hook-moss.
“Sequencing gives an overall picture of the microbial diversity of the moss – that is, how many and what kind of microbes are present. We also want to know which microbial genes are active under various conditions, so that we can understand how microbes could be used more widely in bioremediation.”
The metals and microbes accumulated in the moss tissue are identified through sequencing and traditional microbiological cultivation methods. After identification, metals and microbes are localized in the moss tissue. Identification and localization are used to determine microbes adapted to metal-rich conditions, which could potentially be applied in purification processes. Purification processes are enhanced by adding microbes to the moss tissue.
To analyse microbiome composition, Lehosmaa has used the computational resources of the Finnish ELIXIR node’s CSC – IT Center for Science and its Chipster software.
Read more:

Gut microbiota have an effect on human health, but how they work is not known well enough. The behaviour of microbiota at protein level is studied at the University of Turku by means of metaproteomics.
Previous studies have mainly focused on discovering the composition of microbiota, while there is still very little information about the behaviour of gut microbiota at protein level. Proteins perform most cell operations, and studying them in more detail may help us understand the interaction between cells and their environment. Metaproteomics is a good method for studying human gut microbiota, because it can identify and categorise the proteins involved.
“Previously only the composition of gut microbiota has been profiled. However, it tells us nothing about what is really going on in the gut. There may be dead bacterial mass involved, or the bacteria are not otherwise active. In order to discover how they function, we need metaproteomics. This has only recently become technically possible to measure,” says researcher Tomi Suomi.

Suomi works at the University of Turku in Professor Laura Elo’s research team that has developed a new method for studying microbiota at protein level. Suomi says that the key question is which processes can be detected in gut microbiota. Only now is information becoming available on the metabolic activity of bacteria at protein level. This makes it possible to analyse many dietary factors, such as how baby formulas affect the gut microbiota.
“This enables us to measure and study like never before what happens in the intestine. How do bacteria interact with each other and what determines which bacteria are active? The method we have developed makes use of the latest mass spectrometry technology and computational methods. It enables us to comprehensively measure the protein levels of complex microbial samples.”
The importance of gut microbiota for human health and their role in various illnesses has been identified in recent studies. Potential diseases that can be studied by means of metaproteomics include Crohn’s disease, ulcerative colitis, colorectal cancer or diabetes.
“Other areas of application may include allergies. There are a number of conditions with which a link with gut microbiota has been suggested at least on some level. The methods we have developed can be applied directly to such studies.”

Mass spectrometry is used to identify isolated proteins. Proteins in a sample are broken down into smaller amino acid chains, or peptides, which are then analysed with a mass spectrometer. Proteins are identified by deducing the amino acid chains of the peptides on the basis of measured masses by means of computational methods. The measured masses are compared to databases containing, for example, known protein sequences that best match the peptide masses.
In the new DIA methods (data-independent acquisition), the aim is to measure and fragment all peptides contained in the sample for identification. However, identification is more difficult than normally, because individual spectrums may represent more than one peptide. A similar DIA-based mass spectrometry has not been used before in connection with metaproteomics. The research team has developed algorithms to identify peptide sequences and to search for them from databases.
According to Suomi, this is computationally difficult, because the new DIA methods attempt to measure everything in the sample: all peptides originating from various types of microbes contained in it.
“Metaproteomics requires massive computing power, because identification is so difficult. As reference, we have millions of possible proteins from thousands of bacteria, which we are trying to identify in the samples. However, we get really accurate and comparable measurements as a result.”
The research team is using the computing power of Finland’s ELIXIR node CSC – IT Center for Science. The virtual computers of CSC have been connected as an extension of the local computing cluster of the University of Turku.
According to Suomi, this measurement method seems to be working really well for clinical samples. Future applications may include comparable measurements of stool samples in biobanks.
“We may have hundreds of samples from different individuals. At the moment, using this method alone, we are able to perform sufficiently comparable measurements between individuals, for example as part of studying how diseases develop. With the new method, it will be possible to study large cohorts.”
The new method offers a variety of applications, Suomi says.
“In theory, the methods could be used more extensively in metaproteomics applications, such as ocean and wastewater study or the analysis of soil samples, but we have not yet tested our methods for them.”
Ari Turunen
16.11.2023
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2023). Artificial intelligence helps researchers find suitable drugs based on patient’s genetic data and cancer cell samples. https://doi.org/10.5281/zenodo.10796468
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Kaisa Lehosmaa studies the potential of Arctic microbes and plants to purify industrial wastewater. Such method can help to reduce environmental contamination caused by the mining industry. In addition to mining, nitrogen and heavy metal loads to waterbodies come from wastewater treatment plants, stormwater, agriculture, and peat production. The suitable plant-associated microbes for bioremediation are mainly determined by sequencing methods, but also by isolating microbes from the moss.
Industrial processes such as mining have led to increased concentrations of nitrogen and heavy metals in soil and water. Post-doctoral researcher Lehosmaa works in Anna Maria Pirttilä’s research group at the University of Oulu, studying endophytic microbes living inside moss and other plants and their suitability for water purification. The use of microbial ‘symbionts’ in bioremediation is a relatively little researched field. Some plants can store or even evaporate harmful substances such as metals and nutrients into the atmosphere. Plant-associated microbes play an important role in nutrient and metal uptake and transformation within the plants.
In particular, Lehosmaa has studied floating hook-moss, Warnstorfia fluitans, which grows in Finland in low-nutrient peatlands and in groundwater-dependent spring ecosystems. The moss also grows in Pyhäjärvi in areas around the Pyhäsalmi mine. Pyhäsalmi is Europe’s deepest base metal mine, yielding copper and zinc.
“We have found this moss in a mining area, which is naturally adapted to the harsh conditions. The microbes found in floating hook-moss tissue can be used with the moss to improve purification of mining wastewater in cold climate conditions,” Lehosmaa explains.
Mining activities create acidic and metal-rich wastewater that is made mobile by gravity. The wastewater is highly acidic, and contains high levels of metals considered harmful – zinc, aluminium, copper and cadmium. Such wastewater must be treated and cleaned carefully, as it is harmful to the environment.
In Lehosmaa’s research, floating hook-moss proved to be an effective accumulator for metal-rich waters even at low temperatures. When combined with a woodchip bioreactor, the combined unit removed nitrogen particularly well. Lehosmaa and her colleagues used sequencing methods to identify the microbial symbionts of floating hook-moss.
“Sequencing gives an overall picture of the microbial diversity of the moss – that is, how many and what kind of microbes are present.
We also want to know which microbial genes are active under various conditions, so that we can understand how microbes could be used more widely in bioremediation.”

To analyse microbiome composition, Lehosmaa has used the computational resources of the Finnish ELIXIR node’s CSC – IT Center for Science and its Chipster software.
“The microbiome of the moss is quite unknown – the microbial symbionts of plants in general are relatively poorly known. We have identified symbionts in the moss using amplicon and genome sequencing.”
Amplicon sequencing targets the specific gene regions, in this case, the 16S and ITS ribosomal RNA (rRNA) gene regions. The 16S and ITS rRNA gene regions have remained the same over millions of years in evolution for bacteria and fungi, which is why these regions can be used to identify different species. The 16S and ITS rRNA gene regions are sequenced and identified through publicly accessible databases.
“After identifying the microbes, the next step is to find out what they do. We already have preliminary results that interesting processes take place within the moss tissue.”
According to Lehosmaa, it is important to know what happens inside the moss and how the microbes are able to process metals.
“Acidic water usually contains metals in soluble form. It is not possible to remove metals from water, since they are inorganic compounds. However, we can use microbial symbionts to change the solubility of metals. Bioremediation often uses live microbes to precipitate metals into particulate form, thereby making them easier to control and remove.”
It is crucial that the microbes found can also be grown in the laboratory.
“We can’t use microbes in purification purposes if we can’t grow and thus add them to the same or different plant species to promote metal uptake processes,” Lehosmaa says. Microbial symbionts help plants to survive in difficult biological conditions.
In addition to moss, one effective plant used in bioremediation is the common reed. Like moss, it absorbs harmful substances effectively. The common reed creates a large amount of biomass and grows easily. The algae in its rhizome binds soil structures and prevents blue-green algal blooms. The common reed is also used for metal recovery; called phytomining or agromining. Lehosmaa and her colleagues have mapped the accumulation of copper and zinc in the common reed at the Pyhäsalmi mine.
The next goal is to assess the ability of other natural plants, fungi, and bacteria adapted to northern climate conditions to remove nitrogen and metals from water.
“Since the indicator microbes can be cultivated and added, the next step is to expand the research to other mosses. It is interesting to find out whether the microbes can function as well in other plants than floating hook-moss.”
Ari Turunen
31.10.2023
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2023). Purifying mining wastewater with plant-associated microbes. https://doi.org/10.5281/zenodo.13690962
For more information:
University of Oulu
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Vilja Pietiäinen, senior scientist and adjunct professor (docent) in cell and molecular biology at the Finnish Institute for Molecular Medicine (FIMM), wants to make cancer treatments more individualised.
With the help of microscopic imaging, researchers can inspect how the drugs influence the cancer cells. Machine learning models allow researchers to more effectively analyse the images of cancer cells. The artificial intelligence used by the research team has been trained with the Finnish ELIXIR node’s, CSC’s high-performance computing clusters.
“We call this phenotypic imaging. Microscopic imaging allow us to identify hundreds of different cell characteristics from images of drug-treated cancer cells. This information is important for further training the machine learning model. If we are able to clearly identify certain phenotypes, we can also teach the machine to do the same by showing it how certain cells have responded to a certain drug. After this, we can provide the machine with a new dataset, in which case it will be able to classify the cells by how they show up in the images. On the other hand, artificial intelligence, especially deep-learning solutions, can also help us to discover traits or phenotypes that we as humans are not able to either detect or classify.”
The iCAN-project utilises the SD Connect service provided by the Finnish ELIXIR node CSC for transferring sequencing data to the Academics environment.
The data is encrypted using Crypt4GH, a protected standard encryption method developed by the Global Alliance for Genomics & Health for sharing human genetic information.
“This ensures that the information can be used in all of the services included in CSC’s SD service suite, and may even be potentially shared with other service providers who possess similar information.”
Read more here
The Bioinformatics Center at the University of Eastern Finland (UEF), led by Virpi Ahola, is developing new applications for analysing biomedical and multimodal data.
Translational medicine uses basic research in clinical trials, but also patient samples and disease models to identify disease mechanisms and drug targets. The research approach is interdisciplinary, which provides a good starting point for research but also improves treatments for patients.
“What is delaying the era of translational medicine is that we simply don’t know enough. The idea behind combining several different data sources is to obtain more information. The integration is very much computational, and requires CSC – IT Center for Science’s resources and infrastructures like ELIXIR.”
Ahola is a tireless advocate for the openness and reuse of data, and for the development of methods and infrastructures that facilitate and encourage this.
Referring to Biocenter Finland, Ahola says that more should be done together. The centre brings together seven biocentres from different Finnish universities. It should not be impossible to increase collaboration between different biocentres and internationally, for example through the Finnish ELIXIR node CSC.
“ELIXIR is an avenue for us to network and learn from the experiences of other bioinformatics core facilities, and to be part of discussions where research infrastructure issues are brought up and new initiatives are taken.”
Because new technologies produce large and complex data sets, research infrastructures should also include data science experts, not research equipment alone.
“To make effective use of data, the computing capacity offered by CSC, for example, is not enough. Data processing and reuse also requires staff with expertise in the field. As I see it, better resourcing and systematic collaboration between biocentres could substantially facilitate and improve the processing, integration and reuse of large omics data sets.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Professor Merja Heinäniemi and her research team at the University of Eastern Finland use computational methods to interpret cancer samples in order to determine which cellular processes are defective and how cells behave during drug treatments.
In 2019, Heinäniemi and other researchers collected a large dataset of more than 10,000 patient samples. This data on haematological malignancies (HEMAP) will continue to be shared with researchers through the Finnish ELIXIR node, CSC – IT Center for Science. They were able to deduce more than 30 types of cancer from this dataset by computational methods. In addition, new disease biomarkers and new drug candidates were discovered when the data was combined with drug target databases.HEMAP, hosted by CSC, now enables target gene analysis of drug molecules.
Now Heinäniemi’s team has started to use the neural network models they have developed. Data is collected from different studies for this model.
“Our leukaemia project focuses on childhood cancers. It’s a very rare type of cancer: if we don’t get the data combined, the data sets will be very small. The aim is to use CSC’s infrastructure for these projects. This would allow the data to be processed and made available to the scientific community.”
Sharing data requires building trust. In practice, this means working with the patients involved in the projects.
“It’s really important that they are involved and that their data is stored securely, and that researchers are able to do their work with the data. This is what CSC is enabling at national and EU level through its involvement in the ELIXIR infrastructure.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Researchers at the University of Turku discovered a microRNA that may be an early indicator of the risk of developing type 1 diabetes.
Professor Laura Elo and her research group for computational biomedicine at the University of Turku are developing tools for the diagnosis and treatment of complex diseases, such as diabetes, cancer and rheumatoid arthritis. The group screens patient data using computational methods to find signs of diseases and their risk factors.
Elo, Research Director at the Turku Bioscience Centre, is mining patient data for various biomarkers that may help predict the onset of diseases or tell something about the response to treatment. A biomarker is a feature that indicates a change in a biological state, in genes or proteins, for example.
According to Elo, future study of diseases should make use of the various ‘omics’, such as genomics (DNA), proteomics (proteins), transcriptomics (RNA) or metabolomics (metabolism). Elo’s group has been using the computing resources of Finland’s ELIXIR Center CSC to process the extensive data.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
A single tissue sample of a patient can with modern technology be used to profile thousands, sometimes even tens of thousands of cells. We can reliably profile which type of cells it contains.
“Even if the cells may look identical under the microscope, their function may turn out to be quite different once we are able to view gene expression with single-cell accuracy,” says researcher Päivi Saavalainen.
Saavalainen works at the Folkhälsan Research Center and she is also the CEO of a company called SCellex that specialises in single-cell technology. Saavalainen, who together with the Finland ELIXIR Node of CSC has organised single-cell analytics courses for researchers, considers single-cell technology one of the most revolutionary methods in biosciences over the last few years.
“In cancer research, for example, it is important to obtain information about a single cell. The fact is that cancer cells change all the time, meaning that each cell begins to be different. There are also gene mutations, as a result of which certain genes are activated and others deactivated.”
The datasets are huge and the services of Finland ELIXIR Node of CSC can help to perform the computation. Saavalainen says that the AI models are vital.
“If a sample contains tens of thousands of cells and all of them are subject to tens of thousands of gene measurement results, then not only the data on the microbeads but also the actual biological RNA data is immensely complex. You need AI to analyse it. AI can find such new information that simply would not be possible with traditional analysis tools. I think the computing power of CSC is sufficient even to solve our challenging AI models.”
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Celiac disease is considered an autoimmune disease, in which the body’s defence system mistakenly targets its own tissues. Although the exact causal mechanism of celiac disease is not yet known, there is a hereditary predisposition to its onset.
Immunologist Helka Kaunisto from the Celiac Disease Research Center at Tampere University studies dermatitis herpetiformis, the skin manifestation of celiac disease.
Kaunisto studies immune cells and the immune response to find out why some people with celiac disease develop dermatitis herpetiformis.
“It should be remembered that the gut and the skin have different layers that function in different immunological ways. Dermatitis herpetiformis is a really good target for studying the extra-intestinal symptoms of celiac disease. This information can then also be used to investigate other autoimmune diseases. For example, how can rheumatism start in one place and then spread elsewhere and become systemic?”
The study involves analysis of sensitive data obtained from patient’s tissue samples. Patients consented to participate in the study. As this is information subject to the European Union’s General Data Protection Regulation (GDPR), the data will be processed and used in CSC’s sensitive data services (SD Desktop and SD Connect).
The sequencing has been done in collaboration with the University of Helsinki, and the data is stored encrypted in SD Connect and analysed with SD Desktop.
Kaunisto did not have previous experience of the high-performance computing or sensitive data services.
“I started using the Sensitive Data Services because I needed more computational power than what was available for me through the university, and I needed a secure environment for this computational power. I find the services very easy to use as the online guides are very thorough. If I have a problem I can’t solve myself the helpdesk is always very helpful.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Geneticist Petri Auvinen and his research team are using DNA samples to find out what has been happening in the Baltic ecosystem during the past 10,000 years.
“Our aim is to collect sediments samples as far down as possible in the Baltic Sea bed in order to study the history of the Baltic Sea basin. We also take samples deep in marshland, providing us with information about the history of the soil,” says Auvinen.
Petri Auvinen is a research director at the Institute of Biotechnology, University of Helsinki. His research focuses on genomics and metagenomics. While genomics looks at the entire genome of an organism, metagenomics can study and sequence a number of organisms, such as microbes, from a single sample at the same time. The study of micro-organisms has advanced in leaps and bounds. A sequence sample can be taken from any environment, soil or gut to determine the microbiota composition. The term used is microbiome, denoting the microbiota of a specific habitat and its genome, that is, the metagenome.
Some of the data is analysed using the CSC – IT Center for Science’s ePouta hardware, but some must be done with the team’s own.
“The software can be so complex that it cannot be run on the CSC system. We use some virtual computers, but they have their limitations, too. We also have runs that may continue uninterrupted for months. CSC has thousands of users, and the CSC environment obviously has some maintenance outages. For example, when we were working on the genome of the Saimaa ringed seal, the first major assemblies took a thousand hours.”
“Currently we are able to work on large genomes a hundred times better than a few years ago. But there is more and more data all the time, and we must be able to store it efficiently and in a way that it is comparable to other data. We will continue to work with CSC in data storage, transfer and calculation.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Organoids grown from stem cells enable new ways to model a variety of diseases such as cancer. At the University of Oulu, new techniques to engineer embryonic tissue are used to find cancer genes.
Professor Seppo Vainio says that the aim is to improve legislation to allow safeguarding the anonymity of private individuals in research activities such as creating human organoids and the patient records related to them. Currently, university hospitals and the Finnish Social and Health Data Permit Authority Findata are responsible for the administration of clinical test results of operational patient care.
“Researchers can reuse the data on gene-level changes associated with certain human diseases observed in stem cells and the organoids grown from them. This is basic research and produces experimental data such as image analysis data and gene-level data. CSC –IT Center for Science already provides the framework needed to store such digital material.”
According to Vainio, anonymisation is not as relevant when data on experimental cell lines is produced, which is why it would be possible to manage such data through CSC.
“If there was a need to link this data to patient records, it could be done with Findata cooperation. Creating organoids from samples donated by patients, for example, could also be made subject to licence in Finland.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
SD Connect is a service for collecting and storing sensitive research data during the active phase of a research project while SD Desktop users can directly access and manage that data in a virtual computing environment. The services are accessible via a web user interface, from the user’s own computer.
”Once users upload their sensitive data to CSC, these data are always kept encrypted when stored, transferred or processed within our services. Decryption is done only when data is made available for authorised users within the SD Desktop service”, says Francesca Morello from the Finnish ELIXIR node, CSC.
”Researchers can access the workspace with just a few clicks. While SD services are suitable for managing sensitive data from any research field, we are working on further facilitating the use of the services, from fully automating data encryption to streamlining the computing environment customisation.”
The services are available to researchers and students affiliated with Finnish academic organisations, research institutes, and their international collaborators. Using CSC services requires to register a CSC account. While SD Connect and SD Desktop have been designed to facilitate collaboration between organisations, the data is always stored in CSC’s cloud services in Finland.
Data is stored in CSC:s Allas and ePouta platforms. Metadata can be sent outside Finland.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Metagenomics means the study of microbial DNA directly in their natural living environment. The term generally refers to bacterial genomes in a sample, but it also means the genomes of other microorganisms, such as those of archaea and fungi and also the genomes of the eukaryotes inhabiting the sample of interest. Metagenomics can thus be used to study and sequence multiple organisms simultaneously from a single sample. Jenni Hultman, Senior Scientist at the Natural Resources Institute Finland (Luke), and her colleagues sequenced millions of genomes from tundra soil microbiomes. To do this, they needed the computing power of the Finnish ELIXIR Node CSC – IT Center for Science, because the volume of data in the materials runs in terabytes.
“We have learned what kind of microbes live in the subarctic area and what they do. Processing the data material has taken up an incredible amount of computing power. We discovered several previously unknown microbiota and genomes.”
DNA sequences obtained from the samples have been analysed, aiming to identify new species and their relationships. By analysing RNA sequences, they have learned what the microbiota were doing at the time of sampling.
“What I’m particularly excited about is being able to visit the same sites in wintertime. Now we are able to learn about seasonal variation, that is, what takes place in the microbiota in the summer and winter and how warm autumns contribute to the microbial activity.”
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org
https://www.elixir-europe.org
“Good scientific practice involves making sure that data is well documented and remains usable throughout the research process, and in such a way that results can be verified later. It is important that researchers and information systems are able to find and access compatible and reusable research outputs. To ensure this, the FAIR principles were set out by a consortium of scientists and organisations in 2016,” explains CSC’s data management specialist Minna Ahokas.
“With instructions and tools provided by ELIXIR, it is easier for researchers to make their data findable, accessible, interoperable and reusable.”
The RDMkit website, created in cooperation with the ELIXIR nodes of the member countries, aims to support and harmonise data management practices in Europe.
RDMkit includes instructions and tips concerning the entire life cycle of data – from data management planning and data analyses right up to publication and reuse.
“RDMkit has been implemented in a way that anyone dealing with data is able to access the tools. It offers not only instructions but also links to services that researchers and any support personnel may need at various stages of data management.”
Finland’s ELIXIR node, CSC, is one of the parties producing content and maintaining the toolkit.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Post-doctoral Researcher Guilhem Sommeria-Klein at the Academy of Finland is developing mathematical models of microbial communities.
Guilhem Sommeria-Klein wants to develop more efficient methods for analysing and interpreting the data. Rather than solely specialising in mathematics or biology, his research aims to connect various disciplines.
“This represents the core area of our research team focusing on computational analysis, and the work of Sommeria-Klein supports it well”, says Leo Lahti, Associate Professor in Data Science at University of Turku, whose team develops machine learning models for screening microbial communities.
“Microbial ecology is in desperate need of this type of basic computational studies. These models will help break complex microbial ecosystems down to a few basic structures. Ocean microbiome research could also be useful in monitoring the changes in the state of the Baltic Sea. Models based on statistical reasoning can take into account any prior information and describe the uncertainty in the results. The high-performance computing services of CSC – IT Center for Science are needed to fit these models to the data.”
In the future, Sommeria-Klein wants to continue studying ecosystems that differ from each other.
“We want the perspective on microbial ecology to be consistent across ecosystems, as it is of major importance for various societal issues, such as human health, the ocean food chain and the global carbon cycle.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Katariina Pärnänen from the University of Turku uses metagenome sequence data that is stored in extensive open databases. She uses supercomputers at CSC – IT Center for Science (the Finnish ELIXIR node) to analyse the data,allowing her to identify different species of bacteria and their resistance genes.
“In some species of bacteria, it is possible to identify genes that appear in its genome only once. By comparing these with genes in the databases of other species, we can identify the species in question. We look for matches with resistance genes stored in databases. Then we can say that the person has ten resistance genes in their stool sample, or that they carry a certain amount of coliform bacteria.”
Katariina Pärnänen would like to extend her research to the world’s entire human population and study the gut microbe samples that have been metagenome sequenced and are openly available in databases.
“It would be interesting to discuss the technical feasibility of such a project with CSC experts.It would also promote open science, because the identified resistance genes and species found in microbiomes could also be stored for use by other researchers.”
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The PerMedCoEproject combines clinical patient data with data related to the operation of genes, proteins and cells. The goal is to develop tools that can be used in precision medicine. The modelling of cells in detail is, however, a major undertaking, requiring a lot of computing power by supercomputers.
CSC – IT Center for Science and the Barcelona Supercomputing Center (BSC), alongside ten other academic and commercial players, kicked off in October 2020 a European Commission Centre of Excellence a project called HPC/Exascale Center of Excellence for Personalised Medicine (PerMedCoE). The project develops cell-level modelling software suitable for high-performance computing. Thanks to high-performance computing, biological data such as genomics and proteomics can be made part of precision medicine, because data can be analysed much faster. Diagnoses, for example, should in future be possible to make within hours or days. PerMedCoE is part of the ELIXIR Finland’s development programme.
The results and tools of PerMedCoE are open to all researchers.
“When the project ends in summer 2023, we will have updated versions of modelling tools developed from open source code, and these will be made available to the research community. The project will also create new expertise to support the use of precision medicine tools in CSC computing environments,” says Jesse Harrison from CSC.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
“We need the resources provided by CSC to obtain comprehensible information from the large-scale sequencing data that we can then analyse statistically. We increasingly use these services as a platform for cooperation. We can build workflows with other research groups and make the data available through CSC, and the data analysis platform is also in one place, on the CSC servers. It is also important that the bioinformatics data resources provided by ELIXIR can be accessed via CSC. We are also increasingly using these services to provide training in computational research methods.”
Leo Lahti, Associate Professor of Data Science from the University of Turku, is developing the machine learning models with his research group and partners for screening microbial signatures from large-scale data collections.
‘The microbiome is just one element in development of diseases, but it is an element that we haven’t been able to study as extensively before, because the effective data collection methods we have now have become available only very recently,” says Lahti.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The large-scale FINRISK population survey of risk factors for chronic, noncommunicable diseases has been used to collect health data on the population every five years since 1972.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Researcher Heidi Marjonen took part in a study in which more than 3,000 persons were given information of their 10-year disease risk for the most common diseases in Finland. She works as a genome expert at THL Biobank, processing genomic data of all THL Biobank cohorts. When genetic data is combined with clinical data, one can predict individual´s disease susceptibility. The overall 10-year risk evaluation was based on the genetic data and other traditional risk factors, such as gender, age, body mass index, blood pressure and cholesterol levels. The genetic risk was calculated as a personal polygenic risk score, taking into account millions genetic variations.
This data was stored on the ePouta platform for sensitive data in Finland’s ELIXIR node CSC, which enables a secure transfer between the portal’s user interface and the database.
The polygenic risk score, says Heidi Marjonen, is a major research trend. The risk score is a single value that reveals the genetic burden of a disease.
“Researchers receive information about the genome in a convenient way and allows to study the effect of genome on a disease or other traits in an individual.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The iCAN-PEDI study, investigating drug treatments and drug responses in children with cancer, is a part of the large-scale iCAN Flagship project of Academy of Finland. The study combines genetic and epigenetic information on the patients’ cancer with data on the drug testing of patient derived cancer cells. Together with collaborators, the team also develops artificial intelligence (AI) -guided analysis for the drug testing data. The project aims to deliver findings that may affect treatment approaches back to the doctors. This helps the doctors to construct more individualised treatment approaches.
Vilja Pietiäinen, senior scientist and adjunct professor (docent) in cell and molecular biology at the Finnish Institute for Molecular Medicine (FIMM), leads the iCAN-PEDI project with Minna Koskenvuo, a clinician (in Pediatric Hematology and Oncology) at the HUS New Children’s hospital and at Turku University hospital. In addition, many clinicians from HUS’s New Children’s Hospital and researchers from the University of Helsinki are involved in the project. Pietiäinen says she wants to make cancer treatments more individualised.
“From a medical perspective, the way we treat childhood cancers is already individualised. However, studying children’s tumours on a molecular level can help us find more effective drugs for specific types of cancers. The solid tumours in children are often heterogeneous and difficult to diagnose only based on pathology. In addition, some of these tumours are very rare. The types of cancer seen in children generally include fewer genetic changes, which means more molecular-level data is required for a diagnosis. The diagnosis, in turn, will affect the treatment approach.”
According to Pietiäinen, collecting large amounts of data on individual patients can significantly improve the diagnostic process and help find new ways to classify different cancers. It will also allow researchers to understand how much variation can be found even within well-known types of cancer.
“We want to better understand why a certain patient responds to drugs the way that they do. This will allow us to develop better and more individualised ways for choosing a treatment approach for a specific patient.”
She and her team combines patient’s molecular-level cancer data with cell models that represent each patient’s individual cancer cells. Exome sequencing allows researchers to examine the information of roughly 20,000 genes in a single run. Transcriptomics, in turn, makes it possible to analyse thousands of RNA molecules simultaneously. This process provides information on how different genes are expressed. Tissue imaging serves to illustrate the biomarkers expressed by different types of cancer tissue. The resulting data is stored in HUS Acamedic, the secure environment used by the iCAN project.
Pietiäinen says that often, genetic data alone is not enough to determine how an individual patient’s cancer will respond to a specific drug.
“We need to study the drug responses of individual patient’s cancer cells with the help of microscopic imaging in the laboratory. Cancer is a very heterogeneous disease: not all cells will necessarily respond to the same drug(s). However, we are also interested in those cells that do not show a response, and have developed the resistance to the used treatments. A combination of different drugs may be required to eradicate all the cancer cells.”

Once the study-consented patient has been operated on, a cancer tissue sample is sent directly to a pathologist, who do the diagnosis and will forward the additional sample directly to the researchers involved in the study. Drug susceptibility is tested with a multi-well cell culturing plate in a process that utilises robotics. The wells are very small, which means that only a small amount of the valuable cell samples is required. A single plate can be used to test dozens of drugs at a time.
Researchers fill the wells with different concentrations of specific drugs and cancer cell samples. With the help of microscopic imaging, they can inspect how the drugs influence the cancer cells in the wells of the plate. Machine learning models allow researchers to more effectively analyse the images of cancer cells. The artificial intelligence used by the research team has been trained with the Finnish ELIXIR node’s, CSC’s high-performance computing clusters.
“We call this phenotypic imaging. Microscopic imaging allow us to identify hundreds of different cell characteristics from images of drug-treated cancer cells. This information is important for further training the machine learning model. If we are able to clearly identify certain phenotypes, we can also teach the machine to do the same by showing it how certain cells have responded to a certain drug. After this, we can provide the machine with a new dataset, in which case it will be able to classify the cells by how they show up in the images. On the other hand, artificial intelligence, especially deep-learning solutions, can also help us to discover traits or phenotypes that we as humans are not able to either detect or classify.”
Once the hundreds of analysed traits are fed into the artificial intelligence, it will be able to differentiate between different drug responses at single-cell level. The data can also be used to sort patients into groups based on the drug responses they exhibit.
Identifying the optimal drug response requires several different data sources. As an example, Pietiäinen mentions a large European project (ERA PerMed) that the project researchers were previously involved in.
“We know that there is currently no targeted drug treatment for up to 90 per cent of cancerous gene mutations. Therefore, we were only able to partially determine the efficacy of different drugs and drug targets for different drugs based on genetic information. However, drug testing did show that patients’ cells responded to certain drugs.”
Pietiäinen considers it crucial to be able to compare drug testing data from cancer studies to the response shown by healthy cells, for example.
“This way, we will be able to see such things as whether a particular patient’s cells respond particularly well to certain drugs. This information can then be compared to patients’ genetic and gene expression data. For instance, we could find out that a specific patient has a mutation that makes the cancer more susceptible for a certain drug, causing them to respond better to that drug. On the other hand, non-mutational information, such as how genes are expressed, how signal pathways are activated, or how epigenetic changes arise, may help us better understand how cells respond to different drugs. These different types of data can then be used at the individual level but also to divide patients into different subgroups to find more suitable treatment approaches.”
The patient’s blood samples or cerebrospinal fluid samples can be used for fluid biopsies and used to inspect how the tumour’s DNA is expressed. This will show how well the patient is responding to the drug, or if the cancer has recurred.

The iCAN research project, which is funded by Academy of Finland, covers most currently known types of cancer. Several research groups who concentrate on different types of cancers and research groups working on improving the relevant research methods are involved in the project. Information on the cancer is compared with the patient’s other health data in the secure HUS Acamedics environment.
“All the data we upload on Acamedics is available to all researchers participating in the iCAN project. We have a wealth of material that we can compare our findings against. This allows us to identify, for example, patient group and patient specific genetic markers in the genetic and other omics data.
All data, which includes data types such as drug testing data, genetic data and transcriptomics data, are combined using a powerful tool called an Integrated Molecular Tumour Board system (iMTB). In their research project on children with cancer, Pietiäinen and her colleagues are also evaluating how doctors can quickly make use of the results of recent or ongoing research.
“We aim to report clinically relevant findings to the doctors, thereby hopefully helping to choose a better treatment approach if they have ran out of recommended approaches.”

The iCAN utilises the SD Connect service provided by the Finnish ELIXIR node CSC for transferring sequencing data to the Academics environment.
The data is encrypted using Crypt4GH, a protected standard encryption method developed by the Global Alliance for Genomics & Health for sharing human genetic information.
“This ensures that the information can be used in all of the services included in CSC’s SD service suite, and may even be potentially shared with other service providers who possess similar information.”
The sheer magnitude of the iCAN project is illustrated by the fact that the accumulated data is expected to reach three petabytes by 2026.
“All of this data makes it possible for us to understand the molecular makeup of different types of cancer and patients’ drug responses.”
Ari Turunen
29.9.2023
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2023). Artificial intelligence helps researchers find suitable drugs based on patient’s genetic data and cancer cell samples. https://doi.org/10.5281/zenodo.10796468
Institute for Molecular Medicine Finland (FIMM)
FIMM is part of HiLIFE Helsinki Institute of Life Science -research center.
https://www.helsinki.fi/en/hilife-helsinki-institute-life-science/units/fimm
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The Bioinformatics Center at the University of Eastern Finland (UEF), led by Virpi Ahola, is developing new applications for analysing biomedical and multimodal data. These can be used to study cancers, metabolics, cardiovascular and neurodegenerative diseases.
Ahola has had a long career in bioinformatics. She was part of professor Ilkka Hanski’s metapopulation biology research group, which sequenced the whole genome of the Glanville fritillary butterfly that was the first reference genome solved in Finland. At the Karolinska Institute in Hong Kong, Ahola analysed gene function in different diseases at the single-cell level, and thereby studied how stem cells can be used to develop new drugs and treatments. She now heads the UEF Bioinformatics Center.
The Bioinformatics Center integrates different types of omics data (genomics, proteomics, transcriptomics) with clinical data and, in the future, possibly also imaging data.
“In addition to the usual omics analyses, we carry out multimodal data analysis for different research groups. This entails combining the analysis of different types of data in order to provide more information than if they are analysed separately.”
Analysis of the multimodal data varies depending on whether the data originates from different patients.
Omics is a research method that aims to analyse all genetically determined variables of a research subject simultaneously. Genomics analyses genetic variation and the function of genes, proteomics focuses on proteins, and epigenetics on the regulation of gene function and the storage of hereditable information without changes in the DNA sequence. Metabolomics, for its part, analyses changes in metabolism caused by disease, diet or medication.
“We are developing bioinformatics services in collaboration with biomedical experts. One focus at the University of Eastern Finland is on understanding the molecular basis of key chronic diseases and improving their prevention and treatment,” Ahola says.
Translational medicine uses basic research in clinical trials, but also patient samples and disease models to identify disease mechanisms and drug targets. The research approach is interdisciplinary, which provides a good starting point for research but also improves treatments for patients.
“What is delaying the era of translational medicine is that we simply don’t know enough. The idea behind combining several different data sources is to obtain more information. The integration is very much computational, and requires CSC – IT Center for Science’s resources and infrastructures like ELIXIR.”
One example Ahola gives is single-cell technologies.
In transcription, the genetic code in DNA is copied into RNA. This is the first phase of protein synthesis. Transcriptomics provides precise information about the gene expression in an individual cell at a given moment.
“The use of single-cell transcriptomics is still expensive. The principles of open science exist, and therefore all data must be shared when it is published. This allows data to be reused, and different data sources to be combined.”
However, the challenge is that data is produced using different technologies.
“Different data sources may have different numbers of cells or different cell types. Which methods should be used to combine the different data? If this could be solved, we could analyse more effectively cell development and specialisation.”
Ahola’s aim is to provide more assistance in the use of computational methods.
The University of Eastern Finland’s Bioinformatics Center provides researchers computing capacity and helps researchers in data pre-processing and analysis, and also assists in the use and installations of different computational methods and software.
“If there are no bioinformaticians in the same team or collaborative teams, researchers are expected to be proficient in computational methods and processing big data.”
Ahola admits that the requirements are tough, for example for postgraduate students. Fortunately, the University of Eastern Finland has taken up this challenge by providing a Computational Biomedicine as an orientation option.
“One example of the reuse of data is Finnish biobanks, which contain the genomes of over half a million Finns. It’s not a simple matter to analyse the biobank data, because the amount of data is insane.”
Ahola is referring to the FinnGen research project, which was launched in autumn 2017. Its main goal is to increase understanding of the causes of diseases and promote their diagnosis, prevention and the development of treatment methods. FinnGen uses samples collected by all Finnish biobanks. By June 2023, more than 553,000 samples were collected for the FinnGen survey. The first phase of the research project lasted six years. There are only a few research projects of this scale in the world.
The research projects can combine genomic data with data from national health registries. Indeed, Finland has exceptionally good resources to carry out genetic research covering the entire population.
Clinical data from longitudinal studies combined with genetic data offers many opportunities. But there must be a lot of data.
“Data collections are needed because no single researcher can collect data from 10,000 or 100,000 individuals. If the dataset is smaller, it may not provide reliable information for studying genetically complex diseases.”
There are many research projects using different data sources underway at the University of Eastern Finland. A project on Alzheimer’s disease at the University of Eastern Finland and Kuopio University Hospital will combine clinical data collected from patient visits with FinnGen data. In this way, researchers are aiming to understand the biological mechanisms leading to the onset of Alzheimer’s disease.
“FinnGen’s biobank is a unique resource that could be used much more in research,” Ahola says.
“Another example of research on Alzheimer’s disease is a project with Rappta Therapeutics and UEF professors Mikko Hiltunen and Annakaisa Haapasalo. This project uses transgenic cell lines to study the effect of different Alzheimer’s treatments on protein function.”
One interesting collaboration project is underway with Academy of Finland researcher Kirsi Ketola.
“The study investigates carboplatin treatment resistance mechanisms in prostate cancer. Carboplatin produces DNA cross-links, which lead to activation of a mechanism that repairs DNA and causes resistance, allowing cancer cells to divide again. The research uses single-cell techniques to measure both gene expression and chromatin changes at the single-cell level.”
Chromosomes are located in the nucleus, in the form of long chromatin strands.
According to Ahola, careful data integration and analysis could promote development of personalised treatment.

Ahola is a tireless advocate for the openness and reuse of data, and for the development of methods and infrastructures that facilitate and encourage this. She cites the European Genome-phenome Archive (EGA) as an example. This is a data archive that makes it possible to share and, with permission, access biomedical data that has already been published.
“The archive contains human genomic data, combined with clinical and other metadata. Since in principle it may be possible to identify a person by genome and phenotype, data sharing is strictly regulated.”
According to Ahola, the EGA allows for data sharing in the appropriate way. This makes it possible to reuse valuable biomedical research data, for instance for creating or testing new research hypotheses.
“Existing data can be approached from a different perspective – for example, patients can be selected using different criteria than in an already published study, or data can be used as part of a larger data set.”
Referring to Biocenter Finland, Ahola says that more should be done together. The centre brings together seven biocentres from different Finnish universities. It should not be impossible to increase collaboration between different biocentres and internationally, for example through the Finnish ELIXIR node CSC.
“ELIXIR is an avenue for us to network and learn from the experiences of other bioinformatics core facilities, and to be part of discussions where research infrastructure issues are brought up and new initiatives are taken.”
Because new technologies produce large and complex data sets, research infrastructures should also include data science experts, not research equipment alone.
“To make effective use of data, the computing capacity offered by CSC, for example, is not enough. Data processing and reuse also requires staff with expertise in the field. As I see it, better resourcing and systematic collaboration between biocentres could substantially facilitate and improve the processing, integration and reuse of large omics data sets.”
Ari Turunen
1.9.2023
Read article in PDF
Citation
Turunen, A., & Nyrönen, T. (2024). Improving breast cancer treatment prognoses with liquid biopsy. https://doi.org/10.5281/zenodo.13691344
More information:
Bioinformatics Center, University of Eastern Finland
https://uefconnect.uef.fi/en/group/bioinformatics-center/
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Professor Merja Heinäniemi and her research team at the University of Eastern Finland use computational methods to interpret cancer samples in order to determine which cellular processes are defective and how cells behave during drug treatments. The aim is to find effective medicinal treatments for leukaemia.
Leukaemia, or blood cancer, occurs when the precursors of white blood cells in the bone marrow turn into cancerous cells. Unlike other cancers, leukaemia is not a single tumour but instead consists of cancer cells in the bloodstream and bone marrow. In children, leukaemia treatments are so effective that the prognosis for recovery is as high as 90 per cent. Heinäniemi points out that the disease may recur, however.
“Even if leukaemia is treatable after a relapse, it still means that chemotherapy, a long course of cytostatic drugs, takes several years. Therefore, more effective treatments are important, and for some patients the treatments could be reduced. There are patients for whom it is difficult to find treatments, but on the other hand, treatments for childhood leukaemias last long.”
A professor of computational biomedicine, Heinäniemi studies how defects in gene regulations influence the development of cancer cells.
“Blood cancers include various types of leukaemias, acute and chronic leukaemias. Of these, myeloid leukaemias are mainly adult diseases, whereas lymphoblastic leukaemias are children’s diseases,” Heinäniemi explains.
In acute leukaemia, the blood stem cell genome in the bone marrow is altered and white blood cell precursors begin to divide uncontrollably. In children, the commonest form of leukaemia develops from a precursor of B- and T-cells of the immune defence system, and is called acute lymphoblastic leukaemia.
Heinäniemi’s research group has carried out gene expression profiling analyses related to cancer. Gene expression refers to a sequence of events in the cell in which the code in DNA is copied into RNA and then into a protein under the control of this messenger molecule. The gene that promotes cancer development can be activated or inactivated. Regulatory regions that affect gene function also play a role in the development of DNA damage in childhood leukaemia and more mature cancer of the lymphoid tissue.
In 2019, Heinäniemi and other researchers collected a large dataset of more than 10,000 patient samples. This data on haematological malignancies (HEMAP) will continue to be shared with researchers through the Finnish ELIXIR node, CSC – IT Center for Science. They were able to deduce more than 30 types of cancer from this dataset by computational methods. In addition, new disease biomarkers and new drug candidates were discovered when the data was combined with drug target databases.
For example, subtypes of childhood leukaemia were found that behave differently at the molecular level.
“We can already see from the data clustering that even subtypes of the disease have unique genetic profiles and can be identified from the data. The combined data revealed to us, at the molecular level, the heterogeneity of the disease, and the similarities between different diseases.”
Heinäniemi has used the data to look for patients for whom treatment could be less rigorous. Cytostatic drugs are medicines used to destroy cancer cells. However, they can cause many side effects. Patients with a poor treatment response also tend to have an increased risk of relapse.

Together with Olli Lohi from Tampere University, Heinäniemi’s team mapped the entire human genome to see how different genes could act as predictors of childhood leukaemia.
“Potential biomarker genes for childhood leukaemias were identified in the HEMAP data. We initially set out to identify poor drug response, but in the follow-up genomic study we discovered the traits of leukaemia patients who respond well to drug treatments.”
One such sign of a good response is cells in the cell cycle. Cell life usually follows a rhythm – the cell cycle – in which cell division, i.e. mitosis, alternates with an intermediate phase, or interphase. The goal of the cell cycle is usually to produce two identical cells by cell division.
“By mapping individual cells, we determined the number of cells in the cycle. We were able to separate the different cells into phases, and the number of cells in the cycle seemed to be an important marker of a good response. Single-cell analysis is a good way to study how a cancer cell behaves. From there, the rarer surviving cells are revealed among the rest of the mass. It is important to study how drug treatment affects the cancer cell behaviour,” Heinäniemi says.
Single-cell RNA sequencing (scRNA-seq) measures the activities of all genes in each cell separately, giving a more accurate view of cellular differences. This is important information, because cancer cells try to escape from immune cells by mutating.
“In cancer research, it is important to obtain data at the level of a single cell. The fact is that cancer cells change all the time, meaning that each cell begins to be different.”
Now, single-cell technology for studying leukaemia can measure the profile of up to 10,000 cells in a single bone marrow sample. As single-cell technology becomes more common in cancer research, it will be easy to measure even millions of cells.
Heinäniemi’s group in the research project found that leukaemia treatment rapidly triggers different kinds of escape pathways in cells by altering gene reading. This enables the cancer cell to evade the treatment given. The RNA molecular profiles that are read affect the construction of the functional part of the cell. In a way, this indicates the current state of the cell and what it is trying to do.
“A leukaemia cell is a type of bone marrow stem cell that still has the potential to change its phenotype. It can take on different phenotypes to try to hide from the treatment. For example, making the cell not divide so wildly is one escape route. During initial treatments, the cell can also switch between differentiation states and find a drug-resistant state.”
In Heinäniemi’s group, a broad molecular profile measured from cells can be grouped using computational methods. This makes it possible to distinguish normal bone marrow cells from leukaemia cell profiles, and to identify different leukaemia cell phenotypes based on measurements. Models can also be trained to learn the relationship between different measurements collected from the same sample.
“The diagnostic stage does not fully reveal what kind of escape mechanisms those cancer cells may have. This is where single-cell technology has helped, because now we can measure those rare, resistant cell types during treatment. So we get new information, and we can then think about how we can prevent the cell from escaping, or find out where it is hiding.”
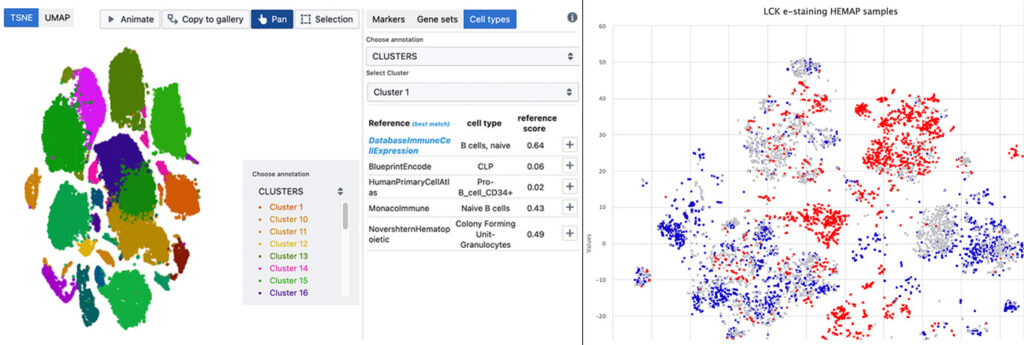
Now Heinäniemi’s team has started to use the neural network models they have developed. Data is collected from different studies for this model.
“Our leukaemia project focuses on childhood cancers. It’s a very rare type of cancer: if we don’t get the data combined, the data sets will be very small. The aim is to use CSC’s infrastructure for these projects. This would allow the data to be processed and made available to the scientific community.”
It is not possible to conduct a study using Finnish data only.
“We have a long history of Nordic collaboration. Now other European countries have also joined in. The aim would be to enable data provision for researchers who cannot process the data themselves. We bring together profiles that have already been collected and measured, because collecting them from public databases is laborious. The aim is to bring the single-cell data together in one place and in an accessible format. When we can bring the results of different research groups around the world together in one place, we can quickly compare which drug candidates might work.”
Sharing data requires building trust. In practice, this means working with the patients involved in the projects.
“It’s really important that they are involved and that their data is stored securely, and that researchers are able to do their work with the data. This is what CSC is enabling at national and EU level through its involvement in the ELIXIR infrastructure.”
Ari Turunen
15.8.2023
Read article in PDF
Citation
Nyrönen, T., & Turunen, A. (2023). Better treatments for leukaemia. https://doi.org/10.5281/zenodo.10020637
More information:
Bioinformatics Center
University of Eastern Finland
https://uefconnect.uef.fi/en/group/bioinformatics-center/
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Pölönen P, Mehtonen J, Lin J, Liuksiala T, Häyrynen S, Teppo S, Mäkinen A, Kumar A, Malani D, Pohjolainen V, Porkka K, Heckman CA, May P, Hautamäki V, Granberg KJ, Lohi O, Nykter M, Heinäniemi M.Cancer Res. 2019 May 15;79(10):2466-2479. doi: 10.1158/0008-5472.CAN-18-2970. Epub 2019 Apr 2.PMID: 30940663
Data-driven characterization of molecular phenotypes across heterogeneous sample collections.
Mehtonen J, Pölönen P, Häyrynen S, Dufva O, Lin J, Liuksiala T, Granberg K, Lohi O, Hautamäki V, Nykter M, Heinäniemi M.Nucleic Acids Res. 2019 Jul 26;47(13):e76. doi: 10.1093/nar/gkz281.PMID: 31329928
Immunogenomic Landscape of Hematological Malignancies.
Dufva O, Pölönen P, Brück O, Keränen MAI, Klievink J, Mehtonen J, Huuhtanen J, Kumar A, Malani D, Siitonen S, Kankainen M, Ghimire B, Lahtela J, Mattila P, Vähä-Koskela M, Wennerberg K, Granberg K, Leivonen SK, Meriranta L, Heckman C, Leppä S, Nykter M, Lohi O, Heinäniemi M, Mustjoki S.Cancer Cell. 2020 Sep 14;38(3):380-399.e13. doi: 10.1016/j.ccell.2020.06.002. Epub 2020 Jul 9.PMID: 32649887
Laukkanen S, Veloso A, Yan C, Oksa L, Alpert EJ, Do D, Hyvärinen N, McCarthy K, Adhikari A, Yang Q, Iyer S, Garcia SP, Pello A, Ruokoranta T, Moisio S, Adhikari S, Yoder JA, Gallagher K, Whelton L, Allen JR, Jin AH, Loontiens S, Heinäniemi M, Kelliher M, Heckman CA, Lohi O, Langenau DM.Blood. 2022 Oct 27;140(17):1891-1906. doi: 10.1182/blood.2021015106.PMID: 35544598
Kuusanmäki H, Dufva O, Vähä-Koskela M, Leppä AM, Huuhtanen J, Vänttinen I, Nygren P, Klievink J, Bouhlal J, Pölönen P, Zhang Q, Adnan-Awad S, Mancebo-Pérez C, Saad J, Miettinen J, Javarappa KK, Aakko S, Ruokoranta T, Eldfors S, Heinäniemi M, Theilgaard-Mönch K, Wartiovaara-Kautto U, Keränen M, Porkka K, Konopleva M, Wennerberg K, Kontro M, Heckman CA, Mustjoki S.Blood. 2023 Mar 30;141(13):1610-1625. doi: 10.1182/blood.2021011094.PMID: 36508699
Mehtonen J, Teppo S, Lahnalampi M, Kokko A, Kaukonen R, Oksa L, Bouvy-Liivrand M, Malyukova A, Mäkinen A, Laukkanen S, Mäkinen PI, Rounioja S, Ruusuvuori P, Sangfelt O, Lund R, Lönnberg T, Lohi O, Heinäniemi M.Genome Med. 2020 Nov 20;12(1):99. doi: 10.1186/s13073-020-00799-2.PMID: 33218352
Semisupervised Generative Autoencoder for Single-Cell Data.
Trong TN, Mehtonen J, González G, Kramer R, Hautamäki V, Heinäniemi M.J Comput Biol. 2020 Aug;27(8):1190-1203. doi: 10.1089/cmb.2019.0337. Epub 2019 Dec 2.PMID: 31794242
Sequential drug treatment targeting cell cycle and cell fate regulatory programs blocks non-genetic cancer evolution in acute lymphoblastic leukemia. Malyokova A, Lahnalampi M, Falqués-Costa T, Pölönen P, Sipola M, Mehtonen J, Teppo S, Viiliainen J, Lohi O, Hagström-Andersson AK, Heinäniemi M*, Sangfelt O* co-senior, BioRxiv https://www.biorxiv.org/content/10.1101/2023.03.27.534308v2

MicroRNAs are short RNA strands, of which more than 2,300 have been identified in humans. Their abnormal expression contributes to the development of many diseases, such as cardiovascular and immunological diseases and cancer. Researchers at the University of Turku discovered a microRNA that may be an early indicator of the risk of developing type 1 diabetes.
Professor Laura Elo and her research group for computational biomedicine at the University of Turku are developing tools for the diagnosis and treatment of complex diseases, such as diabetes, cancer and rheumatoid arthritis. The group screens patient data using computational methods to find signs of diseases and their risk factors.
Elo, Research Director at the Turku Bioscience Centre, is mining patient data for various biomarkers that may help predict the onset of diseases or tell something about the response to treatment. A biomarker is a feature that indicates a change in a biological state, in genes or proteins, for example.
The onset mechanisms of type 1 diabetes have been investigated for a long time in Finland. Type 1 diabetes is caused by the destruction of cells that produce insulin. The pancreas does not produce the insulin hormone needed by the body, causing the blood sugar level to rise.
“We aim to predict as early as possible which children will get type 1 diabetes.
Finland is the ideal country for this type of study, because the country’s type 1 diabetes incidence is the highest per capita in the world.”
Both genetic and environmental factors play a role, and Elo’s group is seeking biomarkers from diabetics in order to learn something about the development of the disease.
Data is obtained from a variety of sources. One key data set consists of follow-up studies of children. It was already in 1994 that Finland began an ambitious and extensive research project called “Diabetes, Prediction and Prevention” (DIPP) to predict and prevent diabetes. Blood samples collected in the project are studied to discover factors contributing to type 1 diabetes. Children with a genetic risk of developing diabetes are invited to a follow-up study.
“With the parents’ consent, the children are monitored since their infancy until they either get diabetes or turn 15.”
Samples are taken every 3 months, and from the age of 2 onwards, every 6 or 12 months. The university hospitals of Turku, Tampere and Oulu are taking part in this screening.

Samples have also been taken from children who undergo seroconversion at some point. Seroconversion means that autoantibodies begin to appear in the blood. Some of these children develop the disease. The follow-up study includes children with a genetic risk of falling ill.
“The majority of these children never fall ill nor develop autoantibodies. Our goal is to predict as early as possible which children will develop the disease. This is why we study both those who fall ill later and those who remain healthy throughout the follow-up period.
At some point, some children develop autoantibodies, indicating that the body is attacking itself, resulting in the destruction of the beta cells in the pancreas. This can be measured in the follow-up samples,” says Elo, but reminds that a large percentage of the children monitored never fall ill nor develop autoantibodies.
The method is to compare the samples of children who have fallen ill to samples from healthy children with as many similarities as possible. The method uses blood samples, and the idea is that blood also reflects disease processes in other parts of the body. In the case of diabetes, for example, it is difficult to get samples from the pancreas.
Thanks to this comparison, Elo’s research group found a promising biomarker, a specific microRNA.

“MicroRNAs are very short RNA strands that can be considered epigenetic regulation – they regulate the operation of cells without coding the proteins. MicroRNAs can be identified in the blood.”
MicroRNAs have been linked to various diseases, such as diabetes. When comparing various samples, the study discovered a microRNA (6868-3) that seems rather promising.
“We compared the different sample groups to find microRNA associated with falling ill and not falling ill during the follow-up period. In this case, one microRNA clearly appeared to be linked with falling ill.”
This result was studied in more detail in laboratory tests.
“We were able to identify this marker in our material at a very early stage and, in fact, predict earlier than with the currently used markers who will eventually fall ill and who will not.”

Laura Elo emphasises that the computational methods her group have developed are also suitable for the study of diseases other than diabetes.
“We have also analysed, for example, protein levels in various autoimmune diseases and cancers. A diagnosis is not usually made until clinical symptoms have appeared. In the development of the computational methods, we are motivated by the fact that with the aid of long-term follow-up measurements, we can find very early markers for diseases.”
Elo says they have begun to realise even better that it is not worth taking just a single measurement.
“Follow-up studies create, over time, a kind of reference of the person, enabling us to follow changes in the body, and to learn more about disease processes. The marker may be a molecule that is associated with a disease. MicroRNA is one example of such marker.”
According to Elo, future study of diseases should make use of the various ‘omics’, such as genomics (DNA), proteomics (proteins), transcriptomics (RNA) or metabolomics (metabolism). Elo’s group has been using the computing resources of Finland’s ELIXIR Center CSC to process the extensive data.
”We recently published a new longitudinal modelling method in the Nature Communications journal.
The goal with our method is to discover as reliable markers as possible in longitudinal materials, and the focus was on protein measurements. One of the key questions was how to reliably analyse noisy data. We compared previously-used methods and got good results in both simulated and real data. We are now able to more reliably find proteins associated with diseases, for example.”
Going to the laboratory to confirm findings is a long and expensive process, which is why it is important to find reliable changes and markers.
Ari Turunen
10.6.2023
Read article in PDF
Citation
Nyrönen, T., & Turunen, A. (2023). MicroRNAs may reveal type 1 diabetes. https://doi.org/10.5281/zenodo.10017409
More information:
Turku Bioscience Centre
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The revolutionary single-cell RNA sequencing (scRNA-seq) measures the activity of all genes separately in each cell, giving a more accurate picture of how cells differ from each other. This technology will produce a vast amount of information on diseases, such as cancer. There are millions of cancer cells in a tumour. Data analysis requires increasingly more computing capacity and efficient algorithms as the number of cells and samples analysed increases.
RNA transports the manufacturing instructions in the cell from the DNA to the proteins. RNA contributes to gene expression, that is, the production of a protein that corresponds to the gene. Single-cell technology enables the measurement of RNA levels in all genes of each individual cell.
A single tissue sample of a patient can with modern technology be used to profile thousands, sometimes even tens of thousands of cells. We can reliably profile which type of cells it contains.
“Even if the cells may look identical under the microscope, their function may turn out to be quite different once we are able to view gene expression with single-cell accuracy,” says researcher Päivi Saavalainen.
Saavalainen works at the Folkhälsan Research Center and she is also the CEO of a company called SCellex that specialises in single-cell technology. Saavalainen, who together with the Finland ELIXIR Node of CSC has organised single-cell analytics courses for researchers, considers single-cell technology one of the most revolutionary methods in biosciences over the last few years.
“In cancer research, for example, it is important to obtain information about a single cell. The fact is that cancer cells change all the time, meaning that each cell begins to be different. There are also gene mutations, as a result of which certain genes are activated and others deactivated.”
Saavalainen considers single-cell resolution important also in the study of healthy tissues as we can find out what kind of cell types are found.
“Single-cell resolution has created a huge amount of new information for basic research alone. It was thought for a long time that humans have about 200 types of cell, but single-cell analyses have already identified more than 500.”
Thanks to the new technology developed by SCellex, it is possible to determine from the tissue structure what type of cancer cells the tumour contains and whether a mutation, for example, has only affected a certain part of the tumour.
“Now we are able to find out accurately whether, for example, a tumour contains some cancer cells that are drug-resistant and what they are actually like,” says Saavalainen.
According to Saavalainen, immunotherapies for cancer have also developed dramatically. This works by helping the body’s own immune cells, T-cells, to identify and kill cancer cells. A T-cell is one of the two types of lymphocyte, along with the B-cell. They identify foreign structures and help kill cells infected by a virus and also cancer cells in which mutations have changed their own genome and consequently the proteins.
“Cancer cells are trying to escape from T-cells. They keep their changed structures hidden and secrete cytokines that silence T-cells. The goal with drug treatment is to allow T-cells to penetrate tissue, identify cancer cells aggressively and kill them. Now we are able to find out, for example, what a cancer cell next to a T-cell is doing. Is it creating some gene product that silences the T-cell, and how is the T-cell reacting to that?”
According to Saavalainen, the best-case scenario is that we understand the cancer cell types of each patient and find an effective drug to which the patient responds well. This mean we may be able to find means for individual treatments.

Single-cell analytics is generally performed by gently separating the cells from the tissue and transferred individually into a solution, followed by sequencing of the RNA contained in them. However, the problem with this approach is that the original location of the cells and their order in the tissue will be lost, meaning we do not know which cells were originally next to each other. Thanks to modern spatial techniques, the cells no longer have to be separated in individual solutions; you simply slice layers only one cell in thickness from the tissue, with RNA obtained directly from such layers. As the RNA is sequenced, we know which cell and which part of the tissue the RNA originated from.
“This means we can sequence tissue and still know their location and maintain their original order. Spatial sequencing is one of the hottest things at the moment,” says Saavalainen.
SCellex is developing a patented technology to determine the location of cells with machine learning models and microscopic colour beads. The beads are placed into a 160,000 picowell chip array platform and their random combinations create visual coordinates for the wells that can be calculated from the microscopic images by means of an AI model. The synthetic DNA codes attached to the microbeads are combined with the RNA molecules released from the tissue slices on the chip array platform, thereby linked to the well coordinates.

“We are using an AI model that can compute automatically which beads are in each well. In other words, AI creates a map. After this the actual tissue section can be connected to the chip, after which the synthetic DNA strands glued to the colour beads are attached to the RNA from the tissue. The RNA molecules are attached to these strands and identification is made.
“When RNA is sequenced in large batches, the data can be analysed to determine which colour bead combination the RNA matches, and then compare it with the original microscopic image and the AI computation. This way we can arrange the RNA data in their proper places.”
The datasets are huge and the services of Finland ELIXIR Node of CSC can help to perform the computation. Saavalainen says that the AI models are vital.
“If a sample contains tens of thousands of cells and all of them are subject to tens of thousands of gene measurement results, then not only the data on the microbeads but also the actual biological RNA data is immensely complex. You need AI to analyse it. AI can find such new information that simply would not be possible with traditional analysis tools. I think the computing power of CSC is sufficient even to solve our challenging AI models.”
Saavalainen says that the single-cell method is not yet mature enough to be used for diagnostics or prescription of drug treatments. However, at the moment it is a good tool for research purposes.
Ari Turunen
16.5.2023
Read article in PDF
Citation
Ari Turunen, Päivi Saavalainen, & Tommi Nyrönen. (2023). Single-cell RNA sequencing enabling individual disease treatment. https://doi.org/10.5281/zenodo.8181234
More information:
SCellex
Folkhälsan
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Celiac disease is a condition in which gluten in cereal-containing food products containing rye, wheat and barley cause inflammation and damage to the small intestinal mucosa. Because of the damage, nutrients are not properly absorbed. RNA sequencing of patient tissue samples and cells makes it possible to study the immune system, and whether it’s functioning properly.
Celiac disease is considered an autoimmune disease, in which the body’s defence system mistakenly targets its own tissues. Although the exact causal mechanism of celiac disease is not yet known, there is a hereditary predisposition to its onset.
Immunologist Helka Kaunisto from the Celiac Disease Research Center at Tampere University studies dermatitis herpetiformis, the skin manifestation of celiac disease.
“I have always been interested in autoimmune diseases. People who get them are at risk of developing other autoimmune diseases.”
Dermatitis herpetiformis is a common extra-intestinal manifestation of celiac disease, causing an itchy rash with small blisters due to intake of wheat, rye and barley -derived gluten. Both celiac disease and dermatitis herpetiformis are associated with a strong hereditary predisposition.

“Half of my thesis work is RNA sequencing. We are not looking for a specific gene or protein in the sequencing, but want to investigate what types of changes gluten intake causes in the RNA profile of people with dermatitis herpetiformis.”
According to Kaunisto, it is possible that gluten intake leads to altered expression of certain RNA molecules in people with dermatitis herpetiformis. It shows the effect of gluten on the immune system, for example on cellular metabolism or inflammation.
At the same time, it is possible to explore how in celiac disease the immune response can spread from a local reaction in the gut to a systemic reaction that spreads to the skin or other organs. In other words, the disease is complex, and this complexity includes immunological abnormalities.
Kaunisto studies immune cells and the immune response to find out why some people with celiac disease develop dermatitis herpetiformis.
“It should be remembered that the gut and the skin have different layers that function in different immunological ways. Dermatitis herpetiformis is a really good target for studying the extra-intestinal symptoms of celiac disease. This information can then also be used to investigate other autoimmune diseases. For example, how can rheumatism start in one place and then spread elsewhere and become systemic?”
Around 10 per cent of people with celiac disease have dermatitis herpetiformis. Celiac disease and dermatitis herpetiformis can be investigated by measuring antibodies in the blood. In people with celiac disease and those with dermatitis herpetiformis, gluten triggers the formation of tissue antibodies.

“Celiac disease is known as an intestinal disease, but it has many other symptoms that are not related to the intestines at all. There may be neurological and skin problems too. Is there a difference in immunity between people with celiac disease and people with dermatitis herpetiformis? How can an immune response spread from the gut to the skin – and why does a rash develop?”
The diagnosis of celiac disease exploits the analysis of antibody levels. Transglutaminases are enzymes that bind proteins together in tissues. High levels of antibodies to transglutaminase 2 (S-tGAbA) suggest the person has celiac disease. Transglutaminase 2 modifies the structure of gluten in the body. This causes the lining of the small intestine to become inflamed and damaged.
The Celiac Disease Research Center in Tampere has conducted a study in which patients with dermatitis herpetiformis who were on a gluten-free diet were exposed to gluten for a short period. Before and during exposure, samples were taken from the small intestine and blood. These samples are studied to find out how gluten affects the expression of RNA in blood cells and the small intestine.
“Although some people with celiac disease have the same serum-based antibodies as people with dermatitis herpetiformis, not everyone will get dermatitis herpetiformis “, says Kaunisto.
“Patients with dermatitis herpetiformis have antibodies to transglutaminase 2, but they also have antibodies to a related enzyme, transglutaminase 3. Transglutaminase 3 (TG3) antibodies are also found in the skin, close to the rash, and are thought to be involved in its development. TG3 antibodies are also found in the bloodstream of patients with dermatitis herpetiformis. Although some people with celiac disease also have antibodies to transglutaminase 3 in the bloodstream, not all people with celiac disease develop dermatitis herpetiformis. Why this is so is what we want to solve.”

According to Kaunisto, the research is of great benefit to clinical science.
“For example, if celiac disease is not well managed – the person doesn’t stick to a strict gluten-free diet, say – is there a greater chance of developing extra-intestinal symptoms?”

The study involves analysis of sensitive data obtained from patient’s tissue samples. Patients consented to participate in the study. As this is information subject to the European Union’s General Data Protection Regulation (GDPR), the data will be processed and used in CSC’s sensitive data services (SD Desktop and SD Connect).
The sequencing has been done in collaboration with the University of Helsinki, and the data is stored encrypted in SD Connect and analysed with SD Desktop.
Kaunisto did not have previous experience of the high-performance computing or sensitive data services.
“I started using the Sensitive Data Services because I needed more computational power than what was available for me through the university, and I needed a secure environment for this computational power. I find the services very easy to use as the online guides are very thorough. If I have a problem I can’t solve myself the helpdesk is always very helpful.”
To avoid the long-term effects of gluten, treatment can begin as soon as possible when celiac disease is detected early. However, since gluten is present in many foods, gluten-free diets are difficult to maintain.
“At the moment, the only treatment is a strict gluten-free diet, but there is also a lot of research underway into pharmaceuticals as possible future treatments. The Celiac Disease Research Center also collaborates extensively with companies developing new potential drugs. Drugs may be able to prevent intestinal and other damage in patients in the future, but initially they will likely be adjunctive therapies alongside gluten-free dietery treatment. A preliminary study carried out in Tampere University found that the experimental-phase drug ZED1227 inhibited transglutaminase 2 activity, and its use reduced gluten-induced intestinal damage in patients.”
Ari Turunen
12.4.2023
Read article in PDF
Citation
Ari Turunen, Helka Kaunisto, Tommi Nyrönen, & Francesca Morello. (2023). Tissue samples analysed with Sensitive Data (SD) services provide new information on celiac disease and other autoimmune diseases. https://doi.org/10.5281/zenodo.8154655
Read article:
More information:
Celiac Disease Research Center, University of Tampere
https://www.tuni.fi/en/research/celiac-disease-research-center
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Geneticist Petri Auvinen and his research team are using DNA samples to find out what has been happening in the Baltic ecosystem during the past 10,000 years.
Samples obtained from the seabed by drilling can be used to study past and present species and their habitats. This is useful in the study of biodiversity and climate change. Samples are obtained from the sediments –that is, layered soil that has been moved by water, wind or ice. If DNA can be isolated from the sediment samples, it can be used to learn about organisms located at certain depth of the sediment.
“Our aim is to collect sediments samples as far down as possible in the Baltic Sea bed in order to study the history of the Baltic Sea basin. We also take samples deep in marshland, providing us with information about the history of the soil,” says Auvinen.
Sediments are not created everywhere, but they can be found in the Baltic Sea and on marshland.
“These sediments have never been studied as extensively as we are doing now. At best, we may be able to reach back to samples dating from the Ice Age, which was when sediments began to accumulate in the sea.”
Petri Auvinen is a research director at the Institute of Biotechnology, University of Helsinki. His research focuses on genomics and metagenomics. While genomics looks at the entire genome of an organism, metagenomics can study and sequence a number of organisms, such as microbes, from a single sample at the same time. The study of micro-organisms has advanced in leaps and bounds. A sequence sample can be taken from any environment, soil or gut to determine the microbiota composition. The term used is microbiome, denoting the microbiota of a specific habitat and its genome, that is, the metagenome.
“Analysing the sediment, we are able to determine when it was created. We analyse the sediments to find what microbes, other organisms and plants lived in a specific period.”
Auvinen’s research group has been studying environmental samples from soil and composts for a long time. For example, by isolating DNA samples from composts, they have been able to identify thousands of species of bacteria.
Auvinen has spent a lot of time studying the genetic origin of microbes.
“We published our first microbiome study of the Baltic Sea in 2010. We were already then using next-generation sequencing methods. These methods can determine up to billions of DNA sequences at the same time from a single sample.”
As the Baltic Sea is a shallow basin mostly filled with brackish water, it suffers from eutrophication, toxic blue-green algae blooms, and oxygen deficit, all of which have an impact on the community. The research group carried out thorough sequencing to determine the structures of bacterial communities in the northern part of the Baltic.
Previously research focused on one molecule at a time, but now we are talking about a sequencing volume a million times greater. Next generation sequencing can be used to determine microbes in a sediment sample.

Microbes can provide a surprising amount of new information about climate change and biodiversity.
“It would not surprise me if we discovered from sediments that as the environment has changed, so have the microbes. It is worth keeping in mind that practically all matter used by organisms has been dissolved from sediments by microbes. This means that if microbiota undergoes major changes in the environment, it is possible that some other ecological services change as well.”
By this Auvinen refers to “services” provided by nature, such as pollination, conversion of nutrients suitable for humans, and clean water.
“If the environment changes, then forests, for example, may disappear or be damaged for long periods, meaning that these services would no longer be available. Humans can manage up to a point with technology, but at some stage life may become difficult or even impossible. On the other hand, you may look at the situation from the viewpoint that as ecological services change or are reduced, the environment can no longer support such a large number of people.”
Auvinen’s group has researchers from a range of fields. You need experts from different fields in order to arrive at an accurate analysis of bygone environments.
“Our plan is specifically to study reconstruction, that is, how DNA and RNA data can be combined with sample dating, enabling us to know the precise age of samples.”
Auvinen mentions stable isotopes that can be used for reconstructing the dating of environmental conditions. Furthermore, botanists can analyse the DNA of, say, pollen, and this can be combined with isotope dating, enabling us to see what the environment was like thousands of years ago. As sediments contain both old and new DNA, and it is not possible to tell them apart, dating is crucial.
“In order for us to know which way the environment is heading, we should know what it was like earlier. We are able to tell, going back 10,000 years, what has been happening in the environment. This can be used as reference material for what will happen in the future.”
Important areas of study include not only biodiversity loss but also excessive use of chemicals. They affect not only us but generations that come after us.
“Man-made chemicals not originating from nature will circulate in the environment. We have pharmaceuticals and detergents that may never disappear from nature. We do not know how these chemicals will affect the environment in the long run. The spread of microplastics in nature is one manifestation of excessive use of chemicals.”

Some of the study’s sediment and data samples are analysed, while some are placed in storage in order to study other aspects of them later. In addition to microbiome study, the Institute of Biotechnology conducts plenty of research on human sequence samples and other species. All of this requires a huge amount of computing power.
“Our institute alone produces 8 terabytes of sequence data on one device in a week. This is a lot more than it used to be ten years ago. When all start doing research this way, I see a big challenge ahead in data processing.”
For example, the DNA sequence that covers the entire genome is in separate parts in the cell, and these must be assembled in the proper order. Then there is annotation to do, seeking the genes and their function in the sequence.
“When assembling genomic data, you must have plenty of RAM, because all sequences must be analysed in the same memory space. We will also need more disk space for data storage. We have a great number of highly trained people that use most of their day to copy data from one place to another.”
The greatest bottleneck is the storage of sensitive data.
“The storage space we are using is too small.”
Another challenge is the software used for the calculations. Some of the data is analysed using the CSC – IT Center for Science’s ePouta hardware, but some must be done with the team’s own.
“The software can be so complex that it cannot be run on the CSC system. We use some virtual computers, but they have their limitations, too. We also have runs that may continue uninterrupted for months. CSC has thousands of users, and the CSC environment obviously has some maintenance outages. For example, when we were working on the genome of the Saimaa ringed seal, the first major assemblies took a thousand hours.”
“Currently we are able to work on large genomes a hundred times better than a few years ago. But there is more and more data all the time, and we must be able to store it efficiently and in a way that it is comparable to other data. We will continue to work with CSC in data storage, transfer and calculation.”
Ari Turunen
20.3.2023
Read article in PDF
Citation
Ari Turunen, Petri Auvinen, & Tommi Nyrönen. (2023). DNA isolated from Baltic Sea sediment shedding light on climate change and biodiversity. https://doi.org/10.5281/zenodo.8154641
More information:
Institute of Biotechnology (University of Helsinki)
https://www.helsinki.fi/en/hilife-helsinki-institute-life-science/units/institute-biotechnology
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Organoids grown from stem cells enable new ways to model a variety of diseases such as cancer. At the University of Oulu, new techniques to engineer embryonic tissue are used to find cancer genes.
Professor of Developmental Biology Seppo Vainio’s research group uses organoids to study the genes that cause kidney cancer. Stem cells can be manipulated to create organs – such as three-dimensional cell cultures resembling kidneys – that include nearly all the cell types found in real organs. Organoids can also be clusters of cancer cells grown from the cells harvested form a patient’s cancer tumour. Organoids are created with a few cells harvested from tissue or with stem cells.
“We can create models of the development of internal organs like kidneys. Our toolkit also includes methods to create, or replicate, the gene-level changes caused by different diseases found in human embryonic stem cells. This is based on gene targeting techniques,” says Vainio.
Of all kidney cancers, 90 per cent are caused by renal cell carcinoma. Factors causing the disease include smoking, obesity and genetics. In particular, the research at the University of Oulu focuses on the similarities between kidney development and cancer development. The research group studied gene expression to determine whether some of the genes involved with kidney development are also involved with cancer development. A variety of organoids were experimented with. Some were created with renal cells and cancer cells from mice and some were a combination of mouse renal cells and human cervical cancer cells.
“When we combined the embryonic renal cells and the renal cancer cells to create a single organoid, the embryonic renal cells did not create the tubular structures typical of kidneys. However, when we blocked the expression of certain genes associated with kidney growth in the cancer cells, cancer cell growth slowed down and we found that the tubular structures developed normally,” says researcher Anatoliy Samoylenko.
The research group found that deactivating certain cancer cell genes resulted in the embryo being able to produce new structures normally. The organoid model developed at the University enables a new way of examining the detrimental signals that cancer cells send to their surroundings.
Embryonic stem cells have revolutionised disease-oriented research, and stem cells can be used to create in vivo models. Organoids can be used to identify the initial stages of tumour growth, cell proliferation and specialisation, cell migration and cell death.

According to researcher Ilya Skovorodkin, organoid research is a revolutionary development.
“True science starts with experimentation. In a way, classical medicine can never be a true scientific endeavour, since you cannot use people as test subjects.”
Skovorodkin says that organoids will change this. They enable studying human diseases with experiments. They can be taken advantage of in the development of new medicines and treatment methods.
“Naturally, we are still far from being able to study all the interaction that goes on within an organism. However, we can start by studying the cell-cell interaction and the way cells signal each other.”
Organoids can be kidneys, hearts or cancers in miniature form.
“In the best-case scenario we are able to harvest cells from a patient in connection with an autopsy or from the patient’s skin, for example. The cells can be reverted to their embryonic state and then they can be used to create miniature organs. This enables us to experiment. What kind of medicine would be best for this patient? Our primary focus is on organ development and embryonic kidney development in particular. Organoids are a very impressive tool. The laboratory in Oulu was one of the first laboratories to have the capability to create a kidney organoid.”
According to Skovorodkin, the next step in biomedicine is to create an organoid with blood circulation.
“With a single organoid, we can study cell-cell interaction and how organs work. But in real life, organs are connected to the whole organism via blood circulation. Blood circulation brings cells all the materials they need, and interaction happens with cells and organs. We can already grow blood vessels of chicken embryos in organoids.”
Skovorodkin aims to model the interaction between cells and organs. Microfluidics, or the control of microscopic fluid and gas flows, can be used to create artificial blood vessels and to study blood circulation in organs.
This type of modelling would significantly benefit cancer research.
“A cancer does not grow in isolation; it is always connected to the whole organism through blood circulation.”
According to Professor Vainio, the aim is that organoids could be used with 3D bioprinting techniques to find new cell and tissue therapies.
“Organs that will not be rejected are greatly needed for transplantation. Successful attempts at this have already been made,” Vainio says.
In Finland, biobanks are responsible for the legality and information safety of human sample collection. The biobanks encode each sample, which ensures that donors stay anonymous.
“But the need to apply for licences has increased the bureaucracy of getting samples and their clinical data for use in research.”
Vainio says that the aim is to improve legislation to allow safeguarding the anonymity of private individuals in research activities such as creating human organoids and the patient records related to them. Currently, university hospitals and the Finnish Social and Health Data Permit Authority Findata are responsible for the administration of clinical test results of operational patient care.
“Researchers can reuse the data on gene-level changes associated with certain human diseases observed in stem cells and the organoids grown from them. This is basic research and produces experimental data such as image analysis data and gene-level data. CSC –IT Center for Science already provides the framework needed to store such digital material.”
According to Vainio, anonymisation is not as relevant when data on experimental cell lines is produced, which is why it would be possible to manage such data through CSC.
“If there was a need to link this data to patient records, it could be done with Findata cooperation. Creating organoids from samples donated by patients, for example, could also be made subject to licence in Finland.”
Ari Turunen
27.2.2023
Read article in PDF
Citation
Ari Turunen, Seppo Vainio, Anatoliy Samoylenko, Ilya Skovorodkin, Susanna Kaisto, & Tommi Nyrönen. (2023). Organoids grown from stem cells boost cancer research. https://doi.org/10.5281/zenodo.8154628
More information:
University of Oulu
Development Biology Laboratory
Findata
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The ELIXIR Node in Finland, CSC, is working together with the universities of Linköping and Uppsala to build a database of pathological data, consisting of a secure authorisation mechanism for receiving and storing pathological images and data that describe them. The data description also plays a key part in the authorisation process. BIGPICTURE relies on ELIXIR AAI’s technologies regarding the authorisation of imaging data. The organisations taking part in the project are committed to producing and sharing image data.
The six-year BIGPICTURE project that began in February 2021 will collect three million scanned, digital pathology slides from various European hospitals, research organisations and pharmaceutical companies.
BIGPICTURE is a European consortium, the purpose of which is to create a secure storage place and platform under European data security principles. Whole slide images and machine learning algorithms can be stored in the platform, enabling image analysis by means of artificial intelligence.
The project is participated in by 45 organisations from 15 countries. Finland’s contribution to the project is provided by Helsinki University Hospital (HUS), mainly through the Helsinki Biobank, and CSC – IT Center for Science.
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Francesca Morello works as Customer Liaison Officer for CSC’s sensitive data services. Morello and her colleagues are developing tools and services to analyse, share and publish data. CSC will also host the Finnish part of the Federated EGA (European Genome-phenome Archive), a distributed network of repositories for sharing human sensitive biomedical data.
SD Connect is a service for collecting and storing sensitive research data during the active phase of a research project while SD Desktop users can directly access and manage that data in a virtual computing environment. The services are accessible via a web user interface, from the user’s own computer.
”Once users upload their sensitive data to CSC, these data are always kept encrypted when stored, transferred or processed within our services. Decryption is done only when data is made available for authorised users within the SD Desktop service”, says Morello.
This cloud-computing environment is easy to use. Morello is very enthusiastic about the fact that accessing the workspace does not require specific technical expertise.
”Researchers can access the workspace with just a few clicks. While SD services are suitable for managing sensitive data from any research field, we are working on further facilitating the use of the services, from fully automating data encryption to streamlining the computing environment customisation.”
The services are available to researchers and students affiliated with Finnish academic organisations, research institutes, and their international collaborators. Using CSC services requires to register a CSC account. While SD Connect and SD Desktop have been designed to facilitate collaboration between organisations, the data is always stored in CSC’s cloud services in Finland.
Seppo Vainio is Professor in Developmental Biology at the University of Oulu, and his research area is organoids. An organoid is a simple and small version of an organ, grown from stem cells. The study of organoids involves the processing of plenty of sensitive data.
“Organoids can be used to model cellular and molecular changes that are either normal or abnormal in terms of organ functioning. The crucial thing is that researchers have developed methods based on the use of human cells to create cells capable of multiple tasks. These can then be used to channelled in various pathways using developmental biology signals. This means we have methods – recipes, if you like – to make cells do different hings, such as create organoids that model the normal development of a kidney. We are able to create development models of these and other organs. We also have methods to create similar genetic changes in human stem cells found in the human genome.”

According to Seppo Vainio, organoid study is a current scientific megatrend. Now we are able to model human disease processes in a new way. When organoids are combined, we can also experimentally study the interaction of tissues and organs during their formative phase in humans. Organoids are useful in studying human diseases and developing new drugs and treatments.
“We have human stem cell libraries in Europe, and in principle, we could create a stem cell storage of every human being in a biobank. Where necessary, we could then draw on this to create a personal disease model for the study of the health and disease of each person.”
Vainio says that as the goal is to have personalised health technology and data, both the stem cell and cell biobank, we have realistic means of finding out how diseases develop. However, all this requires investments in research. Vainio hopes that the Finnish biobank system could be developed in this direction.
One interesting area involves a type of stem cell called an induced pluripotent stem cell (iPSC). Human embryonic stem cells were first grown in the late 1990s, and iPSCs, which are very similar to these, were first generated in 2007. These iPSC lines can be created from, for example, a patient’s skin or blood cells, and they can be programmed to differentiate as desired.
“Because iPSCs originate in individuals, the projects also involve a large amount of processing of sensitive patient material. Our goal is to link observations in organoids better and better into patient records. In this context, the Finnish Social and Health Data Permit Authority Findata offers ways for the utilisation of Finland’s numerous register data system.”
For example, the goal with the FinnGen research project study is to gain a better understanding of disease mechanisms and the development of new treatments by combining genomic and health data. It contains the genetic data of more than 500,000 Finns. The data is returned, as agreed, into Finnish biobanks, from which it is freely available to researchers. FinnGen has identified a number of genetic variants related to diseases.
“Researchers have modelled variants identified in organoids experimentally in order to study in more detail genetic changes associated with diseases, or pathogenesis. Once this research is later connected to automated drug screening performed with various chemical libraries and biomarkers, the process will create a foundation to accelerate the development of new treatments with organoids.”
Sensitive data include human personal data, ecological data or confidential data. Processing of personal data is regulated by the European General Data Protection Regulation.
”The data controller is an organisation or a legal representative who takes all the decisions on how the data is used. With SD services, we aim at providing all the tools for researchers and their organisations to manage the data access during collection, analysis and reuse,” says Francesca Morello.
Processing health or register data for secondary use is strictly regulated by national laws. SD Desktop is a certified secondary use environment that has been audited against Findata (the Finnish Social and Health Data Permit Authority) regulations. In this case, data access and data exports are managed by Findata and by CSC’s helpdesk. According to Morello, these services are designed so that they provide researchers and data controllers all the instruments to keep their data safe but the services remain flexible and user friendly.

Sequencing, storing and processing genetic sequences is a time consuming process. As a first step, DNA sequences can be uploaded by the sequencing facility directly into researchers’ workspace in SD Connect. The encrypted data can be easily shared with other researchers, via a URL. When the data collection phase is over, researchers can spin up a virtual computer with SD Desktop and analyse the data stored in SD Connect via data streaming. They can also decide to give read only access to their collaborators from other organisations, for example, to analyse their data together.
When researchers have created their results from their genetic analyses, they can publish their data under controlled access using the Finnish Federated EGA service. In this case, the dataset will be assigned a permanent identifier and the data will be advertised internationally via EGA for reuse. The data remains in Finland while approved researchers can access the data via data streaming using the SD Desktop service.
Only one copy of the same dataset is uploaded to CSC and used during all the different stages of research. The Federated EGA (European Genome-phenome Archive) together with its fully compatible American counterpart dbGAP are the primary global resources for access of sensitive human biomedical data consented for research use.
Ari Turunen
19.12.2022
Read article in PDF
Citation
Ari Turunen, Francesca Morello, & Tommi Nyrönen. (2022). Sensitive Data (SD) services for Research: with a few clicks a researcher can launch a personal secure computing environment. https://doi.org/10.5281/zenodo.8154610
More information:
University of Oulu
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org
My CSC portal
SD Connect
SD Desktop
Federated European Genome-phenome Archive
https://research.csc.fi/-/fega

Jenni Hultman, Senior Scientist at the Natural Resources Institute Finland (Luke), is interested in microbiota in cold climates that may also have an effect on the global climate. For example, there are bacteria and archaea on the tundra that eat methane, a potent greenhouse gas.
It has been discovered that the tundra is also a major source of other greenhouse gases, such as nitrous oxide. However, the diversity of micro organisms that contribute to generating nitrous oxide, is largely unknown to science yet. Even if we found out which types of microbes are involved, it is also important to know what kind of processes their genes are actually involved.
Metagenomics means the study of microbial DNA directly in their natural living environment. The term generally refers to bacterial genomes in a sample, but it also means the genomes of other microorganisms, such as those of archaea and fungi and also the genomes of the eukaryotes inhabiting the sample of interest. Metagenomics can thus be used to study and sequence multiple organisms simultaneously from a single sample. Hultman and her colleagues sequenced millions of genomes from tundra soil microbiomes. To do this, they needed the computing power of the Finnish ELIXIR Node CSC – IT Center for Science, because the volume of data in the materials runs in terabytes.
“We have learned what kind of microbes live in the subarctic area and what they do. Processing the data material has taken up an incredible amount of computing power. We discovered several previously unknown microbiota and genomes.”
DNA sequences obtained from the samples have been analysed, aiming to identify new species and their relationships. By analysing RNA sequences, they have learned what the microbiota were doing at the time of sampling.
“What I’m particularly excited about is being able to visit the same sites in wintertime. Now we are able to learn about seasonal variation, that is, what takes place in the microbiota in the summer and winter and how warm autumns contribute to the microbial activity.”
Hultman compared the sequence fragments found in the microbiota to those available in the databases.
“Most of the time, we could not find the same sequences in the databases. More than 90 per cent of the identified genes were unknown to us.”

Microbial data found in Kilpisjärvi and Pallas in Finnish Lapland was compared to other data sources, such as findings in Alaska and Sweden. Most of the Earth’s organisms are microbes. Although they occur everywhere and in all conditions, most of them are difficult to cultivate in laboratories. New techniques are required for their study, one of them being assembly of the metagenomic sequences. The metagenome-assembled genome, or MAG, is binned from metagenomic data. In other words, using a sample that contains a number of genomes, the genome of an individual species is pieced together. This MAG data gives us new information about microbes that have not been cultured and subsequently sequenced, stored and annotated in databases.
Jenni Hultman’s study found more than 800 different MAGs, and only a small fraction of them were known previously. One interesting MAG was a species of archaea that oxidises ammonia. Ammonia-oxidising microbes play an important role in the nitrogen cycle.
“We first looked for the unknown archaea species in two datasets, from Norway and Kilpisjärvi, Finland. Once we started finding out where similar sequences could be found, we discovered them in Canada and Abisko, Sweden. Following a more detailed study, the particular archaeal genus was eventually found on both polar regions of the globe. It’s a fascinating thought that this archaea is specialised in living in polarr egions. It is important to note that this discovery was made thanks to openness of data and the availability of databases. Arctic regions may be found to have plenty of microbes that can be useful in biochemical cycles.”
These include enzymes – proteins that speed up chemical reactions – that are created by microbes.
“Microbes are efficient in creating enzymes in cold conditions. These are biotechnically interesting communities. We create large, open databases about these species, sequencing them all. We have already discovered more than a thousand genomes. Enzyme processes can be interesting, because in cold conditions it can be more economical to grow microbes that produce enzymes.”
As a case in point, Hultman mentions genes found in microbes that can break down lignin. One potential application is the replacement of fossil materials.
Other interesting targets of study are fungi and actinobacteria.
“We have found fungal families new to science – yet actually very common in our northern samples – which contain genes related to breaking up carbohydrates. Actinobacteria, known decomposers, occur in composts, for example, but there are also plenty of actinobacteria that live in cold conditions but still manage to be very active.

Methane is one of the most significant greenhouse gases. A Tampere University study in which sequencing data was analysed using CSC’s computing power, it was discovered that methane-eating bacteria, or methanotrophs, can be utilised for the manufacture of inexpensive bioproducts. Methanotrophs consume methane for their growth extremely efficiently.
Previously only individual species of microbes were studied, and the assumption was that a specific species only operates in a specific way. According to Hultman, these new findings have disproved this idea. For example, methane-eating microbes can be found among many species, not only in bacteria but also in archaea.
“Some can operate in both aerobic and anaerobic conditions. Previously it was thought that methane is created in anaerobic conditions and consumed in aerobic conditions. Deeper layers of soil contain methanogenic archaea that produce methane, while layers closer to the surface contain methanotrophs which oxidise methane and to which methane is key energy source. How to find the conditions in which these methane-eating methanotrophs thrive and how to make them multiply?”
Jenni Hultman did her thesis on composts and then moved on from such warm environments to the permafrost and Arctic areas. She is interested in how much the Arctic areas are warming and how much carbon is released into the atmosphere, but also how microbes work in carbon sequestration. Microbes play an important role in this.
“At high latitudes, the climate is warming four times as fast as previously thought. Once we learn how Arctic microbiota function, we can improve the climate-change prediction models. This data should be included in the climate models in order to better predict what the changes will cause. Metagenomics and climate sciences are actually rather closely linked. Microbes produce greenhouse gases and also consume them.”
In her work Hultman has been using the Sequence Read Archive (SRA) database, an open-access database composed of DNA sequences. They are maintained by the National Center for Biotechnology Information (NCBI), the European Bioinformatics Institute (EBI) and the DNA Data Bank of Japan (DDBJ).
“Researchers produce a huge amount of data all the time, and it is vital that we have access to databases. I hope ELIXIR can offer us up-to-date databases. When I find a new sequence fragment and I want to know what it is, I don’t have to download all new databases. Instead, they would be offered by ELIXIR. ELIXIR enables a large network that can help researchers to make open publications.
Ari Turunen
30.11.2022
Read article in PDF
Citation
Ari Turunen, Jenni Hultman, & Tommi Nyrönen. (2022). Microbiota in permafrost play an important role in climate change. https://doi.org/10.5281/zenodo.8154600
More information
Natural Resources Institute Finland (Luke)
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Proper data management enables high-quality research. Data management is now guided by the FAIR principles, which have been put in place to ensure that data is findable, accessible, interoperable (capable of being integrated with other data) and reusable. Under these principles, the ELIXIR infrastructure offers useful data management tools that support researchers at various stages of the process.
“Good scientific practice involves making sure that data is well documented and remains usable throughout the research process, and in such a way that results can be verified later. It is important that researchers and information systems are able to find and access compatible and reusable research outputs. To ensure this, the FAIR principles were set out by a consortium of scientists and organisations in 2016,” explains CSC’s data management specialist Minna Ahokas.
“With instructions and tools provided by ELIXIR, it is easier for researchers to make their data findable, accessible, interoperable and reusable.”
The RDMkit website, created in cooperation with the ELIXIR nodes of the member countries, aims to support and harmonise data management practices in Europe.
RDMkit includes instructions and tips concerning the entire life cycle of data – from data management planning and data analyses right up to publication and reuse.
“RDMkit has been implemented in a way that anyone dealing with data is able to access the tools. It offers not only instructions but also links to services that researchers and any support personnel may need at various stages of data management.”
Finland’s ELIXIR node, CSC, is one of the parties producing content and maintaining the toolkit.
Ahokas stresses that the site was designed right from the start transparently in collaboration with researchers and data management experts. Anyone belonging to the ELIXIR infrastructure can participate in the development. Everything has been documented in the GitHub portal designed for software development projects.
“Data can be viewed in RDMkit throughout its life cycle. There are instructions for data collection, description and publication.”
RDMkit was developed in the ELIXIR-CONVERGE project, the aim of which is to help harmonise life science data management across Europe. There was a clear demand for unifying data management, as research projects are as a rule international, with data being transferred across national borders.
“RDMkit is the first major international attempt to unify data management practices and instructions to enable reusable data that is also sufficient in quantity and quality and described in a uniform way. Data management entails the planning of data collection, processing and description: how and where it is stored and how version management is handled. Whether some data should be stored for the long term also needs to be considered. And decisions must also be made about what data can be deleted.”
Ahokas emphasises the importance of offering researchers services that help them comply with good data management practices.
“We are trying to avoid such situations as researchers being presented with some new lists of data management requirements every time they apply for funding, without being offered the services needed to ensure that they can comply with these. If we demand that research project data management follows the FAIR principles, then we must offer sufficient support and services to produce FAIR data.”

CSC, Finnish research organisations and universities have created a national data support network. It supports cooperation between CSC and organisations, and provides a forum for open discussion, questions and peer support.
And at Aalto University, for example, for each discipline is assigned a data agent – that is, data management experts with experience in research. They will collaborate with researchers to manage data.
At the time RDMkit was launched, data management came under a new kind of pressure owing to the COVID-19 pandemic.
“When RDMkit was almost ready, the world was hit by the pandemic. It was then that we realised in the ELIXIR-CONVERGE project that data related to COVID and its requirements also had to be taken into account. That is why instructions were added in RDMkit specifically related to the processing of COVID-19 data, and the COVID-19 Data Portal was set up.”
RDMkit and ELIXIR’s data management instructions have also been adopted as part of the data management of EU’s Horizon Europe financial instruments.The RDMkit toolkit is recommended for use in the biosciences, and has also attracted interest worldwide. There are a considerable number of US users, and that country’s primary federal agency for conducting and supporting medical research, the National Institutes of Health, is interested in collaborating with the ELIXIR infrastructure.
RDMkit is a general collection of data management instructions with links to research data management tools such as IceBear.
“IceBear was originally designed for data management in crystallography and structural biology,” says Lari Lehtiö, Professor of Structural Biology at the Faculty of Biochemistry and Molecular Medicine.

Lehtiö is also the head of the Oulu unit of Instruct, a research infrastructure for structural biology. Structural biology unit of Biocenter Oulu, especially through efforts of Professor Rik Wierenga and the developer Ed Daniel, designed IceBear, a data management application for structural biology. The application has also been developed in the EOSC-Life network coordinated by ELIXIR. Instruct is part of this network. With the support of the EOSC-Life project, IceBear was transferred to the cPouta cloud service maintained by CSC.
Biocenter Oulu crystallises proteins and other macromolecules. The amino acid chain of proteins is folded in a 3D helical structure that is unique to each protein. As there is a huge number of ways in which the folding can occur, researchers need to study protein structures in laboratory conditions through experimentation, by crystallising. The three-dimensional structure of a protein can be determined on the basis of how X-ray radiation scatters from the protein crystal. Using the scattering data, mathematical transformation can be used to calculate the protein’s electron density map, indicating the location of atoms in the protein. These days structural research also makes use of cryogenic electron microscopy. This involves a frozen sample made of proteins being bombarded with electrons, with millions of individual 2D images of the proteins subsequently being combined into a 3D structure.
Automatic imaging equipment is used in the crystallisation of proteins. Proteins are crystallised in various solutions, with crystals being formed under certain conditions.
“A protein is crystallised in a droplet that is followed by imaging. Plates may contain up to 300 droplets, and the plates can number several hundred. With pictures of these being taken every day accumulating a lot of data. Crystallisation is usually carried out by robots,” says Lehtiö.
The crystal samples are picked manually from the microscope and placed in liquid nitrogen tanks. With the IceBear software, it is now possible to automate entries made of the samples and the data they contain.
“Often samples are sent to another infrastructure, to other synchrotrons [cyclic particle accelerators] in Europe. Thanks to IceBear, we can find out that eventually happened to the sample elsewhere. Metadata is transferred between databases used by European synchrotron and IceBear. Samples contain a fair amount of metadata, such as which protein the sample contains how it was crystallised and what the conditions were during crystallisation.”
IceBear does away with manual logs. Data can be transferred without the use of forms, and the links are created securely in the barcodes for each of the samples.
“You only need to do it once. The value of this application is that researchers’ time is saved even years from now,” says Lehtiö.
Ari Turunen
20.10.2022
Read article in PDF
Citation
Ari Turunen, Minna Ahokas, & Tommi Nyrönen. (2022). Reusable, accurately described and high-quality data – tools created by the research community for agile data management. https://doi.org/10.5281/zenodo.8154582
More information:
RDMkit
https://rdmkit.elixir-europe.org/covid19_data_portal
ELIXIR CONVERGE
https://elixir-europe.org/about-us/how-funded/eu-projects/converge
COVID-19 dataportal
https://www.covid19dataportal.org
EOSC-Life
https://elixir-europe.org/news/eosc-life-start
IceBear
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Post-doctoral Researcher Guilhem Sommeria-Klein at the Academy of Finland is developing mathematical models of microbial communities. He aims to build a unified statistical framework for describing the assembly of microbial communities. These models can then be applied to various environments, such as the ocean microbiome or the human gut microbiome. The goal is to better understand the role of microbiota in the functioning of ecosystems or human health. The research, carried out at the University of Turku, will result in open-source computing methods available for other researchers to use in their work.
High-throughput DNA sequencing has had a major impact on studying micro organisms. The method enables using the DNA sequence data sampled from any environment to assess the composition of the microbial community, whether it be in the soil, the ocean or the human gut.
“First we check what information can be found about the DNA sequences of the samples in the sequence databases. If a similar sequence cannot be found in the sequence databases, it will be difficult to know what the organism is. For example, the oceans have many organisms we do not know about. So in the end we are dependent on the sequence databases.”
However, Sommeria-Klein points out that not all species of plankton can possibly be included in the databases.
“We will never be able to describe and sequence all species of plankton. The huge diversity simply makes it impossible.”
However, this problem can be bypassed. Microbial communities can be classified into operational taxonomic units (OTU) using computational methods. The classification is based on DNA sequence similarity, and it is widely used in microbial research. The similarity is usually determined based on the sequence of a certain gene, chosen for its widespread occurrence and stability across the targeted microbial organisms.
“The similarity of the data derived from different ecosystems through the analysis of DNA sequences is striking. Organisms living in the human gut or in the ocean, especially bacteria, do not necessarily differ that much from each other.”
Sequenced genes of a microbiota contained in an environmental sample are analysed in bulk using the same method that is used to analyse the sequenced genes of one species. This approach, called metagenomics, is a common concept in the microbial research.
“We can use metagenomics to compare microbial communities sampled in different locations and study spatial patterns of variation, for instance. We can also determine the functions of certain genes in the microbial communities and how those functions change when the place and conditions are different.”

Unlike on land, microbes make up most of the biomass in the ocean. Phytoplankton are a key part of theoceanic microbial communities, since these organisms can combine water and carbon dioxide using energy from sunlight to form the organic molecules that constitute all living organisms, as plants do on land (photosynthesis).
“Since there are no plants in the open ocean, phytoplankton form the base of the whole ocean food chain,” says Sommeria-Klein.
The process also releases oxygen: phytoplankton are responsible for half of the oxygen in the atmosphere, and have a major impact on the oxygen content of sea water, thus enabling animals to live in the ocean.
“Although phytoplankton need light, they are actually often most abundant at a depth of around a hundred meters, where colder water that transport nutrients from the ocean depths meet the sunlight. The ocean is a three-dimensional environment: we miss a lot by only studying its surface. The biomass in the depths of the ocean, up to thousands of meters deep, is much larger than we thought previously. Due to darkness, there is no photosynthesis there. However, plenty of organic matter falls to the bottom, nourishing the deep ocean ecosystems.”
Guilhem Sommeria-Klein has access to enormous amounts of data on all the oceans of the world and at different depths. The research schooner Tara captured genetic material for DNA sequencing from the world’s oceans in 2009–2013. In total, 35,000 samples were collected from 210 locations around the world. The DNA analysis revealed over 40 million genes, the majority of which were new to science. About 250,000 molecular “species” of plankton were isolated from the samples based on DNA. The analysis was based on the metabarcoding approach, which is the analysis of DNA sequences in a specific gene region to identify different species or individuals.
“The ocean actually harbours a very wide range of microbes beyond phytoplankton. This view was much underappreciated until Tara expeditions. Microbial eukaryotes, in particular, are highly diversified and still poorly known. Moreover, the geographic distribution of plankton is not well known, as studying their habitat is difficult. In a recent study, we analysed the geographic distribution of different groups of eukaryotic plankton around the world and contrasted these distributions in light of the organisms’ main characteristics. ”
Sommeria-Klein’s interests lie in the functioning of these microbial communities and what they are doing in response to their environment in the oceans.
“Plankton are constantly moving with the ocean currents, which reshuffle communities and transport the organisms to different environmental conditions. I am fascinated by the way that these communities continue to interact, specialise and evolve under such challenging conditions.”
The oceans also have an important role to play as a carbon sink. Plankton communities greatly contribute to it by capturing carbon dioxide from the atmosphere through photosynthesis. This carbon is then recycled in the ocean food chain and finally sequestered in the ocean floor as dead organisms sink to the bottom.
“Global warming leads to changes in water temperature but also in ocean currents. These combined changes can have profound effects on the ecosystems, with an impact on the ocean fish stocks and the effectiveness of the ocean as a carbon sink.”

Guilhem Sommeria-Klein wants to develop more efficient methods for analysing and interpreting the data. Rather than solely specialising in mathematics or biology, his research aims to connect various disciplines.
“This represents the core area of our research team focusing on computational analysis, and the work of Sommeria-Klein supports it well”, says Leo Lahti, Associate Professor in Data Science at University of Turku, whose team develops machine learning models for screening microbial communities.
“Microbial ecology is in desperate need of this type of basic computational studies. These models will help break complex microbial ecosystems down to a few basic structures. Ocean microbiome research could also be useful in monitoring the changes in the state of the Baltic Sea. Models based on statistical reasoning can take into account any prior information and describe the uncertainty in the results. The high-performance computing services of CSC – IT Center for Science are needed to fit these models to the data.”
In the future, Sommeria-Klein wants to continue studying ecosystems that differ from each other.
“We want the perspective on microbial ecology to be consistent across ecosystems, as it is of major importance for various societal issues, such as human health, the ocean food chain and the global carbon cycle.”

Ari Turunen
29.9.2022
Read article in PDF
Citation
Ari Turunen, Guilhaume Sommeria-Klein, Leo Lahti, & Tommi Nyrönen. (2022). Gene sequencing used for study of structure and functioning of microbial communities in oceans. https://doi.org/10.5281/zenodo.8154571
More information:
University of Turku
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Antibiotic-resistant bacteria carry resistance genes and often become dominant in microbial populations. Bacteria can also mutate and receive genes from other microbes that make them resistant to antibiotics. Katariina Pärnänen, post-doctoral researcher at the Academy of Finland, searches for and identifies these genes that have been collected from different environments around the world and stored in databases.
Antibiotics, or drugs produced by microbes that kill other microbes, especially bacteria, have been used to fight infections since the 1930s. Use of antibiotics, particularly overuse or inappropriate use, results in bacteria becoming resistant to antibiotics. This is a growing problem, and because of it people die from infections that cannot be treated with antibiotics. Katariina Pärnänen studies antibiotic resistance at the University of Turku.
“When a person has been infected with an antibiotic-resistant bacteria, they will not respond to treatment with antibiotics. These days antibiotic-resistant bacteria are typically resistant not just to one but as many as 15 different antibiotics.”

Antimicrobial resistance genes are a growing problem globally. They are found not just in the human gut but also in soil, wastewater, fish farms and animal farms. Resistance genes found in our environment may ultimately end up in the human gut.
“Everything about microbes is of interest to me. From research perspective, it is quite useful that I have some understanding of what happens in pig farms, fish farms or in a river flowing through a big city in Indonesia. At the University of Helsinki, I studied the infant gut microbiome and the connection between formula or breastfeeding and the number of resistance genes.In a way, my research represents the ‘One Health’ approach,” Pärnänen says.
Discovery of new viral diseases and threats of such diseases in the early 2000s led to the emergence of the One Health and Global Health research concepts: a broad understanding of the need to protect human and animal health and the ecosystem.The movement initiated by physicians and veterinarians represents an interdisciplinary approach that covers the local, regional, national and international levels.
“I want to address antibiotic resistance as a problem that affects human health, animal and food production, and environmental health. Previously I studied the ways in which resistance genes were passed on from mothers to children, and how the duration of breastfeeding affected the number of these genes.Now my focus is on the various factors linked to antibiotic resistance of the gut microbiota.These include overuse of antibiotics, living environment and health history. These may provide an indication of whether the person is likely to carry antibiotic-resistant bacteria.”

The study of environmental bacteria and their genes took a giant leap forward with the introduction of next generation sequencing methods. NGS is based on massively parallel sequencing, where millions of short DNA fragments are cloned simultaneously.The analysis method is referred to as metagenomics, as it allows the genome analysis applied to an individual species to be used for a group of genes that were recovered from the environment and sequenced.The first metagenomics based study of antibiotic resistance was published in 2014.
“Antibiotic resistance has been studied for a long time, but with NGS we can now analyse all resistant genes from a single sample instead of just individual genes,” Pärnänen explains.
Metagenomics related studies have shown that antibiotic resistance genes are commonly found in our environment. The risk of these genes transferring to bacteria that cause infections in humans is significant.
Pärnänen uses metagenome sequence data that is stored in extensive open databases. She uses supercomputers at CSC – IT Center for Science to analyse the data,allowing her to identify different species of bacteria and their resistance genes.
“In some species of bacteria, it is possible to identify genes that appear in its genome only once. By comparing these with genes in the databases of other species, we can identify the species in question. We look for matches with resistance genes stored in databases. Then we can say that the person has ten resistance genes in their stool sample, or that they carry a certain amount of coliform bacteria.”
Pärnänen was one of the researchers in a study where resistance genes from human feces were analysed. The study involved a comparative analysis of bacteria found in wastewater treatment plants in seven countries. Half of the world’s population carry the CrAssphage virus – a bacterial parasite – in their intestines. The gene sequence of this phage was used in the study as a marker to indicate feces-based infection.
“The same resistance genes can be found all over the world. Antibiotic resistance is often called an invisible pandemic, because the same resistance genes are spreading beyond country borders. However, certain genes are more common in some parts of the world than others.”
These genes may be found in larger numbers in India than, for example, in Northern Europe. There are also significant differences between Southern and Northern Europe.
“The E. coli strains that cause urinary tract infections in southern Europe can be very resistant.”
Bacteria do not cause a serious disease in a healthy person, but sometimes an intestinal infection may be caused by a resistant bacterium.
“An infection caused by resistant bacteria is difficult to treat. Usually the most difficult resistant infections found in Finland are diagnosed in persons who had travelled abroad.”
In 2002, stool samples were collected from Finnish adults as part of the FINRISK study by Finnish Institute of Health and Welfare (THL), and the sequence data of the microbes found in the samples was analysed.
“Together with THL researchers we analysed samples from the Finnish population to see what the potential impacts of resistance could be and how a large number of resistant bacteria would affect a person’s health. Are high levels of antibiotic resistance genes associated with risk of death during the follow-up period?
It has been predicted that in 2050 more people will die from antibiotic-resistant infections than from cancer. Infectious diseases would be the most common cause of death.Antibiotics are currently being used more on farm animals than on humans, and at the same time our consumption of animal protein is growing. According to Pärnänen, the best way to fight the resistance crisis is to use antibiotics only to treat bacterial infections and only when there is evidence to prove that antibiotics will help.Similarly, an appropriate diet or lifestyle may reduce resistance genes in the intestinal microbiota.
“In studies conducted recently in the United States, fibre intake was linked to a low rate of resistance genes while animal protein in the diet was linked to a high rate of genes. You could say your gut microbiota is what you eat.”
Katariina Pärnänen works in the research team of assistant professor Leo Lahti. The team develops machine learning models that screen microbial groups from large data collections.
“Antibiotic resistance is one example of a research area where we use new measurement methods and computing capacity in a way that has not been done before.This research naturally combines different measurement environments from the human body to the environmental microbiome.This type of research generates ideas for method development that benefits researchers in various fields,” Leo Lahti comments.
Katariina Pärnänen would like to extend her research to the world’s entire human population and study the gut microbe samples that have been metagenome sequenced and are openly available in databases.
“It would be interesting to discuss the technical feasibility of such a project with CSC experts.It would also promote open science, because the identified resistance genes and species found in microbiomes could also be stored for use by other researchers.”
Ari Turunen
1.9. 2022
Read article in PDF
Citation
Ari Turunen, Katariina Pärnänen, Leo Lahti, & Tommi Nyrönen. (2022). Antibiotic-resistant bacteria are a global problem. https://doi.org/10.5281/zenodo.8154563
More information:
University of Turku
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Modelling cells and simulating how they work gives a boost to personalised treatment plans. The PerMedCoE project combines clinical patient data with data related to the operation of genes, proteins and cells. The goal is to develop tools that can be used in precision medicine. The modelling of cells in detail is, however, a major undertaking, requiring a lot of computing power by supercomputers.
Personalised medicine will open up great opportunities in the future. The goal is to be able to combine a patient’s clinical data with genetic data to create personalised treatment plans. The PerMedCoE project (HPC/Exascale Centre of Excellence in Personalised Medicine) is working to improve the compatibility of personalised medicine modelling software with next-generation exascale supercomputer systems. Their theoretical computing power is as much as 10^18 operations per second. The project involves researchers from several European universities and hospitals, and focuses on four cellular-level modelling software systems based on open source code. Further to working on software development, the research project aims to make precision medicine tools easier to use and compatible with a number of European high-capacity computer centres.
“Our aim is that these four software systems could be used in many supercomputers,” says Project Manager Sampo Sillanpää from CSC – Finnish IT Center for Science.
“At the moment this is technically very challenging to implement, because all high-performance computing environments are unique in terms of their system architecture.”
The plan is to achieve seamless operation of software and data masses through jointly agreed technologies. In the PerMedCoE project, this is done by means of workflow software and what is known as container technology. Workflows are used to automate different steps required to analyse data. Further to the actual modelling step, a workflow may involve, for example, data pre-processing and steps relying on results produced by the modelling tool. Container technology can be used to specify a standardised environment in which scientific software is run in each high-performance computing environment taking part in the project. Once the software code and libraries and settings are placed in the container, they can be transferred from one computer to another.
“The software and data are, in a manner of speaking, packaged in a single box, making it easy to move it from one environment to another. CSC have several container technology experts, so the tools can be transferred from one platform to another,” says Sillanpää.
“The containers enable experts to create user-friendly workflows. Workflows in the PerMedCoE project consists of several building blocks, each performing a specific precision-medicine calculation. One building block may pre-process data, a second one carrying out the actual analysis, and a final one delivering the outcome to the end user. This means that the users may not even have to know how many building blocks the automation contains, but focus on analysing the results.”
The usefulness of technologies built in the project is assessed by means of various use cases. Workflows are used to analyse which disturbances may be caused at the cellular level by diseases and how the drugs that have been administered actually work. The models can be used to examine cellular metabolism and signal transmission.
“In PerMedCoE use cases, we make use of publicly available genomic data. Now we can study samples taken from coronavirus patients and look for markers in the genomic data to learn which patient groups are particularly susceptible to the dangerous forms of the disease.”
The project models the skin’s epithelial tissue that reacts to a coronavirus infection by calling various immune cells to work against the virus. This may help to identify patient groups that are susceptible to the serious form of the disease.
“The idea is that we are able to run several models simultaneously for individual patients. This results in efficient analysis of sufficiently large amounts of data, so that the modelling results can be used for personalised medicine,” says Senior Data Scientist Jesse Harrison of CSC.
When modelling the COVID-19 use case, cellular-level RNA sequence data is used. Single-cell RNA sequencing (scRNA-seq) may reveal regular interaction between genes, cell lineages, cellular difference and the cellular frame of reference in its environment.

Another key use case of the project concerns cancer diagnostics. The goal is to create modelling tools for the prediction of cancer tumours and the development of patient-specific treatments. The material has been collected by the Wellcome Institute and the Massachusetts General Hospital Cancer Center. The database contains more than a thousand tumour-tissue cell lines.
“The project aims, among other things, to identify new drug combinations that could be useful in cancer treatment,” says Jesse Harrison.
This will hopefully lead to more personalised cancer treatments and faster diagnostics.
“In order for this to become reality, closer collaboration will be needed between high-performance computing centres and medical organisations. This is because we are talking about large amounts of data, and analysing such large-scale personalised data sets is simply not possible with a desktop computer.”
The results and tools of PerMedCoE are open to all researchers.
“When the project ends in summer 2023, we will have updated versions of modelling tools developed from open source code, and these will be made available to the research community. The project will also create new expertise to support the use of precision medicine tools in CSC computing environments.”

The EU is funding a number of projects that will enable personalised patient treatment in the future. Cancer is an example of a disease that is extremely individualised, whether it is breast, lungs, liver or prostate cancer.
For example, the Conquering Cancer: Mission Possible programme under Horizon Europe will – according to Esa Pitkänen, researcher at FIMM (the Institute for Molecular Medicine Finland) – pave the way to future cancer research and treatments. The ambitious programme strives to understand the mechanisms that lead to cancer, to discover methods for early detection of cancers, and to achieve breakthroughs in personalised cancer medicine.
“What is common to all these goals is a versatile and comprehensive use of health data by means of new calculation methods. Artificial intelligence algorithms based on machine learning have indeed already achieved some encouraging results in terms of, for example, digital pathology. The next leaps will be made by combining various data sources in order to make individualised cancer screening and treatment recommendations,” says Pitkänen confidently.
Within the programme, cancer patients become active participants in cancer treatment development, for example by being able to send their health data securely to researchers. This also gives the patients new research data about their own illness.
“It is important that as treatments develop, people are given an equal opportunity to benefit from new treatments regardless of their background. I am glad to see that this has been taken into account in the programme’s recommendations. There is also special emphasis on cancers among children and young people.”
Ari Turunen
23.8.2022
Read article in PDF
Citation
Ari Turunen, Sampo Sillanpää, Esa Pitkänen, & Tommi Nyrönen. (2023). Personalised medicine against cancer and viruses. https://doi.org/10.5281/zenodo.8154548
Mode information:
HPC/Exascale Centre of Excellence in Personalised Medicine
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Together, microbes and their interactions with the host are called a microbiome. Microbiome composition is unique to each person. The microbiome helps the body’s defence system fight infections, for instance. If the microbiome is disturbed, the body could be exposed to diseases such as diabetes.
Leo Lahti, Associate Professor of Data Science from the University of Turku, is developing the machine learning models with his research group and partners for screening microbial signatures from large-scale data collections.
‘The microbiome is just one element in development of diseases, but it is an element that we haven’t been able to study as extensively before, because the effective data collection methods we have now have become available only very recently,” says Lahti.

Lahti’s research group has characterized microbes in different habitats and ecosystems together with experimental researchers.
”Scientists are starting to have a pretty good idea of what kinds of species and groups of bacteria can be found in different environments. We have learned quite a lot about their function, tasks, and role in metabolism and the chemical compounds they produce.”
All bacteria and single-celled archaea are microbes. Microbes also include algae, protozoa, yeasts and moulds. According to Lahti, microbial research has advanced rapidly after the prices of sequencing technologies started coming down. The DNA of microbial samples is sequenced and the species of microbes in the sample can be identified if they have been characterized earlier.
‘The sample can come from the environment, parts of the human body, or just about anywhere. We study bits of DNA and try to put the puzzle together. That way we can determine the bacterial composition of the given sample. We can even trace completely new bacterial genomes and discover previously unknown species,” says Lahti.
The most diverse microbial ecosystem in our bodies thrives in the gut. According to current knowledge, a typical adult carries around 1 to 2 kg of bacteria on average, which is slightly more than the amount of human cells. There are many levels in the human microbiome, and the levels form diverse ecosystems. The composition of a person’s microbiome is affected by their genome and living environment. Habits such as diet and an outdoorsy lifestyle have been shown to affect the composition of a person’s microbiome. A person’s microbes can be used to deduce whether the person is a vegetarian or an omnivore, for example. Lahti’s research group has also been involved in studying the microbial compositions of different population groups.
“Identifying the species in the human microbiome in general was a huge task. The composition can vary geographically, i.e. according to where people live and whether they belong to the indigenous population, or whether they live in the city or the countryside, or their standard of living.”
Studying the composition helps with understanding how microbes are connected to a person’s health.
“In this context, linking a person’s current state of health with their future health status takes centre stage. Is it possible to infer something on a person’s current or even future state of health by looking at their microbiome? And if it is possible, can the person’s health be affected by modifying the microbial composition, and what kinds of risks or ethical questions are related to this? Computational and machine learning techniques are in a key role in extracting information from the complex data sets that are now being generated.”
In Finland, stool samples were collected in connection with the Finnish Institute for Health and Welfare’s (THL) FINRISK population survey in 2002. The many years’ worth of follow-up data from these individuals is now enabling the study of the links between microbiome composition and long-term changes in health status. Lahti says the extensive Finnish population cohort is unique at a global level.
‘The data is very valuable; carrying out a study like this would be hard in many countries because similar comprehensive population register data is often not available. We now have a huge number of samples and associated health information that we can use to study the link between microbial composition and population health.”
According to Lahti, some microbial analyses can be used in diagnostics. They could be used to identify specific diseases or to identify microbes that predispose humans to the risk of certain cancers. For example, Helicobacter pylori found in the stomach may increase the risk of stomach cancer.
“Certain groups of bacteria are found in the gut that are statistically linked to the risk of developing a disease later on. We have recently discovered that they can indicate an increased risk of mortality, liver disease, and type 2 diabetes, for instance. We do not yet understand the causal relationships between these observations, but we can see the signals years before a person gets sick.”

Research results have been published on mortality rates, liver diseases, and type 2 diabetes.
‘These are significant disease groups that are studied frequently anyway. Despite the long research traditions associated with them, microbiome studies have brought a new perspective into understanding these diseases . Microbes play a part in our metabolism. The compounds produced by microbes in the body can have a significant role in these diseases and in immune systems.”
“After we learn more about what these microbes do and what microbes are found in our bodies, we will have a better chance at understanding the mechanisms that affect the development of diseases. This can help with developing new ways to curb the effects of diseases or prevent the risk of developing them in the first place.”
According to Lahti, there is currently huge medical interest in microbiomes, because lifestyle changes and many common diseases have been found to be linked to changes in the microbial balance. In addition to this, antimicrobial resistance is a growing health problem, among others. It refers to the increased ability of bacteria to withstand antibiotics, and it is predicted to be the leading cause of death in the coming decades.
Lahti’s research group extracts information from big data and merges information from different sources. The size of datasets is constantly increasing, and they need to be structured and organized in order to make them understandable. Such analyses have many computational steps. First, the data must be pre-processed and the DNA fragments must be combined in order to see which species they come from and in which proportions they occur in different samples. After this, the connections between the microbial composition and the living environment or state of health of the host organism can be studied in more detail.
“Data can be complicated. It can be hierarchical and have temporal or spatial structure. So, we need new computational methods. For example, machine learning methods are useful because they reduce the need for human intervention, which means we can automate a significant part of the processing and transfer it to machines.”
According to Lahti, methods that can assist people in making quantitative conclusions play a big part in biomedical research.
“Data is collected in databases. And when we analyse new samples, we want to combine the sample data with the data already in the databases. The newly observed data must be interpreted in the context of the previously collected data and accumulated knowledge.

According to Lahti, when studying species of microbes, it is important to understand how they work together as an ecosystem and interact with the human body. Sequencing the genomes of groups of microbes and data integration can require massive computing and storage resources.
“We need the resources provided by CSC to obtain comprehensible information from the large-scale sequencing data that we can then analyse statistically. We increasingly use these services as a platform for cooperation. We can build workflows with other research groups and make the data available through CSC, and the data analysis platform is also in one place, on the CSC servers. It is also important that the bioinformatics data resources provided by ELIXIR can be accessed via CSC. We are also increasingly using these services to provide training in computational research methods.”
Ari Turunen
30.6.2022
Read article in PDF
Citation
Ari Turunen, Leo Lahti, & Tommi Nyrönen. (2022). Studying the human microbiome is a key towards holistic understanding of our health. https://doi.org/10.5281/zenodo.8154534
University of Turku
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The large-scale FINRISK population survey of risk factors for chronic, noncommunicable diseases has been used to collect health data on the population every five years since 1972. The data can be analysed to identify risk factors for chronic diseases. More and more genetic data is also being collected, and when combined with registry data it makes it possible to develop measures to prevent diseases and to create more effective treatments.
There are many Finnish survey datasets, but according to Kati Kristiansson of the Finnish Institute for Health and Welfare (THL), FINRISK contains exceptionally rich and diverse data on the health of the Finnish population. The participants are randomly selected from populations in different regions of the country. They are asked about their lifestyle, family history of illness, mood and other factors related to health and wellbeing. Registry and survey data can be combined with genetic samples.
“When all these laboratory measurements and questionnaire data are combined with health registry data, we can learn about people’s medical histories, what medications they have taken and all the causes of death in the population.”
In 2015, the FINRISK data collections were transferred to the THL Biobank. Two years later, the FINRISK and Health 2000 surveys were combined into a new FinHealth population survey. Naturally, there are also other biobank datasets containing genomic data and data from health examinations. What makes the FINRISK data of exceptional quality is its time span.
“Various values are measured at the person’s first health examination. When this baseline has been established, the same person can be tracked in the registers for up to 30 years to see the developments over time regarding their health. You can see what kind of risk factors were present at the beginning, what kind of illnesses occur over the years, or the cause of death.”
According to Kristiansson, what is particularly valuable in the FINRISK population data is the monitoring over time after the initial measurements have been taken.
“This kind of analysis helps in determining the factors that increase the risk of future illness. Hereditary and lifestyle factors are a very useful aid in this regard.”
FINRISK data is stored in the THL Biobank. New data from different research projects is being added all the time. Kristiansson’s research relies heavily on biobank data. She leads the Public Health Research team in the Population Health Unit at THL and is involved in collecting data for the Terve Suomi (Healthy Finland) research project, which began in 2022 and will provide up-to-date information on the health and wellbeing of adults living in Finland.
Kati Kristiansson has used the FINRISK data throughout her research career.
“In my early days as a researcher, I studied the genetic determinants of diseases affecting broad segments of the population, with a particular focus on risk factors for cardiovascular disease. This is what I have been doing ever since – I have always had a keen interest in identifying risk factors for public health diseases and assessing possible preventive measures.”
In the FINRISK study, 10,000 people are asked to participate in health examinations every five years. The most recent examination also involved taking a stool sample, and sampling the person’s microbiome – the population of microorganisms inhabiting their body. In addition, a blood sample was taken to study each research participant’s metabolome, which is the full range of hundreds of metabolites that are present within the body.
“We also obtain data on obesity in the population, through body mass index measurements and a lot of other information that is not otherwise in the registers.”

Kristiansson has studied risk factors for type 2 diabetes and coronary heart disease. She is particularly interested in the use of genomic data for the prevention of diseases affecting broad segments of the population.
“Finland has excellent registers, and combining them with different data makes for a very high standard of research. But surveys, health examinations or registers alone are not enough. To identify hereditary risk factors for common diseases, in addition to registries it’s essential to have genomic data.”
Blood samples are also taken at health examinations, in addition to the aforementioned comprehensive measurements. This allows the isolation and storage of the person’s DNA and the measurement of various lipid values, lipid cholesterol and blood sugar.

In Finland, DNA has been isolated from samples taken at FINRISK health examinations since 1992. The samples have now been subjected to whole genome genotyping, which uses DNA microarray technology to determine the genetic information contained in the DNA. Genotyping involves reading hundreds of thousands of chromosomal loci, and then extrapolating the information to millions more using statistical methods. These sites contain many genetic variants associated with different diseases.
FINRISK data has also been combined with data collected in other countries. International research projects often collect as many samples as possible in order to conduct comprehensive genome-wide association studies (GWAS). The purpose of these is to examine the genetic loci associated with various diseases and traits. The FINRISK database contains a wealth of research data on cardiovascular disease and its risk factors, including obesity, diabetes, blood cholesterol level, blood sugar and genetics.
Kristiansson is interested in the biomarkers found in the data that indicate changes in biological status. One focus area in analyses of the FINRISK data is peptides, the relatively short proteins that each consist of a sequence of less than 50 amino acids. Natriuretic peptides are peptides that induce natriuresis, which is the excretion of sodium ions and water by the kidneys. The effect of this is to reduce blood pressure.
Two of these small proteins, atrial natriuretic peptide (ANP) and (atrial peptide) and brain natriuretic peptide, also known as B-type natriuretic peptide (BNP), are secreted from the heart into the bloodstream and function as hormones.
The secretion of these peptides is regulated by the pressure load on the muscular layer of the heart wall. Heart failure involves an increase in the concentrations of these peptides in the blood plasma. This makes them effective clinical biomarkers of cardiac stress.
The study in which Kristiansson was involved found interesting differences in the amount of peptides in the blood of different people. The research team investigated the effect of gene variants on peptide levels and compared the effect of the these variants on blood pressure. The studies took into account place of residence, age, gender, whether the person smoked and how heavily, blood pressure, and the glomular filtration rate (GFR), the filtration rate of capillaries in the kidneys. Deteriorating kidney function is manifested as a drop in the GFR. Genetic research can shed further light on the specific factors and gene regions that affect peptide levels and cause changes in blood pressure.
According to Kristiansson, such findings from analysis of the FINRISK data provide an indication of which gene regions could be targeted for further research and drug development.
“Once we know these regions of the genome that affect biomarkers of disease, further research can be done to try to find a good drug protein.”
According to Kristiansson, cholesterol-lowering drugs such as these are good examples of these innovations. Drug development is carried out through several projects, including the FinnGen research project coordinated by the University of Helsinki. The data generated in the course of the project will be made available to other researchers and companies, both in Finland and internationally.
According to Kristiansson, the FINRISK data is particularly useful for studies of disease prevention.
“One aim of the project is to gather information which can be used to help people adopt a healthier lifestyle.”
Ari Turunen
23.5.2022
Read article in PDF
Citation
Ari Turunen, Kati Kristiansson, & Tommi Nyrönen. (2022). FINRISK: one of the world’s longest-running population survey time series. https://doi.org/10.5281/zenodo.8154515
Finnish Institute for Health and Welfare THL
https://thl.fi/en/web/thlfi-en
FINRISK Calculator
https://thl.fi/en/web/chronic-diseases/cardiovascular-diseases/finrisk-calculator
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
The Finnish ELIXIR node CSC is building an infrastructure in which human data obtained from Finland’s biobanks and research organisations has been pre-processed and described and saved in a secure way. The parties responsible for sharing the data can automate their authorisation process with the CSC platform. This improves licensed availability of data for research and healthcare purposes.
In a project funded by the Academy of Finland, an infrastructure is created that meets the requirements for storing and using sensitive data. The data consists of clinical register data, genomic data and material related to bioimaging. The project is participated in by not only CSC but also the biomaging infrastructure Euro-BioImaging, THL Biobank and the Institute for Molecular Medicine Finland (FIMM).
The data can be stored in CSC’s sensitive data infrastructure. Researchers are allocated a space in which the data and computing power are in the same place. Researcher can only access data to which authorisation has been obtained by the data owner. The project also makes use of federated data management developed by CSC. ELIXIR AAI and REMS are applications developed by CSC for the managing users in the ELIXIR infrastructure.
Read more here: https://www.elixir-finland.org/en/sensitive-data-infrastructure/
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org
https.//www.elixir-europe.org

THL Biobank contains a large amount of data on the health and lifestyles of the Finnish population, the collection of which began already in the 1960s. When this is combined with genetic data stored in the biobank and with national health registers, illness risk factors can be effectively identified and predicted.
“Data in the biobank can be used for practically any health research,” says research manager Kaisa Silander, who has contributed to the descriptions and classification of THL Biobank’s population cohorts.
“We have plenty of data from various population research studies and when these are put together, researchers have an impressively large material at their disposal.”
Silander considers the material to be significant also by international standards.
“I think it is as valuable as UK Biobank or Estonia Biobank. When biobank data is combined with health registers, we have health information about certain people over a period of 40 years. We know exactly, for example, which diseases a certain person has had.”
Silander has been working with cohort data for a long time. Projects funded by e.g. EU Health programmes built the infrastructure. After that she helped combine the metadata for research cohorts managed by the Finnish Institute for Health and Welfare (THL) into a searchable database.
“We built a common infrastructure for data storage, because all these cohorts contain similar data. First we created a catalogue in which these cohorts were described in a uniform manner. We saved the variable metadata into the same database, in order to make them easy to search and to find. “The description of the variables was done in Finnish and English. The same protocol will also be used for future cohort description,” says Silander.
THL Biobank was established in 2014. The population cohorts collected by THL had two traditional lines of research. Between 1965 and 1980, population health examination surveys were carried out by driving around the country on buses modified into mobile clinics. The mobile clinic examined more than 50,000 Finns around the country. THL’s population cohorts are composed of information provided by the participants of the health surveys, of a baseline clinical examination, and of a sample bank. Most of the samples have been turned into data. For some participants up to 40 years or more of health register follow-up can be obtained. The mobile clinic was followed up in 2000–2001 by the nationwide Health 2000 study and a further follow-up of the latter in 2011–2012. The mobile clinic research included pulmonary diseases, heart conditions, anaemia and iron deficiency, diabetes, kidney and urinary tract conditions, thyroid conditions, calcium metabolism diseases and coronary artery disease. Later the range of diseases has been extended, and today biobank studies can be conducted on any disease.
The FINRISK study has collected information mainly about risk factors for cardiovascular diseases and diabetes, originally in eastern and central Finland. Later the study was expanded to cover several other areas in Finland, and the range of diseases studied was also increased. In 2017, these two lines of research were combined in the FinHealth study.
“The cohorts contain plenty of lifestyle data obtained with survey questionnaires,” says Silander.
The questionnaires include questions about smoking, alcohol use and diet, sleeping and exercise habits. The health checks have included height, weight, blood pressure and other measurable matters. Blood, stool and urine samples have also been taken.
“The samples are used to determine biomarker levels, such aslipids and inflammation (C-reactive protein, CRP). CRP is a protein produced by the liver, and its concentration in the body increases rapidly during inflammation.”
Kaisa Silander hopes that in the future we would be able to obtain much more biomarker data describing changes in a person’s body that may give an indication of an illness.
“There are good methods for high throughput biomarker analysis. Currently we have at the biobank information about more than two hundred biomarkers which were analyzed by NMR spectroscopy. However, there are laboratories that can produce thousands of biomarkers from a single serum sample. This type of information, combined with the FINRISK material, for example, would be valuable. There are still suitable serum samples for many of the FINRISK participants.”

Thanks to the FinnGen research project, THL Biobank may offer genetic data for other biobank studies.
The goal of the FinnGen project, started in autumn 2017, is to collect the genome data of half a million Finns. The project utilises samples collected by all Finnish biobanks. Genome data is combined with data available in national healthcare registers. This gives a better understanding on how diseases develop, and identify new treatments. Phenotypes used in FinnGen are age, gender, height, weight and smoking. Genome data created in the FinnGen project is returned annually to biobanks.
“If you can combine health register, questionnaire and genome data, it certainly enables extensive research areas,” says postdoctoral researcher Heidi Marjonen. She works as a genome expert at THL Biobank, processing genomic data of all THL Biobank cohorts.
THL Biobank’s sample collections contain genotype data of densely mapped single variants of DNA, and also data of whole genome sequence, and exome sequence data of the protein coding regions of DNA.
All exome and whole genome sequence data related to the FINRISK and Health 2000 were produced at Washington University and the Broad Institute/Massachusetts Institute of Technology, which are among the leading laboratories for genome studies. The FINRISK cohort consists of 10,000 exome sequences and 4000 whole genome sequences. Combining this data with Finnish health data, enables more accurate study of diseases.
“Now it will be possible to create personalized treatment methods. When lifestyle data is combined with genetic data, better drug treatments can be developed,” says Marjonen.
According to Marjonen, DNA samples can also be used to produce epigenetic data.
Epigenetic inheritance means the transfer of hereditary data to an offspring of a cell or organism without the data being coded into the DNA or RNA sequence. Epigenetic factors are affected by many external factors, such as dietary habits.
Another interesting dataset is the microbiome, which is data about the microbes in the human intestine. Stool samples from the 2002 FINRISK cohort have been studied to determine the sequence data of all microbes. This data can be utilised to see how the microbiome affects human health.
Heidi Marjonen took part in a study in which more than 3,000 persons were given information of their 10-year disease risk for the most common diseases in Finland. When genetic data is combined with clinical data, one can predict individual´s disease susceptibility. The overall 10-year risk evaluation was based on the genetic data and other traditional risk factors, such as gender, age, body mass index, blood pressure and cholesterol levels. The genetic risk was calculated as a personal polygenic risk score, taking into account millions genetic variations.
This data was stored on the ePouta platform for sensitive data in Finland’s ELIXIR node CSC, which enables a secure transfer between the portal’s user interface and the database.
The polygenic risk score, says Heidi Marjonen, is a major research trend. The risk score is a single value that reveals the genetic burden of a disease.
“Researchers receive information about the genome in a convenient way and allows to study the effect of genome on a disease or other traits in an individual.”
Ari Turunen
8.4.2022
Read article in PDF
More information:
THL Biobank
https://thl.fi/en/web/thl-biobank
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Diabetes is a major chronic disease that comes with a number of associated conditions that pose remarkable challenges. Such diseases include diabetic kidney disease, diabetic retinopathy, coronary heart disease and strokes. Now, a Finnish research team sequences the genomes of thousands of individuals with diabetes to look for genetic risk factors.
Individuals with diabetes have a higher risk of cardiac diseases in comparison to the rest of the population. A third of individuals with type 1 diabetes, earlier also called juvenile diabetes, develop kidney disease that has a remarkable impact on mortality and the risk of cardiac disease. On the other hand, diabetic retinopathy is the most significant cause of blindness among working-age population.
Finnish children and young adults have the highest risk of type 1 diabetes in the world. Type 2 diabetes is often considered a disease of the western life style, but the highest patient concentrations are found in middle-income countries with China and India topping the statistics of individual countries.
“Diabetes is associated with remarkable and severe complications. The associated conditions have a major impact on the quality of life and life expectancy,” explains Niina Sandholm, Genetic Epidemiologist at Folkhälsan Research Centre. Sandholm is involved in the FinnDiane research project, the objective of which is to identify hereditary and environmental risk factors predisposing to diabetic complications. FinnDiane-study is a collaboration project between University of Helsinki, Helsinki University Hospital (HUS) and Folkhälsan Research Center
According to Sandholm, genetic data could be beneficial to young patients in particular, already at an early stage prior to the emergence of risk factors.
“Currently, the use of genetic data in clinical treatment is mostly associated with rare diseases, but our results and earlier research suggest that extensive genetic data could also be utilised in the early prevention of common diseases.”

FinnDiane, established by Professor Per-Henrik Groop in 1997, is a follow-up study participated in by almost 8,000 individuals with type 1 diabetes. The participants are recruited at 80 hospitals and health centres across Finland. It is one of the most extensive research materials on type 1 diabetes and associated complications in the world. Now, this material is used for sequencing the genome of 1,700 patients.
Sandholm has experience in research projects where genome-wide association study (GWAS) is applied as research method. The method is particularly useful when the genetic background of the studied disease is complex. It enables identifying genetic variants that either increase the risk of diabetes or protect from the disease. The GWAS method involves identifying genetic variants from the participants’ blood samples. The number of these variants ranges from hundreds of thousands to millions, and the number of patients may vary between thousands and hundreds of thousands.
A GWAS study on 5,600 FinnDiane participants with type 1 diabetes revealed, for example, a new genetic locus related to cardiac diseases close to the DEFB127 gene. It is the most extensive study of its kind to date. Locus means the location of a DNA sequence on a chromosome. The variation of a sequence is called an allele.
The same study that identified the DEFB127 gene also revealed other genetic factors predisposing to cardiac diseases.
“Many predisposing genetic factors have been identified for cardiac diseases and other common medical conditions. One of the most significant factors lies within the region of genes CDKN2A and CDKN2B. Diabetics have a remarkably higher risk of cardiac disease than the rest of the population, and there is little knowledge of the related genetic factors. Our research indicated that the same genetic area CDKN2A/B affects the risk of cardiac disease also in individuals with type one diabetes.”
A third of individuals with type 1 diabetes develop a kidney disease. Some may develop renal failure that may, at worst, lead to the need for dialysis treatment or a kidney transplant.
Another study analysed various data sources to identify connections to kidney disease in 27,000 diabetics. GWAS is a fast and economic method, but it cannot identify all variants. Therefore, the research team have turned to sequencing the entire genomes of patients.
“Variants identified with the GWAS method are common, and individual variants’ impact on the risk of developing a condition is quite moderate. The objective of sequencing is to identify rare variants that may have a significant impact on developing a condition at the level of individual patients. The worst case scenario is that such a variant prevents the functioning of an entire protein.”
According to Sandholm, the research results may help predict the risk of developing a condition or lead the way in developing new medicines.
“The broader goal of genetic research is to identify variants that affect the risk of developing a disease or directly cause a disease. This enables a better understanding of the causes of diabetic complications.”
The ultimate goal is to learn to prevent and find cures to diseases associated with diabetes.
“Our aim is to read the entire DNA sequence of all patients. This will result in a huge amount of data,” says Sandholm.
“The sequencer produces DNA data in strands of 150 base pairs. To verify the data, our goal is to read each one of the three billion DNA base pairs an average of 30 times. This means that there will be 600,000 strands of 150 base pairs per patient.”
To map the entire sequence, the sequenced strands must be placed in the correct order with the help of a reference human genome. This requires enormous computing capacity which is provided by the ELIXIR centre at CSC, the Finnish IT Centre for Science.
“The purpose is to organise the data so that it would enable identifying how each base pair variant affects a given disease at the level of individual patients. Our aim is to identify rare variants that cannot be identified with the GWAS method. Only a few patients in the sample exhibit rare variants.”
Single nucleotide polymorphisms (SNP), or variants in the DNA base pairs, are a sort of end result of the data processing.
“The variation in the DNA sequence is expressed as SNPs. Each patient has either no alleles or one or two variants. They are markers that indicate which diseases the variant may cause.”
The research group has already sequenced the entire genomes of 600 patients.
“Based on the initial results, we identified individual variants that are clearly associated with the risk of stroke, for example. There are also variants in genes that have previously been associated with congenital kidney diseases. Now it appears that variants in the same genes also affect the development of diabetic kidney disease.”
Along with her colleagues, Niina Sandholm studies the protein-coding parts of the gene and gene regulatory regions that may have links to risk factors contributing to diabetes.
“The area between genes – 95% of the genome – contains plenty of regulatory regions that determine which gene appears in which tissue. As such, the DNA sequence is the same in each human cell, but gene regulation causes eyes to develop into eyes and kidneys into kidneys. In this respect, gene regulatory regions and their changes play a key role.”

This study is one of the world’s first to sequence the entire genome this extensively with regard to a specific disease. For the time being, the sequencing of the entire genome is relatively rare.
“The current trend is to sequence the exome with a focus on the protein-coding parts. However, it is only a matter of time that sequencing the entire genome becomes more common. ELIXIR, for one, invests in the development of full genome sequencing and genomic data processing methods.”
CSC provides the ePouta service for processing sensitive data. In the ePouta cloud service, virtual private servers operate on CSC’s computing platform under increased data security. The users receive dedicated cloud resources which are separated from CSC’s other computing environments. The FinnDiane research group uses the computing cluster of Institute for Molecular Medicine Finland (FIMM) which is connected to CSC’s sensitive data computing platform via the ePouta light path. By scaling the computing resources, the light path enables faster processing of the project data. In addition, the researchers have been allocated a remarkable amount of storage space to store the genomic data.
Ari Turunen
3.3.2022
Read article in PDF
Citation
Ari Turunen, Niina Sandholm, & Tommi Nyrönen. (2022). Finnish research team sequences the genomes of thousands of individuals with diabetes to look for genetic risk factors. https://doi.org/10.5281/zenodo.8154493
More information:
Folkhälsan
FinnDiane
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
Glaucoma is a progressive disease of the optic nerve that causes damage to the optic nerve head and nerve fibre layer. Artificial intelligence models are currently being developed for early detection of glaucoma.
Researcher and project manager Ara Taalas specialises in data science, artificial intelligence and machine learning algorithms in medicine. One of his research objectives, in a joint project involving the Institute for Molecular Medicine Finland (FIMM) and Terveystalo health clinic, is to develop effective learning algorithms for glaucoma detection. Previously, Taalas modelled stem cell differentiation processes and worked in drug design.
When developing the artificial intelligence model, Ara Taalas focused on how the nerve layers of the fundus appear in the photographs. The algorithm will help to detect changes in the fundus pictures that can indicate damage to the nerve fibre layer.
Taalas uses the computing services of Finland’s ELIXIR node CSC. He develops models together with researchers in FIMM’s Machine Learning in Biomedicine team, and the same source code can be used on the computing servers of both CSC and Terveystalo.
“Finland is at a high level in terms of data management, but individual healthcare actors typically do not have a comprehensive picture of their patients – patient data is often scattered between various service providers. When customers go to a different organisation, the data does not follow them, which may make diagnosis and treatment more difficult. From the viewpoint of a researcher, the ideal thing would be to have a site for the entire country where each patient’s medical history could be found in its entirety.”
Data description should also be standardised.
“The structure of patient data systems has a major effect on the usability of any data entered into it. Fields where data can be entered in free form may be convenient for the person typing it in, but cause a lot of trouble to data analysts when trying to utilise it. Analysts often have to do a lot of work to standardise the data and to identify entries that contain errors. Modern patient data systems have in this respect become better in that they are much more structured.”
Read more here
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The six-year BIGPICTURE project that began in February 2021 will collect three million scanned, digital pathology slides from various European hospitals, research organisations and pharmaceutical companies. One of the participating scientists is pathologist Yossra HS Zidi-Mouaffak, co-coordinator of BIGPICTURE’s Finnish node. She is focusing on how artificial intelligence can be used in pathology.
The project is participated in by 45 organisations from 15 countries. Finland’s contribution to the project is provided by Helsinki University Hospital (HUS), mainly through the Helsinki Biobank, and CSC – IT Center for Science. The BIGPICTURE platform is being built collectively by pathologists, researchers, AI developers, patient advocates, and industry representatives. The files are stored in a repository, which enables the creation of new, efficient AI applications to promote digitalization of diagnostic pathology and to bring novel methods for tissue analyses. Samples can be analysed with artificial intelligence.
Yossra HS Zidi-Mouaffak currently works as a pathologist for Helsinki Biobank (HUS) and is also a PhD student and researcher at the University of Helsinki in Professor Olli Carpén’s research group. One of Yossra Zidi-Mouaffak’s projects involves digital pathology and colorectal cancer.
“Colorectal cancer (CRC) is the second most deadly and the third most commonly diagnosed cancer in the world. It is also the second most common type of cancer in Finland. Most of the CRC patients are treated with surgery and oncological treatments depending on the stage of the disease,” says Zidi-Mouaffak.
Oncological treatments can involve chemotherapy and radiation therapy.
”In our project, we are focusing on a particular set of patients with stage II colorectal cancer for whom the risk benefit ratio of adjuvant chemotherapy in often marginal.”

Stage II colorectal cancer is considered as an early stage of the disease where tumor invasion remains “local” without metastatic dissemination to other distant parts of the body. The tumor will penetrate through the entire intestinal wall and may also extend to adipose tissue or an adjacent organ, but it does not yet spread to lymph nodes or to distant organs. About 75% of patients with stage II will remain cancer-free 5 years after surgery.
“Unfortunately, 25% of the patients will not and these patients could benefit from post-operative chemotherapy”, says Zidi-Mouaffak.
“The question thus is: how to assess which patients are at high risk of recurrence?” Our project’s ultimate aim is to provide a predictive tool in stage II colorectal cancer, requiring ideally a considerable amount of data and images for more reliable results. BIGPICTURE is providing both large amounts of data and AI tools for researchers. This obviously makes progress much faster in this area of research.”
Zidi-Mouaffak selects, annotates and analyses scanned microscope images obtained from cancer patients’ surgical tissue samples stained with Hematoxylin and Eosin (H&E). This results in tissue parts dyed according to their PH. H&E is a routine stain for pathologists that allows them to analyse through the microscope, the morphology of the cells as well as the other components of the tissue.
Two Finnish biobanks, Auria and Helsinki Biobank, are among institutions providing datasets, which include whole slide images and associated curated metadata. Such datasets are used to create machine learning models by means of convolutional neural networks. The artificial intelligence models analyse the images that have been previously selected and annotated.
“As a pathologist, I believe that machine learning has the potential of improving pathologists’ output. The machine learning algorithms can be used as diagnostic tools to achieve routine tasks where they would be obviously faster, and more accurate than a human eye.”

Zidi-Mouaffak gives a few simple examples of AI diagnostic tools: recognize and count cell divisions (called mitosis), count the number of certain immune cells in specific areas, or accurately assess cell proliferation indexes.
“However, AI tools used for making predictions of a disease’s outcome based on image data, are very challenging to develop. They still require long phases of testing and validation before they can actually be used in clinical practice.”

The BIGPICTURE project first creates a storage infrastructure that enables processing, storage and sharing of extremely large image files. Pathological images may be up to several gigabytes in size. Slide images are provided with metadata. This material can be used to develop artificial intelligence tools, such as algorithms. Deep-learning algorithms are taught to classify morphologically similar cohorts, that is, they analyse shapes and structures in the samples. Artificial intelligence is able to detect cancer signs, or biomarkers, and they can then be verified.
“Based on recent studies, we believe that artificial intelligence applied on pre-selected digital slides from well-curated cohorts, could provide an interesting alternative to the existing molecular and morphological predictive markers.”
The purpose of the research team that Zidi-Mouaffak is part of is to develop and verify a new, “predictive marker” that could facilitate the stratification of stage II colorectal cancer patients. The focus is on the morphological features of the tumour.
According to Zidi-Mouaffak, deep-learning algorithms can make surprisingly accurate predictions of certain types of cancer, but in many cases it is not known how the algorithm reaches its decision.
“It is a sort of black box. This clearly deserves more research, and here is where repositories like the ones developed by BIGPICTURE, become extremely relevant. This kind of research requires huge databases with very big numbers of high quality digital slides and metadata, which is the aim of BIGPICTURE.”
Ari Turunen
10.2.2022
Read article in PDF
Citation
Ari Turunen, Yossra HS Zidi-Mouaffak, & Tommi Nyrönen. (2022). BIGPICTURE helps pathology go digital. https://doi.org/10.5281/zenodo.8154477
More information:
BIGPICTURE
HUS Helsinki University Hospital
Helsinki Biobank
https://www.helsinginbiopankki.fi/en/
Auria Biobank
https://www.auria.fi/biopankki/en/
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org

Sharing biomedical data collected of humans is a prerequisite for disease prevention and treatment in the modern world. The Finnish ELIXIR node CSC is building an infrastructure in which human data obtained from Finland’s biobanks and research organisations has been pre-processed and described and saved in a secure way. The parties responsible for sharing the data can automate their authorisation process with the CSC platform. This improves licensed availability of data for research and healthcare purposes.
Personalised drug treatments are only possible if patient data is available and it has been stored and pre-processed correctly. In a project funded by the Academy of Finland, an infrastructure is created that meets the requirements for storing and using sensitive data. The data consists of clinical register data, genomic data and material related to bioimaging. The project is participated in by not only CSC but also the biomaging infrastructure Euro-BioImaging, THL Biobank and the Institute for Molecular Medicine Finland (FIMM).
The project creates solutions to facilitate quick and easy access to various data for by researchers. The data can be stored in CSC’s sensitive data infrastructure. Researchers are allocated a space in which the data and computing power are in the same place. Researcher can only access data to which authorisation has been obtained by the data owner. The project also makes use of federated data management developed by CSC. ELIXIR AAI and REMS are applications developed by CSC for the managing users in the ELIXIR infrastructure.
Secure transfer of data will revolutionise healthcare in the next decades. The project supports researchers developing artificial intelligence algorithms by offering them computing services, more advanced research use of health data, and data management technologies. The data material’s compatibility with international standards is also verified.

The sequencing capacity of the Institute for Molecular Medicine Finland and Helsinki University Central Hospital is improved with a direct connection to CSC’s computing and data services. Genomic data is transferred to CSC on a superfast and secure optic cable. Data pre-processing and quality assurance are fast, because the data is located at CSC.
As the sequence data is physically closer to the computing services, the pre-processed data will be available to the researcher more quickly. The capacity can be used to sequence exomes, genomes and transcriptomes efficiently.
Combining genetic and clinical data still requires a lot of data storage and computing capacity. The European HPC Center of Excellence for Personalised Medicine (PerMedCoE), a joint project by CSC and the Barcelona Supercomputing Center (BSC), brought the data analysis methods of personalised medicine into the supercomputer environment. Algorithms developed in the project can significantly reduce the computing time required for analysis. Analysis of genetic and protein data is becoming faster, facilitating and speeding up disease diagnosis and identification of the appropriate treatments. Disease diagnosis by utilising molecular biology can in future be done within hours or days.

CSC, together with the Finnish biobanks, the National Institute for Health and Welfare and Euro-BioImaging, which operates from Turku, are developing an artificial intelligence algorithm for mining medical data.
Euro-BioImaging Finland offers image storing services and data services, such as image collections. Terabytes of images have been stored in the collections, and these can be used as reference data, for example. The material ranges from plankton imaging to cancer cells.
Euro-BioImaging Finland also offers medical imaging material. Free access to imaging services is provided by six universities and three university hospitals in Finland. These use Open Microscopy Environment (OMERO) services, enabling researchers to view, organise, analyse and share material from anywhere with internet access.
”Turku already has two new OMERO services in production use for image data, one for research and the other for teaching purposes. Both also serve, to a limited extent, the entire country. Now would be a good time to plan how these could be linked with CSC services,” says Pasi Kankaanpää, Senior Scientific Manager at Euro-BioImaging.
Kankaanpää has submitted articles to the Nature Methods publication concerning recommendations for image data management and its metadata.
“This increases cooperation and also emphasises the importance of managing sensitive data. Data management and processing are key aspects at Euro-BioImaging Finland – and indeed what this project funded by the Academy of Finland emphasises,” says Kankaanpää.
At the moment, the transfer and utilisation of genomic data does not work across borders. CSC is developing standards for genomic data technologies (such as GA4GH.org Passport, Cloud, Beacon), which are also relevant outside Europe, such as North America, Japan and Australia. The purpose of the ELIXIR infrastructure is to adopt global standards for the responsible sharing of genomic data. Europe also has a strong desire to create a federated data security infrastructure for sensitive genomic data. The plan is to create what is known as European Health Data Space (EHDS).
“ELIXIR has been developing good tools for a long time for researchers – improving usability by creating new tools. ELIXIR’s cooperation with the Global Alliance for Genomic Health has created a fine vision on how this global cooperation could work, and also concrete tools and models,” says THL Biobank’s Director Sirpa Soini.
The aim is to make biobanks operate in a compatible, federated data infrastructure that transcends national borders. This is connected to the ‘1+million genomes’ and ‘Beyond million genomes’ projects funded the EU member states and the Commission. In the ‘Beyond million genomes’ project, CSC is in charge of the technical infrastructure work.
THL Biobank’s part in the project is to design management processes for national health data for research. The objective is to enable researchers and students easier access to material in Finnish biobanks. This would also mean that data could be transferred securely from biobanks to CSC’s sensitive data environment and sharing it with those who have been authorised to access it.
Sirpa Soini is very well aware of the concerns and regulations concerning the use of sensitive data. She nevertheless feels that GDPR is too often blamed for any problems, although it is in fact many member states that themselves restrict the transfer of sensitive data in their own legislation or interpretations. Soini is also a lawyer by training and thinks that many issues can be solved provided there is enough political will.
“At the moment it seems that people are simply saying that various things cannot be done because of GDPR. But that’s not the real reason. It not the reason in Finland or elsewhere, and solutions are available.”
According to Soini, GDPR does not restrict data use, but in fact enables it, but in a responsible way, and taking account of the risks. National legislation is required to support certain use cases.
According to Soini, in secondary use of data it is difficult to predict subsequent use. But in cases like this, the premise should be that medical and applied research and product development is possible under GDPR, based on the law.
“This would mean that consent would not have to be obtained. Our law prescribes use for the general good, complete with the appropriate data protection and data security measures. You do not need full, detailed consent as such, although transparency should be promoted.”
She also says that there are no absolute legal obstacles for transferring data abroad. THL Biobank, for example, has made agreements about data transfer to the United States and Australia.
“I suggested a cooperation agreement to the US and Australian lawyers, emphasising which responsibilities each partner has in terms of risk management. It is important that the agreements have precise restrictions and that the material is pseudonymised. It is also always specified where the data can be stored.”
One such place could be the European Genome-phenome Archive (EGA). To protect the identities of data providers, data made available for research has been pseudonymised. Only an authorised party, such as the Finnish Institute for Health and Welfare, may decrypt the pseudonymisation.
Soini speaks of a dream cloud in which the data itself would not be moving.
“Data could be stored securely in an international database. Direct searchers and identification would be possible within the Trust Federation Network, provided the datasets were ready. This would put the controller in control of its data, assessing requests to use the register. In an ideal situation, permits could cover several datasets around the world, in effect creating a type of federated solution: the data itself would not more anywhere, rather the researcher would be given access to a “dream cloud”. This could be accessed by researchers from various locations.”
Ari Turunen
30.12.2021
Read article in PDF
Citation
Ari Turunen, Pasi Kankaanpää, Sirpa Soini, & Tommi Nyrönen. (2021). Sensitive data infrastructure. https://doi.org/10.5281/zenodo.8135532
For more information:
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org

Glaucoma is a progressive disease of the optic nerve that causes damage to the optic nerve head and nerve fibre layer. The risk of developing glaucoma increases with age. Some 2% of people over the age of 50 have glaucoma, and more than 5% of those over 75. There are estimated to be more than 60 million patients with glaucoma globally, of which around 6 million are categorised as being visually handicapped.
The challenging thing about glaucoma is that in its early stages it exhibits no or very few symptoms. Early diagnosis is very important because any damage that has already occurred cannot be reversed. The objective with treatment is to prevent any visual handicap caused by glaucoma. With most patients, the condition advances gradually over many years. However, with a small percentage of patients, the disease may lead to damage in a shorter period.
For purposes of glaucoma detection and identification of progressing speed, it would be best if healthcare systems found the high-risk cases as early as possible. Artificial intelligence models are currently being developed for early detection of glaucoma.
Researcher and project manager Ara Taalas specialises in data science, artificial intelligence and machine learning algorithms in medicine. One of his research objectives, in a joint project involving the Institute for Molecular Medicine Finland (FIMM) and Terveystalo health clinic, is to develop effective learning algorithms for glaucoma detection. Previously, Taalas modelled stem cell differentiation processes and worked in drug design.
According to Matti Seppänen, chief physician and Head of Ophthalmology at Terveystalo health clinic, glaucoma diagnosis and classification are based on the examination of the optic nerve head, nerve fibre layer and anterior chamber angle, intraocular pressure measurement, and a visual field test.
“The pathogenesis of glaucoma is not known, but damage to nerve cell structures probably contributes to glaucoma damage.”
Probably some 30–50 per cent of patients have intraocular pressure which is considered normal (10–21 mmHg). Patients have an individual susceptibility to the development of glaucoma damage at different intraocular pressure levels. Some patients develop glaucoma damage at lower pressure levels, while other patients may have minor damage even at higher pressure levels.
“At the moment, a glaucoma diagnosis requires examination by an ophthalmologist and several additional examinations. The optic nerve head can be examined by means of, for example, biomicroscopy and stereo papilla photography. The nerve fibre layer can be examined with colour fundus photography or optical coherence tomography (OCT) of the nerve fibre layer.”
During an examination, glaucoma may be suspected on the basis of the shape of the optic nerve head, for example. The structure of the optic nerve head can be evaluated with a measurement of the cup/disc ratio, meaning that the size of the optic nerve cup is compared to the size of the outer edge of the optic nerve head.
“Damage to the nerve fibre layer may show in an OCT examination as a thinned nerve fibre layer. In colour fundus photography, defects in the nerve fibre layer may also be discovered. A glaucoma diagnosis is often based on several examinations, and currently there is no single method for screening the entire population for glaucoma. Artificial intelligence applications may in the future bring considerable help for screening and diagnostics.”
Esa Pitkänen from the Institute for Molecular Medicine Finland FIMM (University of Helsinki) tells how glaucoma can be studied with the help of algorithms.
When developing the artificial intelligence model, Ara Taalas focused on how the nerve layers of the fundus appear in the photographs. The algorithm will help to detect changes in the fundus pictures that can indicate damage to the nerve fibre layer. The purpose of the model is to find out whether subtle changes in the network in the fundus, as they become darker and more monotonous, can be linked to damage in the nerve fibre layer.
“This is one of the factors the model is designed to focus on. In the future, the model will be taught more nerve fibre patterns in the fundus. The purpose of these algorithms is to find ways that will help doctors to make decisions. And advanced artificial intelligence system may detect changes that not even the most experienced clinician can see.”

Examinations on the eye’s structure and operation involves variations caused by the examination method used, experience of the person assessing the case, and the patient and how serious the disease is. Evaluating the optic nerve head does not always result in sufficient accuracy using the current methods. The result of the visual field examination may be normal even if the optic nerve and nerve fibre layers have been damaged. This is because structural damage usually occurs before any visual field defects. If we are able to develop applications that examine more accurately and more efficiently any structural changes, glaucoma diagnoses may be made earlier.
According to Taalas, one application for the model could be that the artificial intelligence model is always used when performing an eye examination.

“Population surveys have found that up to half of those who have glaucoma have not actually been diagnosed with it. The existing screening methods are not cost-effective enough and a general screening of the population cannot be done for lack of sufficiently good methods. If artificial intelligence applications were able to identify with sufficient accuracy patients that have a higher-than-average susceptibility to develop glaucoma, it would be easier to screen out those among the population that do not yet have the symptoms and offer them early treatment for best results.”
One of the future versions is that during a visit to the opticians or healthcare worker, the examination would include fundus photography, and at the same time artificial intelligence would analyse the patient’s fundus photo. If artificial intelligence indicated that the patient had a higher risk than average to develop glaucoma, the patient would be referred to further examinations at an early stage.
With artificial intelligence applications, the division of work would probably change dramatically in the optical field and the diagnosis of eye diseases. This would also result in significantly higher numbers of patients being treated. As the age structure of the population is changing, the number of glaucoma patients in Finland will double from the current figures by 2030.
Taalas uses the computing services of Finland’s ELIXIR Center CSC. He develops models together with researchers in FIMM’s Machine Learning in Biomedicine team, and the same source code can be used on the computing servers of both CSC and Terveystalo.
“Finland is at a high level in terms of data management, but individual healthcare actors typically do not have a comprehensive picture of their patients – patient data is often scattered between various service providers. When customers go to a different organisation, the data does not follow them, which may make diagnosis and treatment more difficult. From the viewpoint of a researcher, the ideal thing would be to have a site for the entire country where each patient’s medical history could be found in its entirety.”
Data description should also be standardised.
“The structure of patient data systems has a major effect on the usability of any data entered into it. Fields where data can be entered in free form may be convenient for the person typing it in, but cause a lot of trouble to data analysts when trying to utilise it. Analysts often have to do a lot of work to standardise the data and to identify entries that contain errors. Modern patient data systems have in this respect become better in that they are much more structured.”
Ari Turunen
23.11.2021
Read article in PDF
Citation
Ari Turunen, Lila Kallio, Arho Virkki, & Tommi Nyrönen. (2021). Patient data creating better artificial intelligence models. https://doi.org/10.5281/zenodo.8135413
More information:
Institute for Molecular Medicine Finland FIMM, University of Helsinki
Terveystalo
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Medical research will not progress without data and its re-use. Collected data can be used to create artificial intelligence models to make speedier diagnoses and to support care decisions. New data analysis technologies emerge all the time, but how to make data available for all researchers?
One of the strengths of the Genome Center, which is being established in Finland, consists of biobank databases. The Center would be in charge of developing a genome data register, that is, a centralised system for storing and managing genetic data. The aim is to create a high-quality database that describes genetic variation in Finland. Auria Biobank’s Director Lila Kallio believes that good cooperation between the biobanks and the Genome Center can lead to significant results in the screening of gene variants.
“Once the Genome Center has been established and starts operating, genome data created during research can also be stored in the Genome Center. The Genome Center could re-analyse any genome data against all accumulating reference genome data. This would enable, for example, the screening of new identified, clinically significant variants by using previously produced and stored data,” says Lila Kallio.
The Biobank Act was enacted in Finland in 2013, enabling the establishment of biobanks. There are currently 11 biobanks in Finland. In 2020, the biobank network was joined by Arctic Biobank, which stores extensive population data collected by the University of Oulu in the north of Finland. Researchers in Finland can utilise material from all biobanks through the Fingenious online service. Fingenious is a digital tool through which a researcher can send a request for material to be made available to them. This service is provided by the Finnish Biobank Cooperative, FINBB.
“Biobanks store data about samples securely. Data about biobank samples is available to all researchers. A researcher must present a research plan for approval by the biobank steering groups or ethical committee. Biobanks have a process in place for the research use of samples and their related data.”
Finland has exceptionally comprehensive and high-quality health care data resources. The Act on the Secondary Use of Health and Social Data (552/2019) came into effect in 2019 in Finland. Secondary use of data means that customer and register data within social care and healthcare are used for some other than the original purpose. This act on secondary use has also created pressure to amend the Biobank Act from 2013. The significance of data in biomedical research is increasing and the legislation should create the conditions for both research and appropriate data security.
Secondary use obviously requires that data collected of people is managed securely. Identifier data of human samples stored in the biobanks is carefully protected.
“The biobanks remove all personal identifiers and replace them with pseudonym codes. When samples are handed over for research purposes, the pseudonyms are replaced with another code, specific to that particular research. The code key is stored in the biobank. If you need to access the original sample owing to, for example, by some clinically significant detail, this can only be done with the code key,” says Kallio.
The use of a code key enables the data to be re-used in subsequent more research purposes.
“If the sample was anonymised, that is, making impossible to identify, it would be impossible to access it after any findings in biobank research, and no sample-specific data could be added to it later.”
According to Lila Kallio, the real value of samples is in the data created from it.
“Data is created during diagnostics and treatment. Research also results in analysed data, which must be returned to the biobank in possession of the sample, to be appended to it. Biobanks manage not only identifier data but also clinical data and data that has been produced during research.”
The act on the secondary use of health and social data concentrated the permit process management to Findata, a new legal authority. Lengthy processing times of permit applications has become a problem. All applicants are treated equally regardless of the size of material they are making an application for.
Auria’s chief data officer (CDO) and adjunct professor of medical mathematics Arho Virkki points out that material can be used in a number of ways, and that’s why there should be different protection levels based on the purpose of use. According to Virkki, the initially planned data security improvement leap for the secondary use of health data is too great to be taken in one go.
“Extreme protection weakens data availability, which means that data security won’t reach an optimal level. To me, optimal data security means that material is truly available for scientific research, planning of new treatments and controlling treatment processes. In the optimal situation, data is available and at the same time it is adequately protected. The protection level should be set on the basis of the risks involved.”
Because data management is part of the work of doctors and nurses, Virkki says a balance should be found between material availability and its protection. At the moment, the system is lopsided.
“For example, exploring clinical data is part of medical students’ curriculum. One part of their training is the use operational systems to find data in order to learn.”
According to Virkki, the isolated data architecture has been the culprit for a long time. Owing to a defensive approach and regulation in medicine and healthcare, information architecture is more traditional compared to, for example, logistics and the financial sector. This is why various information systems are poorly integrated.
Virkki does admit, however, that hospitals are more complex places than logistics centres, for example. In logistics systems, postal packages follow a pre-defined route and their travel is easily recorded by the system, whereas when a patient arrives at a hospital, the following steps are typically more complex and involve a great number of different options.
The act on secondary use of data, however, makes the assumption that one type of secure data processing environment fits all uses. Virkki says that the legal entity issuing the research permit could provide a range of user environments based on the researchers’ needs.
“There could be a basic environment sufficient for simple data analysis that consists of spreadsheet software and the usual range of statistical programming languages.”
Then again, if researchers need a custom environment, they should be given exact details about data security with assurances that they will comply with them.
“This way, the authorities would set the requirements for data security, but the researchers should be accountable of it, as has been the case up till now. At the end of the day, it is the researchers’ responsibility to ensure that their results are correct, honest, scientific and anonymous.”
According to Virkki, people in the medical field in Finland take great pride in their work and have always strived to process medical material in a proper fashion. Virkki says that data security can be ensured through licensing and training. Data security issues should be part of medical training. Virkki is a regular speaker at the University of Turku on an introductory course on the basics of clinical research about data platforms and data security.
According to Virkki, amendments are in progress for the act on secondary use of data. If the provisions can be made more flexible and the permit processes faster, there will be many opportunities for artificial intelligence research.
“Now that the reform on social care and healthcare was passed in Finland, there is a good possibility to combine the patient data of basic and specialised health care, that is, view patient data as a single entity. This in turn will enable the development of new AI applications for the clinical side.”
The algorithms of AI models can perform text-based analyses, make medical records or learn to identify details in images that can be used in diagnoses.
“In fact, artificial intelligence is just modern statistics, a refined branch of mathematical statistics. AI models make use of complex statistical methods. When you talk of machine learning, you actually mean statistical learning. Today you can calculate predictions in such a precision that is almost feels like magic.”
Virkki has been intrigued by AI models for a long time. In his doctoral thesis, he created an AI model for human respiratory system during sleep. Recently he has been developing a prediction model for pulmonary embolism. The model is used as a tools for decision-making. Pulmonary embolism occurs when a blood clot gets wedged into an artery in the lungs. The most common symptom is sudden shortness of breath. In serious cases of pulmonary embolism, the clot is diluted by injecting an anticoagulant into a vein.
“If there is reason to suspect that a patient in an emergency room has pulmonary embolism, you have to act fast. A machine can quickly go through a set of scanned images and inform the radiologist where they should focus on in any image. After that, the decision is made whether to start diluting or not. If not, another treatment is chosen. You should be able to do all the following in less than 10 minutes: lung imaging, diagnosis and starting the treatment.”
According to Virkki, the pulmonary embolism model was the first scientific test trying to solve a difficult problem with very little data. However, a more comprehensive and more accurate AI model is under development. Scientific publications and dissertations will be published on the subject.
“If realised, the model will speed up decision-making in case situations, but also assist in quality control. For example, we can screen afterwards whether we detected any smaller cases of pulmonary embolism.”
The development of artificial intelligence models requires a lot of data with which algorithms are taught, and plenty of computing power.
The hospital district of Finland Proper uses the ePouta cloud service of Finland’s ELIXIR centre’s CSC, with a dedicated 10 GiB connection. Virkki hopes researchers could have better access to the ELIXIR network.
“It would be great if researchers were given capacity from the ELIXIR infrastructure for their work. The data resource would be made available directly in the ELIXIR environment, and ELIXIR would ensure there was enough computing capacity.”
The ELIXIR Node in Finland (ELIXIR-FI) is hosted at the CSC – IT Center for Science Ltd. CSC operates resources and services that are part of ELIXIR, like pan-European ELIXIR identity and access infrastructure. In ELIXIR the data needs to be managed as a federation, where data providers work as a single infrastructure providing mechanisms where researchers can bring their analysis to where the data is located. The ELIXIR Compute Platform infrastructure will allow life scientists to easily access, share and analyse data from different sources across Europe. The objective is to combine all components of the ELIXIR Compute services into a seamless workflow. A researcher may use the ELIXIR Authorisation and Authentication services to securely create a scientific software analysis environment, and use the environment to access large biological data resources stored in a cloud.
Text written or dictated by a doctor can be utilised in artificial intelligent models used in aid of current care guidelines and diagnoses. Statements and sentences can be constructed into data and teach the algorithm to make deductions. In the project participated in by Auria Biobank, Turku University Hospital and the University of Turku, artificial intelligence was taught to extract data about smoking from about 30,000 patient records. In the project, headed by researcher Antti Karlsson, a language model called ULMFiT was used. The model was trained using the analysis computers of Finland Proper Hospital District, making use of texts from Wikipedia. After this, the model was trained to become a classifier by means of some 5,000 manually annotated sentences. These days there are also more sophisticated, pre-trained models in Finnish, most notably perhaps FinBERT, based on the Google BERT model. This was created by a University of Turku research group led by Filip Ginter, using computing power from Finland’s ELIXIR centre CSC.
Based on the data collected by the artificial intelligence model, the study showed that quitting smoking, even once a cancer diagnosis has been made, may extend the patient’s life expectancy considerably.
“I’m sure that future patient record systems will not be formal, with items picked from drop-down menus, but written more in prose, with systems that automatically structure them,” says Karlsson.
“This also improves work efficiency. I don’t even want to imagine how difficult it must be for a busy doctor to enter complex matters into the systems.”
When you mine a large data mass, you save an awful lot of time and money. The artificial intelligence model trained by Antti Karlsson analysed patient records, searching for smoking-related issues. In the study, the model analysed text date obtained from the records of 30,000 patients. According to Karlsson, models like this can produce analyses that are more than 90% accurate, in hours or even minutes. It’s quite different from manually reading through the texts of 30,000 patients, entering variables into a table.
“In the best-case scenario, such models could be readily available in a data lake, structuring this tobacco data automatically for research purposes,” says Karlsson.
The model does not produce treatment instructions for an individual patient, but creates a good overall picture.
“A believe that at least initially, the automated systems of the future will collect data relevant to reporting and research, while the really important things, such as dosages and allergies must still be checked and filled in manually by experts.”
Ari Turunen
26.10.2021
Read article in PDF
Citation
Ari Turunen, Pasi Kankaanpää, Sirpa Soini, & Tommi Nyrönen. (2021). Sensitive data infrastructure. https://doi.org/10.5281/zenodo.8135532
More information:
Karlsson et al. (2021): Impact of deep learning-determined smoking status on mortality of cancer patients: never too late to quit. Esmo Open Cancer Horizons. Vol 3. Issue 3.
https://www.esmoopen.com/article/S2059-7029(21)00135-6/fulltext
Auria Biobank
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Bio- logy Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.
https://www.elixir-finland.org

Deep learning has revolutionised cancer research. Deep neural networks can automatically detect features within a patient’s sample data that can be used to identify cancers. In the future, learning algorithms will be able to identify potential early-stage cancers from a blood sample. Esa Pitkänen and his research group at the Institute for Molecular Medicine Finland (FIMM) are developing a new generation of deep-learning algorithms.
Algorithms have been used to identify cells in sectional images of tissue samples. For instance, if tissue cells appear atypical, the algorithm will spot this and determine if the cells are cancerous. DNA sequence data from tumours is now being used along with imaging data to identify cancers.
“Until recently, it was difficult to tell from a DNA sequence what kind of tumour an identified sequence came from. Now new technologies and deep learning algorithms have been created,” Pitkänen says.
Pitkänen and his team are developing algorithms that identify short, repetitive snippets of DNA sequences. These algorithms can be used to find DNA sequences that mutate frequently in a particular type of cancer or to which certain proteins involved in gene regulation bind. Analysis of these sequences can be used for various purposes, such as charting the causes of cancer and developing medicines.
“The replication of DNA in cell division is not perfect; mutations can occur during the process. The division of a single cell involves the copying of about six billion nucleotide pairs in DNA, so errors will inevitably occur. Even the slightest probability of errors is enough to guarantee mutations,” says Pitkänen.
“If enough mutations occur in genes that prevent tumour growth, for example, cancer may start to develop.”
An example of this is a point mutation in which one base within the DNA strand is replaced with another. The enzymes involved in copying DNA may make a mistake when a cell is dividing, for instance by incorrectly repairing the part of DNA that was damaged by ultraviolet radiation from sunlight. A typical mutation caused by ultraviolet radiation that can result in skin cancer is that two consecutive cytosine (C) base molecules in the base pairs of human DNA are converted to two thymine (T) base molecules. When skin cancer-specific mutations of this type are detected in sufficient numbers, the algorithms can learn to associate them with a particular type of cancer.
“We try to predict the type of cancer and tumour from the mutations. This also provides information on how treatment can be developed.”
Pitkänen and his group analyse blocks of sequences and train algorithms to pinpoint deviations in them. From these abnormalities, the algorithm can detect tumours and classify them into different cancer types.
“Before joining the Institute for Molecular Medicine Finland, I worked at the European Molecular Biology Laboratory in Heidelberg, where I participated in the Pan-Cancer Analysis of Whole Genomes (PCAWG) project. This project involved the analysis of more than 2,600 entire cancer genomes. My group is using data from the PCAWG project in several of our cancer genomics projects.”
An algorithm developed by the group has been taught the mutations found in the tumour samples of 2,600 cancer patients. This sample set contained about 47 million mutations. Approximately 50 million somatic mutations were found in the sequence data.
“We trained the algorithm to try to deduce the type of cancer from these sequence changes. Once the algorithm is given all the mutations of different tumours and their sequences, in the future it will be able to determine the kind of tumour that has been detected. This deduction process is based on the algorithm learning these connections.”
Through deviations in the sequence data in tumours, the algorithm learns to identify when a given tumour corresponds to a particular type of cancer. It can group tumours based on sequence data alone.
“A researcher in my group, Prima Sanjaya, has developed neural network models for analysing sequence data. Every now and then, researchers come across cases in which a cancer has metastasised without being able to tell where it has spread from. Such cases could in the future be dealt with by means of a liquid biopsy – that is, it will hopefully be possible to determine from a blood sample if the patient has cancer, and if so, what kind.”

Liquid biopsies are based on the fact that cells release into the bloodstream and other body fluids a type of DNA called cell-free DNA (cfDNA). Cancer cells also release DNA, which makes it possible to test for cancer mutations in the blood plasma.
“If a liquid biopsy shows traces of cancer, we don’t know exactly what kind of cancer it is, as it could have entered the bloodstream from anywhere in the body. If we have the means to examine these cases more closely, for example with deep learning algorithms, we could obtain valuable information on where in the patient’s body the tests should be focused. The algorithm may indicate that the source of the cancer DNA could be the large intestine, for example. I believe that such algorithms will become extremely important. Liquid biopsy and algorithms can make it possible to diagnose cancers without surgery.”
In addition to hereditary factors, the development of cancer also depends on the person’s lifestyle. Plentiful research has been conducted at the University of Helsinki on various cancers, such as intestinal cancers.
“It is known that eating red meat contributes to the incidence of colorectal cancer. The mechanisms by which the disease is caused require further research, but in recent years a lot of progress has been made in understanding the significance of DNA alkylation reactions, which are caused by red meat, for example.”
Colorectal cancer is one of the most dangerous cancers in Western countries. In countries such as Finland, it leads to death in 30 per cent of cases. About 15 per cent of colorectal cancers belong to the cancer group that exhibits microsatellite instability (MSI). Microsatellites are sequences of DNA that can vary in length from person to person, and thus function as individual identifiers in much the same way as fingerprints. Microsatellite instability occurs when the post-replication repair mechanism of cellular DNA does not function properly. This causes mutations to begin to accumulate, especially in microsatellites.
“In an MSI tumour, microsatellites are easily vulnerable to single-base additions or deletions. For example, out of eight consecutive adenine microsatellites, one adenine may be lost. When it occurs in a gene, such a change causes a complete transformation in the content of the amino acid chain of the protein that is encoded by the gene. If there are enough changes in genes that are important for preventing uncontrolled cell growth, cancer may begin to develop.”
MSI is often associated with other cancers as well as colorectal cancer, such as stomach, uterine, ovarian and brain cancer. MSI analysis can be used in the prognosis of cancer. The treatment choice may be influenced by the analysis.
“An interesting thing is that the deep neural network is also learning to classify different subtypes of cancers. For instance, it identified the MSI subtype of colorectal cancers,” says Pitkänen.
The ELIXIR Node in Finland, hosted by CSC – IT Center for Science, is one of the main partners in the Personalised Medicine in Europe (PerMedCoE) project. For example, the three-year the HPC/Exascale Centre of Excellence for Personalised Medicine in Europe project is aimed at making effective use of cancer-related data in healthcare and speeding up the process of diagnosis
“Individualised treatments of the future, among them cancer treatments, will be based on a precise understanding of the patient and their illness. This will result from gathering a large volume of data of different types, such as tumour-related data and imaging data during cancer treatment. Many data collection methods produce a mass of data, and the new computational methods developed for analysing it require a very large amount of computational resources,” says Pitkänen.
“Developing a new computational method from the idea stage into a functional healthcare technology is a huge challenge in an operating environment like this. With cancer treatments in particular, it is important that information relevant to patient care be made available to the doctors as rapidly as possible. I’m confident that the results of the PerMedCoE project will provide a basis for deriving relevant information from a colossal volume of health data to help doctors in their work, and thus significantly improve treatment outcomes.”
Ari Turunen
16.9.2021
Read article in PDF
Citation
Ari Turunen, Esa Pitkänen, & Tommi Nyrönen. (2023). Teaching an algorithm to identify cancer from sequence data. https://doi.org/10.5281/zenodo.8135303
1.External factors (e.g. UV radiation from sunlight), 2.Internal factors (e.g. a spontaneous deamination reaction, in which the amine group of a base changes, for example from adenine to uracil) 3. DNA copying errors.
A mutation is a change in the nucleotide sequence of DNA or RNA. A nucleotide consists of a base, a sugar molecule and a phosphate group. The sugar in DNA is deoxyribose and the sugar in RNA is ribose. The four nitrogenous bases that DNA contains are guanine (G), adenine (A), cytosine (C) and thymine (T). RNA has three of the same bases as DNA, but instead of thymine its fourth base is uracil (U).
A mutation may be a change of a single nucleotide – that is, a point mutation – or the change may involve multiple nucleotides. In a point mutation, one base is replaced by another in the RNA or DNA strand. Large mutations can involve thousands of nucleotides, and are called structural changes. A structural change can affect multiple genes at the same time. Cancers are usually caused by several somatic mutations. Mutations of this kind are not inherited, and can occur at any time of life from embryonic development onwards. Mutations may bring about a change in the functioning of a normal cell, causing it to begin to divide uncontrollably.
At the middle of the picture are presented different types of mutations, distribution of mutations on chromosomes and epigenetic information. Epigenetic inheritance is influenced by many external factors, such as nutrition. An example is the development of identical twins so as to become distinct from each other in appearance.
Modelling mutations:
Linear models
Deep neural networks
Transformer models. Transformers are a family of deep learning models that work particularly well with certain types of data, such as textual data. This makes them well suited to machine translation, for instance. In cancer research, transformer models can draw attention to mutation types that are important for identifying a particular type of cancer. For example, in skin cancers that contain many sunlight-induced mutations (C> T, CC> TT), the transformer will focus on these particular mutations.
Ari Turunen
For more information:
HPC/Exascale Centre of Excellence in Personalised Medicine (PerMedCoE)
Institute for Molecular Medicine Finland (FIMM)
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Next-generation analysis methods of genes and RNA molecules enable faster and easier analyses. Data can also be stored well and shared with research teams through the Allas user interface of CSC.
Next-generation sequencing (NGS) methods are used to study variations in the human genome and changes in genetic expression. The analysis of billions of sequence fragments provided by the NGS methods can be performed in a single computer run.
The new methods enable us to study numerous genes and targets from different samples simultaneously. This means that we can quickly analyse individual cells, such as cancer cells. We can also analyse cell-free DNA from blood plasma, indicating quickly and reliably whether the selected treatments have been effective and specifically whether any metastases remain.
Platforms used by the Institute for Molecular Medicine Finland (FIMM) and CSC use a range of algorithms to analyse data produced by sequencing methods (exomes, genomes and transcriptomes). One of the most important ones is Broad Institute’s Genome Analysis Toolkit (GATK). This is used to look for gene variants and identify changes in the DNA or RNA sequence in the cell line. GATK analysis software has become the de facto bioinformatics standard in the scientific community. GATK software can also be run on the superfast Dynamic Read Analysis for GENomics (Dragen) platform. The Finland ELIXIR Node CSC maintains Dragen, in collaboration with FIMM. Dragen perfroms the computing-intensive primary analysis of the sequence data, which is followed by the downstream bioinformatics analysis by the researchers. This make the CSC storage capacity beneficial, because analysed data will not fit into any conventional computer, instead it is shared directly to users through the Allas service. Cooperation between CSC and FIMM is crucial in terms of completing the analyses quickly.
“When we have at our disposal high-capacity sequencing platforms, algorithms and computing power, we get quick results. Today we can analyse one genome in a single day, as opposed to several weeks using older systems,” says Pekka Ellonen.
Mr Ellonen is Head of Laboratory at the Institute for Molecular Medicine Finland (FIMM). The unit uses modern methods to provide the research community genomics (DNA) and transcriptomics (RNA) analyses. The unit receives samples from various research projects.
“We agree with the researchers on the most suitable methods and customise the optimal toolkit to test their hypothesis. Such methods may include exome sequencing, genome sequencing, the sequencing of various RNA molecules (transcriptome), and genetic expression,” says Ellonen.
These methods are able to determine a tissue sample’s genes (genomics) or identify all genes (transcriptomics) and proteins (proteomics) present in the tissue. The sequencing of the exome, that is, the regions that code the proteins can help in the study of hereditary diseases, congenital developmental disorders and cancer. Genetic expression is regulated accurately in cells and any changes may lead to illness. The research can focus on, for example, the differences between cancerous and healthy tissue.

Next-generation sequencing methods enable the study of complex biological systems. According to Ellonen, by far the greatest development in bioinformatics in recent years has been the analysis of single cells. Single cells are analysed in collaboration by the Single-Cell Analytics (SCA) unit of the Institute for Molecular Medicine Finland and the sequencing unit.
Each cell contains the individual’s every gene, but certain genes are only expressed in certain cells and often only under certain conditions. Genetic expression and protein production in cells varies at different stages of development and as a result of illnesses. This causes changes in the cellular and tissue functions. The analytics of a single cell does not actually refer to a single cell.
“Now we are able to study, for example, cancer cells as individual targets. We cannot reach a reliable result merely by determining the base sequence or genetic expression in a single cell; we must study samples of thousands or tens of thousands of cells,” says Ellonen.
Single-cell RNA sequencing (scRNA-seq) can reveal regular inter-gene interaction, cell lineages and differences and the cell’s framework in its environment.
Single-cell sequencing also shows various and even new types of cell and genetic expression data about the functioning. Single-cell DNA sequencing, on the other hand, provides information about mutations taking place in small cell populations among normal cells. Single-cell accuracy provides information on the genetic differences of tumours, which is helpful in their treatment.
“The number of living cells in the sample being studied is verified in the laboratory, after which each cell is separated into its own droplet, enabling single-cell DNA or RNA molecules to be marked with molecule- and cell-specific DNA barcodes. The molecule-specific, cell-specific and eventually the sample-specific DNA barcodes enable both the identification of molecules in each cell and a financially efficient sequencing,” says Pirkko Mattila, head of the Single-Cell Analytics (SCA) unit of the Institute for Molecular Medicine Finland.
“One sequencing run will profile thousands of cells at a time from multiple samples. This results in, from the analysis of thousands or up to hundreds of thousands of cells, a single-cell resolution, enabling us to study the properties of a single cell.”
Liquid biopsy refers to taking a liquid sample, containing cells or parts of cells, from living tissue, such as blood. Liquid biopsy is a promising monitoring tool for cancer treatment without invasive surgical operations.
“We create sequencing libraries from genomic regions interesting from the viewpoint of various cancers,” says Pekka Ellonen.
Liquid biopsy can also be used for identifying cancer in its early stages. A blood sample provides information on tumour blood cells or DNA fragments they have secreted into the bloodstream.
“Tumours are usually in a difficult place, requiring a surgical procedure to remove them or take a sample of them. When tumours grow uncontrollably, there is a higher than normal amount of cell deaths. Dying cancer cells release DNA fragments into the bloodstream. These DNA fragments are collected for sequencing from a blood sample’s cell-free fraction, plasma and serum. Analysis of the sequencing results can show whether the bloodstream contains DNA fragments containing changes typical of cancers,” says Ellonen.

Ellonen says that liquid biopsy is used extensively and it is related to many new research projects. Liquid biopsy can be used not only in basic research but also to make a treatment plan and to monitor treatment effects or cancer recurrence. Being able to take many blood samples at different times will help doctors understand what kind of molecule changes have taken place in the body.
“New genetic markers may be identified and, in the best-case scenario, an accurate treatment method can be selected on the basis of the observed mutations. Alternatively, you may know what you are looking for, that is, you are monitoring for residual signs of the disease in the body, in other words whether the surgical procedure removed the cancer completely.”
Pekka Ellonen is enthusiastic about CSC´s Allas storage service’s user interface, enabling laboratories and research institutions to share pre-processed sequencing results and molecular data with researchers, research teams and consortia. Allas provides 12 petabytes of storage space. The data is securely available through the Web. Data processing can be performed using standard programming interfaces from anywhere.
“Public money produces data, which should be shared in time with the wider scientific community, obviously appropriately pseudonymised. The user interface enables the sharing of large materials, such as the cohort material of useful genomic data.”
Ari Turunen
3.12.2020
Read the article in PDF
Citation
Ari Turunen, Esa Pitkänen, & Tommi Nyrönen. (2020). Efficient processing and sharing of data improving disease diagnosis and treatment. https://doi.org/10.5281/zenodo.8135239
More information:
FIMM
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Data obtained from human genes and proteins enables quick diagnosis of illnesses and more detailed and personalised patient treatment. Those at risk can be screened better and more effective medication can be chosen once the patient’s genome is known. The challenge lies in how data is processed and where it is stored.
Owing to new data analysis methods and improved computing power, the processing of gene and protein data can be accelerated from several days to less than half an hour. But in order to do this, data must be pre-processed, that is, redundancies removed, and be quickly and securely available.
Bioinformatics measurement methods have developed apace, producing huge amounts of data. Now we are in a position to understand the operation of an entire biological system. This means that genes – or the proteins created by them – are studied simultaneously. These ‘omics’, if you like, include determining DNA sequences (genomics), computer modelling of protein structures (proteomics), study of genes in cell tissue (transcriptomics) and metabolic products (metabolomics). Such methods can be used to study molecular interaction and find signs of changes in the human body to identify diseases.
“We can already do a great deal, but combining genetic and clinical data requires a lot more data storage and computing power, and all this in a secure way,” says Katja Kivinen, Head of the FIMM Technology Center from the Institute for Molecular Medicine Finland (FIMM).

The proper pre-processing and modelling of data are requirements for future research, with a view to providing more accurate disease prediction and even personalised ‘silver bullets’ in terms of medication. Kivinen gives two examples: pre-screening of disease risk factors, and drug treatments.
Nature Medicine ran an article in spring 2020 based on data analyses made in the Finngen project. On the basis of genomic data, Professor Samuli Ripatti’s research team at FIMM was able to identify a Finnish population group that had a 60% probability of suffering from cardiovascular diseases or diabetes at some point in their lives. The research material consisted of 135,000 Finnish people providing a blood sample. Data obtained about individual risk factors was combined into a genome-wide risk score.
“Once a person’s genetic background is known, clinicians will be able to offer preventive measures and treatments to improve health as well as save time and money,” says Kivinen.
“In future, population-wide screening can be customised. For example, some women are now screened too early for breast cancer given their small inherited risk, while others are screened far too late. If we use genomic data to take account of inherited risk factors, women could be invited for their first mammography at the optimal time, thereby maximising the chances of discovering cancer at an early stage while minimising unnecessary radiation from repeated screening.”

People react to drugs differently: some do not get the necessary dose, while others suffer from side effects. The reason for varying responses may be our physical characteristics, other medication, or genome. Through the pilot phase of ‘HUS e-care for me’ project, we are optimising treatment for patients with leukaemia and other blood cancers. The project kicked off in summer 2019 and uses artificial intelligence to combine patient-specific biological data produced at FIMM with information about cancer type and status to identify suitable, personalised treatments and stop cancer progression.
“Sometimes a drug may not be effective, or have lost its effect. Cell cultures are made from leukaemia patients’ blood samples, in order to analyse which combination of drugs works best.”
Genome and transcriptome sequencing are performed on the same blood sample. A transcriptome provides information on whether the operation of genes has changed as a result of genetic mutations.
“If a drug no longer has the desired effect, we can find out what kinds of mutations have occurred. Are some genes working, or not, as a result of a mutation? And how do mutations affect metabolic pathways? Now we can see which drug is the best suited for different blood cancer patients directly from blood samples.”
Personalised drug treatments are possible if enough data is available on the patient, and it has been stored and pre-processed correctly. Alongside eleven other academic and commercial players, the Finnish ELIXIR Center CSC and the Barcelona Supercomputing Center (BSC) kicked off the European HPC Center of Excellence for Personalised Medicine (PerMedCoE) project in October 2020. The project is developing algorithms that can significantly reduce the computing time required for analysis. Analysis of genetic and protein data is becoming faster, facilitating and speeding up disease diagnosis and identification of the appropriate treatments. A genome analysis can take weeks or even months. Thanks to super computation and better software, diseases can be diagnosed in future within just hours or days.
Projects like this are important for research teams at FIMM who are routinely working with massive amounts of data.
“The amount of data is increasing at an accelerating pace owing to more powerful equipment and methods,” says Katja Kivinen. “At the moment, we are chronically in short supply of data storage space, and data pre-processing takes too long to enable us to clear up job backlogs and send analysed data to research teams. A secure environment for storing and processing data is vital when dealing with human data. Commercial cloud services offer a secure operating environment, but are too expensive for most researchers. What is more, some data requires a carefully tailored pre-processing and analysis environment and is poorly suited to the options available in cloud services.”
Data processing can be alleviated by dividing work between CSC and FIMM. In the pilot stage, genomic data is transferred from FIMM to CSC on a superfast and secure optic cable. Data pre-processing and quality assurance of analysis are fast, because the data is located at CSC. In future, CSC will distribute data back to research teams on a national basis.
“Previously, it took us 2–3 days per human genome to determine what kinds of changes had taken place. Thanks to this cooperation, the manufacturer of our sequencing equipment has given us access to an optimised computing server that can compress the processing of one genome into 20 minutes. This will help us deal with the backlog of genomic data processing at FIMM and free our bioinformatics specialist to perform other work – such as planning and facilitating various data integration procedures.”
CSC has developed a a new interface to its Allas -service for genomic data from FIMM, to be used by Finnish research teams. The research teams will receive a message once their genomic data is ready, and transfer it to their own project area in CSC’s ePouta environment. Following the pilot stage, the portal’s operating principle will be offered on a larger scale to all research teams in Finnish universities producing ‘omics’ data.
“The interface is vital for us, as data volumes are increasing and data security requirements are becoming tighter, and it is increasingly difficult for us to maintain a data storage and processing environment at FIMM. We must start moving more and more raw data and process data to CSC and make it accessible for research teams.”
Another important area of development, according to Kivinen, relates to data storage and related imaging services.
“Image processing is usually done on the server connected to the actual instrument, which contains the necessary processing software. Owing to slower transfer speeds, moving processing to a cloud service is not always a viable alternative, at least not in all parts of the country. So image processing may also occur on location in the future but, like genomic data, processed data should be shared through CSC .”
Ari Turunen
10.11.2020
Read article in PDF
Citation
Ari Turunen, Katja Kivinen, & Tommi Nyrönen. (2020). Bioinformatics to revolutionise healthcare: Efficient data processing speeds up diagnoses and enables personalised drug treatments. https://doi.org/10.5281/zenodo.8135131
More information:
FIMM
Cleaverhealth
https://www.cleverhealth.fi/fi/ecare-for-me
+1 million genomes’
https://ec.europa.eu/digital-single-market/en/european-1-million-genomes-initiative
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Turku University Hospital and Auria Biobank aim to have all tissue specimens in digital format. The samples would be scanned from glass slides, with diagnostics in pathology performed on computers. They will also develop artificial intelligence models, or classifiers, to identify e.g. cancer from digitalised samples.
Turku University Hospital alone takes 200,000 patient samples every year. Tissue samples are placed in formalin and cast into a paraffin block that can be sliced and subsequently examined under a microscope. The paraffin blocks are stored at the end of the process. Managing the samples is laborious and time-consuming. Systematic digitalisation of the samples makes the job easier.
“Since the samples are so numerous, metadata will enable us to quickly find the samples that we want,” says Antti Karlsson, data analyst at Auria Biobank.
You can, for example, search the database for all samples indicating breast cancer tumours. By using metadata, searches can be narrowed down to pinpoint, say, samples with a certain receptor status from 60-year-old breast cancer patients.
In the digital pathology project, samples on microscope slides are scanned. Then a pathologist can view the samples on a computer screen and describe and classify them. All this annotation data is relevant to teaching artificial intelligence to automatically detect abnormalities such as cancer cells in the samples. This would considerably speed up the work of pathologists. Auria Biobank has invested in data analytics, development of algorithms, and machine learning models.
Turku University Hospital has huge numbers of tissue specimens stored on microscope slides. The problem is that no metadata can be stored on the slides and transferred into databases automatically. The idea now is that pathologists add metadata to new samples by means of graphics software.
Karlsson says that the work is mechanical to begin with. Pathologists use graphics software to indicate the points where e.g. cancer is found on the scanned samples.
Description data is also required. This is where neural-network language models might come in. A pathologist would add information about the sample directly into a computer. This has been studied in cooperation with Filip Ginter’s research team at the Department of Future Technologies of the University of Turku. The research team has focused on how computer software can be used to analyse natural text and speech. From a large amount of unclassified text, the language model learns how a spoken language seems to work statistically. Auria Biobank and Turku University Hospital are interested in how medical statement texts could be formed into classified and structural information by means of language models.
“One application of digital pathology could be to mine various types of information from statements, such as which part of the sample contains interesting tissue, making sample selection for research purposes easier. We could also develop a model that automatically structures regular medical statement texts. Pathologists could speak in ‘prose’, which artificial intelligence would collect and compile into a structured table.”
According to Karlsson, such tables are already used relatively frequently, for example when pathologists have agreed on what aspects of each tumour should be reported.
“At the moment we are experimenting with these models, for example to detect and classify smoking data from among hundreds of thousands of statements, and to detect cancer metastases, symptoms related to hospital infections and various diagnoses.”
The challenge is that data comes in a variety of forms. For example, scanners by different equipment manufacturers produce different kinds of data that should be presented in a systematic way.

Metadata and digitalised sample material are used to develop artificial intelligence applications, for example, which are taught to automatically classify the locations with cancer cells in images. To teach the artificial intelligence system, we require some material classified by pathologists. According to Antti Karlsson, you do not in fact need very many images for the algorithm to start learning.
“A few dozen images is enough to get started. A single, whole slice image may yield a thousand small images that can be used to train the models.”
This means that up to 10,000 small images can be obtained from 20 patients.
“A large image cannot be used with an algorithm, because no computer has the kind of graphics processor memory required to deal with it.”
Karlsson stresses that artificial intelligence models which examine images are different to models examining texts.
“Clearly, they are all manifestations of artificial intelligence, and even of neural networks, but their structures and operating principles are quite different. Artificial intelligence is actually more like a collection of tools, each one of which is useful for a specific application.”

Auria Biobank’s Director Lila Kallio says that, in addition to genome data being used for research purposes, digital pathology making use of data analytics is a focus area at Auria.
“There is growing interest in how digitalised cancer tissue samples can be used to identify various issues. We are involved in studies where we try to use an algorithm to examine an image of a primary cancer tumour and predict how it will respond to treatment, or whether the primary cancer tumour will metastasise. There are indications that the algorithm may be able to predict something that is not otherwise visible from a histological image.”
According to Lila Kallio, Finland has been a pioneering country in data management and sharing. The Finnish Biobank Act has enabled research and the combination of data from various registers. It is critically important that clinical information can be connected to samples.
“Services for researchers have been provided on a one-stop basis. Biobanks take care of the permits, collect the samples and combine all clinical data related to the research. All this can then be combined with other data, such as genetic data.”
Researchers get all the samples through a biobank.
“Biobanks cooperate in Finland. Researchers can request samples from all Finnish biobanks with a single request made to the Finnish Biobank Cooperative.
According to Lila Kallio, the challenge now and in the future is data storage and management.
“Data is stored inside firewalls in hospital districts. If diagnostic samples will be digitalised on a larger scale within pathology, the storage capacity problem must be solved as well. In addition, the image sizes are so huge that they cannot be transferred on ordinary data networks.
The Finnish ELIXIR Center CSC plays an important role in terms of computing power, and safe storage and usage environments.
Ari Turunen
28.2.2020
Read article in PDF
Citation
Ari Turunen, Antti Karlsson, Lila Kallio, & Tommi Nyrönen. (2020). Tissue samples into digital images, interpreted by artificial intelligence. https://doi.org/10.5281/zenodo.8134949
More information:
Auria Biobank
https://www.auria.fi/biopankki/en/
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Digitalisation is revolutionising pathology. Scanners can be used to convert microscope samples into digital format. The scanner captures the sample one view at a time and a computer combines them into a virtual microscopy image.
At Turku University Hospital, pathology samples are digitalised and examined on computer screens. This enables various measurements and AI applications. AI applications developed in cooperation with the Auria Biobank reduce routine work by pathologists and expedite sample analysis.
Markku Kallajoki, Managing Director of Pathology at Turku University Hospital, has done basic cancer-related research based on the study of cell models and cell cultures. He has been a specialist in pathology and Professor of Cellular and Molecular Pathology. One of Kallajoki’s special interests is prostate cancer.
Pathologists view a tissue sample under a microscope to evaluate the aggressiveness of prostate cancer. A malignant tumour is given a so-called Gleason score on a scale of 6 to 10. The higher the score, the more aggressive the cancer. A Gleason score of 7 is considered the threshold between good and bad prognoses. High Gleason scores (8–10) refer to an aggressive tumour and low scores (less than 7 points) to non-aggressive cancer.
“The higher the score, the more aggressive the cancer. Artificial intelligence can identify cancerous areas in tissue samples before examination by a pathologist. It may also suggest a Gleason score. This enables pathologists to focus on providing a second opinion on the sample areas identified by AI. In any case, artificial intelligence facilitates and speeds up the work of pathologists,” says Kallajoki.

Researchers from the University of Tampere and the Karolinska Institute in Stockholm have developed an AI-based method of microscopic diagnosis and classification of prostate cancer. 6,600 prostate biopsies were used as a material for teaching artificial intelligence to distinguish between benign and malignant biopsies. A model was created from the samples capable of identifying and quantifying cancer in tissue samples, and classifying its malignancy.
Studies suggest that 15% of a pathologist’s working time is spent on non-diagnostic work. It takes time to find, process, receive and acknowledge samples and referrals. In addition, analysis of samples often requires consultations with other pathologists. Digitalisation reduces the time needed for such consultations, by enabling pathologists to send images online, rather than microscope slides. They can discuss online images on computers in different hospitals, for example.
“Digital pathology eases our work and enhances its quality. It speeds up work and saves money,” says Kallajoki.
Digitalisation alone allows a pathologist to analyse around 15% more samples than now. When AI is added, this work could become up to 30% faster.
Prostate cancer is the most common malignant cancer in men. It occurs when prostate cells become malignant. Based on biopsies taken from the prostate gland, a pathologist can estimate the cancer’s malignancy based on differentiation of the tumour. The more poorly differentiated the tumour is, the more aggressive the cancer will be.
“A microscope sample is taken if the preliminary clinical examination and findings, laboratory examinations and radiological imaging point to cancer,” states Kallajoki.
“Cancer is not cancer until it is confirmed by a pathologist from a cell or tissue sample. If prostate cancer is suspected, a needle is used to take a sample from the prostate gland via the rectum. In most cases, a total of six biopsy samples, one to two centimetres in length and around a millimetre thick, are taken from both sides of the prostate gland. These cylindrical fragments of tissue are sent to a pathology laboratory for the preparation of histological samples.”
The need for treatment is assessed based on these histological (microscopic anatomy) samples. The samples are fixed in formalin, increasing the tissue’s mechanical strength and preserving it against the destructive effect of the cells’ enzymes. Infiltrated with paraffin, the samples are then embedded in paraffin blocks, from which thin slices of three to four micrometres are cut. These are stained with histological dyes and placed between two glass plates. The samples can now be viewed under a microscope and, if necessary, scanned and digitalised.
High-resolution, digitalised tissue samples reveal the same details as when viewed under a microscope. A digitalised image allows the measurement and automatic calculation of different cell types. In addition, it is easy to return to the samples, as the images can easily be retrieved from an archive for review, for example at meetings where patients’ treatment is being discussed.
Pathologists also learn much from other data. Kallajoki explains that biobanks such as Auria are important. Data is now available from multiple sources, which facilitates the practical work of pathologists. Medical records are a source of patient data, describing the examinations performed and laboratory test results. Imaging data from radiology is also available.
Kallajoki believes new treatments will be developed through the use of data and new methods.
“We are living in exceptional times: cancer treatments are under intense development and new targeted treatments based on molecular changes are forthcoming.”
“Digital images are huge. The image size is 2–3 gigabytes. An enormous amount of data is created when 12 images are taken of one patient in a single examination. Around 200,000 sample slides are made each year at Turku University Hospital. Because this is medical information, two or three backups are required. Multiply the saving of 200,000 microscopic samples by three, and you get a huge data storage requirement.”
Markku Kallajoki explains that the key challenge lies in the fact that the purchase of digital pathology hardware, software and storage systems is planned in different locations, but such systems need to be compatible with each other.
“The optimal system would be a compatible, Finnish-wide one. In digital pathology, the largest single cost item is storage capacity.”
Ari Turunen
9.6.2020
Read article in PDF
Citation
Ari Turunen, Markus Kallajoki, & Tommi Nyrönen. (2020). Digital pathology speeds up diagnosis. https://doi.org/10.5281/zenodo.8131372
More information:
Turku University Hospital
Auria Bio Bank
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

In addition to gene variants there are also genomic variants in the locations of the single base pairs in the DNA stretch. The variations cause differences between individuals, but they can also help localise the disease-causing genes. These single nucleotide polymorphisms (SNP’s) can act as markers indicating the disease. The artificial intelligence model developed at the University of Eastern Finland searches breast cancer interacting SNP’s.
The huge amount of genomic data has made possible that researchers can now calculate what kind of gene variants are among the groups who have cancers. Hundreds or thousands of gene variants can have an impact to a single disease.
With statistical methods researchers can estimate how the gene variants of a single person can increase the disease risk. However, variations are also at the single base pair e.g. nucleotides in DNA, known as genetic variants or SNPs. DNA sequence variations occur when a single nucleotide (adenine, thymine, cytosine, or guanine) in the genome sequence is altered. Each SNP represents a difference in a single nucleotide. For example the nucleotide cytosine (C) can be replaced with the nucleotide thymine (T) in a certain stretch of DNA. It means that the base-pair cytosine-adenine can alter for thymine-adenine. Unlike gene mutations, SNP’s are not necessarily located within genes. They can be also in the non-coding regions of the genes or regions between the genes. There are lots of SNP’s in human genome. They occur almost once in every 1,000 nucleotides on average, which means there are approximately 4 to 5 million SNPs in a person’s genome.
SNP’s can be beneficial when searching the genetic risk factors for cancer. In biomedical research, SNP’s are used for comparing regions of the genome between cohorts with and without a disease.
”When SNP’s occur within a gene or in a regulatory region near a gene, they may play a direct role in disease by affecting the gene’s function. We have a novel machine learning approach to identify group of interacting SNPs, which contribute most to the breast cancer risk,” says researcher Hamid Behravan from University of Eastern Finland. He works in Kuopio at the Institute of Clinical Medicine.
”We have published several findings about identifying the genetic component of the breast cancer risk that would reliably distinguish disease cases from healthy controls. Identifying the breast cancer-associated SNPs that reliably distinguish disease cases from healthy controls may be particularly useful in improving breast cancer risk prediction and developing individual treatment strategies”, says Behravan.
The standard hypothesis testing methods have measured only the association between a single SNP with a disease. However, the studies by University of Eastern Finland have demonstrated that risk factors for breast cancer can be predicted better when SNPs are examined as groups that actually interact with each other.
The idea of genome-wide association studies (GWAS) is to identify SNPs on the DNA, which explains the genetic component of the observed phenotype in genotyped people.
”Genome-wide association studies measure the association between an individual SNP’s with a disease, but ignore the possible correlation among SNPs”, says Behravan.
”To date, population based genome wide association studies often use polygenic risk scoring (PRS), which aggregates the effects of risk alleles with the disease. However, PRS assumes that the disease-associated SNPs are independent of each other and the risk effects are linear and additive. We have shown that instead of evaluating the effect of single components (SNPs) one at a time, it would be particularly useful to improve breast cancer risk prediction by studying groups of interacting SNPs using an machine learning.”

The machine learning method developed in Eastern University of Finland has proven to be efficient.
”We found group of interacting SNPs that have true biological meaning. A biological analysis of the identified SNPs reveals genes related to important breast cancer-related mechanisms, such as Estrogen metabolism and apoptosis.”
Elevated endogenous estrogen levels are associated with increased postmenopausal breast cancer risk. There is also strong evidence that tumour growth is not just a result of uncontrolled proliferation but also of reduced apoptosis.
”So, we found genes behind those identified SNPs by our approach, and built gene interaction maps from those genes, and then we observed several separate networks related to breast cancer, such as Estrogen metabolism and apoptosis network. So not only our system found group of interacting SNPs with highest breast cancer risk predictive potential, but also those identified SNPs were behind a number of important biological entities in breast cancer. Therefore, interacting SNPs indicates both SNPs selected together, and SNPs involve in cancer related biological networks.”

The machine learning approach developed in Kuopio is based on a gradient tree boosting method followed by an adaptive iterative search algorithm. Boosting is the first module and searching the second module.
Boosting is an algorithm and method of converting weak learners into strong learners. Algorithm begins by training a decision tree. Weak classifiers are added sequentially to correct the errors made by existing classifiers towards building a strong classifier.
”The first module evaluates the accuracy of features, in this case the SNPs, on the breast cancer risk prediction. The first module provides an initial list of candidate SNPs with breast cancer-risk predictive features. ”
”The second module then uses the candidate SNPs in an adaptive iterative search to capture the interacting features. The best identified interacting SNPs are then used to predict the breast cancer risk for an unknown individual at the testing phase using a machine classifier. Classifier was trained to distinguish the breast cancer cases (positive samples) and healthy controls (negative samples).”
Since cancer is a multi-factorial disease caused by lifestyle, genetic, and environmental factors, individual analysis of the sources of genetic variants may not be enough to create a comprehensive view of the disease risk. According to Behravan other sources of data is needed.
“We are developing integrative machine learning approaches to combine different sources of data, such as demographic data.”
Ari Turunen
18.5.2020
Read article in PDF
Citation
Ari Turunen, Hamid Behravan, & Tommi Nyrönen. (2020). Searching markers for breast cancer by machine learning. https://doi.org/10.5281/zenodo.8131311
More information:
School of Medicine, University of Eastern Finland
https://www.uef.fi/en/web/laake
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org
Finland’s ELIXIR node is CSC –Scientific Computing Ltd. The Finnish life science research projects have been compiled to one publication. In the pubication different studies, which utilise new methods of bioinformatics, are presented. These are among other drug design, the right doses of the medicine, plant breeding and the Finnish rare diseases.
CSC is participating in building researchers a versatile computing platform (ELIXIR Compute Platform) that includes a number of important services. Once authorised to use the platform, a researcher can make use not only of the computing features, but also various data sources, in addition to storing, transferring and analysing data. In 2019, the ECP had 3,100 users. Read the report.

During metabolism, molecules are created and broken up, and some of these have an effect on health. Their concentrations are measured from blood, urine and tissue samples. Metabolomics enables the detection of biomarkers that can give an indication of a person’s lifestyle, diet, illnesses and the effects of medication and other xenobiotics.
A single measurement yields information about hundreds, possibly thousands of metabolic products (metabolites). The same measurement also reveals external compounds, such as medication, environmental toxins and stimulants.
“Metabolomics enables the comprehensive observation of metabolic phenomena. This gives us an extremely good idea of the body’s biochemical state,” says Professor Seppo Auriola, of the School of Pharmacy of the University of Eastern Finland. Auriola is also the head of the LC-MS Metabolomics Center in Kuopio, which is part of Biocenter Finland’s infrastructure network.
One analytical tool used in metabolomics consists of a combination of liquid chromatography and high-resolution mass spectrometry. Liquid chromatography-mass spectrometry (LC-MS) is used to screen and identify compounds in samples. Liquid chromatography separates compounds on the basis of their fat solubility, while a mass spectrometer is used to measure exact molecular weights. The term ‘molecular feature’ — meaning the signal generated by a compound during ionisation and measurement — is used in metabolomics.
“In metabolomics, we attempt to find the statistically different molecular features be- tween the different groups being studied. These could be ‘ill versus healthy’, for example. Metabolomics also involves trying to identi- fy such molecular features as molecules, by means of various spectroscopic techniques. Our lab uses mass spectrometry for this,” says Laboratory Manager Marko Lehtonen.
Metabolomic measurements can be divided into untargeted and targeted methods. The starting point with untargeted analysis is trying to find as many metabolites as possible from a sample. A targeted analysis, on the other hand, focuses on a limited group of known metabolites.
Untargeted measurements can provide a good basis for creating a hypothesis.
“The first screening reveals metabolic products that have changed, for example after the first exposure. Then we start thinking about the theory and try to understand why this occurred,” says Auriola, who focuses on analytical chemistry and measurement techniques used on samples.

As metabolomics measurement methods become increasingly efficient, more accurate measurement data will be obtained on the effects of people’s lifestyles and environment on their health. Diet is a key external factor affecting a person’s metabolism.
“Metabolomics is ideally suited for dietary studies. Analyses provide clear markers on what a person has been eating and how this affects their endogenous compounds,” says Auriola.
Endogenous substances comprise all compounds produced by the body, such as hormones and transmitters. These include endocannabinoids, steroids and endorphins.
“We can examine whether a positive lifestyle change also affects metabolite levels. This would be an indication that the body is doing better. Metabolomics can also be used to detect disease biomarkers at an early stage, before diseases actually occur.”

Another important area suitable for metabolomics analysis is exogenous compounds – that is, compounds from outside the body – such as medication and environmental toxins. This involves looking for biomarkers to show how a medication is affecting the body.
Auriola thinks it is also important to ask why a certain substance affects us negatively. We can also look for such biomarkers in metabolic products that indicate human susceptibility to a xenobiotic, or the effect of a xenobiotic on humans. These include the effect of pesticides on human health.
“We do not understand the mechanisms of all pesticides. As we develop more advanced methods, we will obtain a clearer picture of how humans are affected by exposure to certain substances. We can measure the level of environmental toxins and the corresponding level of endogenous metabolites in human populations.”
Studies by the University of Eastern Finland and Karolinska Institute examined the effect of polychlorinated biphenyls (PCBs) on mouse offspring. It has long been known that these substances have most effect in the early stages of development. Animal tests have revealed developmental disturbances in various organs. When the metabolomics profiles of offspring were studied, certain changes were found in males. However, such changes were absent from females. The metabolite changes caused by PCB compounds in males affected the liver and nervous system.
“We will be able to monitor changes in the following generation without knowing in advance what we should be looking for,” says Auriola.
“By means of LC-MS equipment and the untargeted metabolomics method, we can find changed molecules among the thousands of molecules we are measuring.”
Molecular characteristics are identified by means of algorithms. The study by the University of Helsinki and University of Eastern Finland involved the analysis of compounds sampled from neonatal umbilical cords. Pre-eclampsia (a type of pregnancy disorder) is one of the commonest causes of premature birth and maternal deaths during childbirth. The precise causes of the condition are unknown. It is known to increase the subsequent risk of cardiovascular disease in both mother and child. However, we do not know how the changed metabolism of mothers with pre-eclampsia affects the metabolism of newborns. Metabolites in the umbilical cord tissue of newborns were analysed with the LC-MS equipment in Kuopio, comparing the results between those who had pre-eclampsia, and healthy controls. The study also made use of material by the Finnish Genetics of Pre-eclampsia Consortium. All Finnish university hospitals contributed to the assembly of the FINNPEC cohort.
“Many different research projects use the services of our laboratory,” says Marko Lehtonen. For example, research samples related to diabetes and Alzheimer’s have been studied in the laboratory. According to Lehtonen, metabolomics will provide more information that can be used to study rare and hereditary diseases.
“Newborns are screened with targeted measurements. This is also an excellent example of an area where metabolomics can be very significant. It will save society money. Based on certain biomarkers found in the body, hereditary diseases among newborns can be identified,” says Lehtonen.
Not all metabolites can be measured using the current equipment.
“Compounds are present in a sample in such small concentrations that we also need targeted methods. As equipment technologies develop, untargeted methods may become efficient enough to reveal compounds that could not be detected earlier. This will ensure that we do not lose other information from a sample. Targeted methods only track specific compounds and are blind to all other data,” says Lehtonen, stressing that the untargeted method provides plenty of data which can be used to investigate new issues.
As equipment becomes more accurate and sensitive, we will be able to observe really small concentrations. We’re talking about picograms and nanograms per litre. One picogram is one trillionth of a gram, and one nanogram is one billionth of a gram.
“We can currently see thousands of compounds, but many important molecules remain below our observation horizon,” says Seppo Auriola.
“For example, more and more steroids will be identifiable in samples as measurement technology improves. This will enable us to study endogenous steroids and their metabolites.”
These include sex hormones, such as testosterone and progesterone, and corticosteroids (e.g. cortisone and cortisol).
“We are involved in a project studying the effect on steroids, and other metabolic characteristics, of exercise and lifestyle choices among children and young people. Other studies involve trying to find compounds that affect steroid metabolism selectively, and may therefore be used as medication.”
Metabolic products studied by means of mass spectrometry are first ionised. These ions are separated from each other on the basis of their mass-to-charge ratio.
According to Lehtonen, the identification of molecular characteristics is the last stage in metabolomics, based on the attempt to clearly identify a statistically different metabolite between two or more groups being studied.
Lehtonen would prefer a model in which laboratory and research data were used as a basis for machine learning.
“Although these spectra can be compared to fragmentation spectra found in mass libraries, the problem is that identification still involves high amounts of manual work. It would be ideal to have a learning algorithm that automatically sought fragmentation spectra and compared them to what was in the library. Such a model could accurately define compounds identified previously in a laboratory. This would be of considerable help to research,” says Marko Lehtonen.

According to Seppo Auriola, we should make more use of measurement data. The problem lies in the availability and uniformity of data.
“ELIXIR has several processes underway to unify the use of various tools in metabolomics, in order to render them compatible. Measurement data should also be archived.”
According to Auriola, in addition to being used for scientific publications, most original measurement data should be made available to other researchers for further analysis.
“The second phase involves adding metadata, determining what kind of data should be available on the samples, how they have been measured and verified, and what kinds of groups have been studied. How will this data be conveyed along with the measurement data? The crucial issue is that data that took a lot of work to obtain could be used for later analyses and comparisons.”
Another challenge involves the available tools: how to pick and identify compounds, and what software is required to calculate the results, to identify molecules and compare their numbers in various samples. How are facts presented? How are changes in metabolite levels obtained, how are they found on the metabolite map, where are the compounds located on metabolic routes, and how are their concentrations changed? How can this be described clearly and how should the result be presented? A fair amount of work is required to unify all this. All the related data and tools are currently fragmented between various people’s software,” says Auriola.
Ari Turunen
8.4.2020
Read article in PDF
Citation
Ari Turunen, Seppo Auriola, Marko Lehtonen, & Tommi Nyrönen. (2020). Metabolomics measures and analyses metabolic changes caused by illness, diet or medication. https://doi.org/10.5281/zenodo.8131264
More information:
LC-MS Metabolomics Center
University of Eastern Finland
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Researcher Raju Gudhe has studied computer science with a focus on intelligent systems. He is now developing deep learning algorithms for breast cancer risk analysis using radiology and clinical data. These algorithms have been trained using massive data sets from Kuopio University Hospital to predict the density of breasts on mammograms
“We try to localise regions of interest on a mammogram and classify the tumour type based on features extracted using deep learning algorithms,” says Gudhe, who works as a data analyst at the Institute of Clinical Medicine of the University of Eastern Finland, in Kuopio.
Mammography, a low dose x-ray imaging technique, is one of the most widely used methods of detecting early-stage breast cancer. The early detection of breast cancer significantly reduces mortality rates. In 1987, Finland was the first country in the world to begin a country-wide cancer screening programme. However, mammography is not perfect. Mammograms are not particularly sensitive and can miss cancer cases, or can appear normal even when cancer is present.
Breasts consist of variable portions of adipose (fat) and fibroglandular (dense) tissues. Being denser, fibroglandular tissues appear white (bright) on mammograms due to the attenuation of x-rays. Most cancers occur in the fibroglandular tissues, the brightness of which hide around 25% of cancers in mammograms.
“This brightness can mask the presence of cancers; it is like finding a snowman in a dense snow cloud”, says Gudhe.
Based on fibroglandular tissue patterns and their distribution, radiologists categorise breasts as either ‘dense’ or ‘fatty’. Women with extremely dense breast tissue have a higher risk of developing breast cancer.

“We researchers at the University of Eastern Finland and Kuopio University Hospital are interested in developing a fully automatic model for estimating breast density. Breast density, one of the strongest risk factors in breast cancer, is a measure of the relative amount of fibroglandular tissues. Accurate segmentation of fibroglandular tissues on a mammogram can reduce the likelihood of false diagnosis.
Algorithms developed by University of Eastern Finland can assist radiologists in the accurate estimation of breast density. The major challenge of using deep learning models is the massive amount of training data they require. In addition, in medicine the acquisition of images with accurate annotations simply adds to the complexity of data.
“We use thousands of mammogram images, which are manually annotated by an expert radiologist to generate accurate training set’s classification (e.g. ground truth labels) for our deep-learning models. We have developed a novel architecture based on the U-Net model, a state-of-the-art solution for the medical image segmentation of fibroglandular tissues”, says Gudhe

Because mammogram images are high-resolution, high computation power is required to train the related deep learning models. In this regard, the services provided by the Finnish ELIXIR centre CSC are used to handle sensitive data efficiently, training the models on graphic processing units provided by CSC.
Raju Gudhe emphasises that, in order to make a robust model for clinical implementations, researchers must integrate different imaging modalities and other clinical features with their algorithms. The next step is to integrate imaging and genomic data in order to analyse the cancer risk.
“Using mammograms, we can identify the density of a breast and, based on the density value, can generate the next image modality. We cannot rely on a single modality for the images, which is why the information cannot be used directly in clinical practice. To obtain an end-to-end model with good classification and diagnosis, we also need genomic data”.
Ari Turunen
1.3.2020
Read article in PDF
Citation
Ari Turunen, Tommi Nyrönen, & Raju Gudhe. (2020). Deep learning algorithms help in breast cancer screening. https://doi.org/10.5281/zenodo.8131233
More information:
School of Medicine, University of Eastern Finland
https://www.uef.fi/en/web/laake
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Breast cancer is the most common type of cancer in women. One quarter of all cancers in women are breast cancers. Until now, genetic risk factors for breast cancer have usually been studied as single factors. Professor Arto Mannermaa intends to explore the big picture and look for more factors which significantly increase the risk of illness when they interact. This is where artificial intelligence comes in.
Mannermaa’s team is developing algorithms that can learn on the basis of genomic and clinical data, and identify and predict risk factors. Learning algorithms are also used in the interpretation of mammography images. Genomic and clinical data are integrated to an AI model that not only helps to determine the risk of illness, but also in drawing up individual treatment plans.
“I’m a biologist and geneticist by training. I engaged in the close study of human genetics and specialised as a hospital geneticist. I have been working on clinical studies in genetics laboratories, but have been interested in cancer throughout my researcher career,” says Arto Mannermaa.
Mannermaa is a Professor of personalised medicine and biobanking at the University of Eastern Finland. In his research, he has focused on the genetics of breast and ovarian cancer. Since its inception, Mannermaa’s team has been involved in the work of the world’s largest genetic epidemiology consortium, the Breast Cancer Association Consortium (BCAC). The consortium has the world’s largest centralised collection of breast cancer tissue samples, collected from over 200,000 patients and controls. The collection includes well-annotated data on factors and clinical results related to breast cancer.
“My research team has been engaged in long-term work to determine the genetic risk variants of breast cancer. We can now identify some 200 variants within normal genomic variation which increase the risk of getting breast cancer. Together with the BCAC, we have also learned about the genetic mutations that are major contributing factors to breast cancer. These include among others mutations of the BRCA and PALB-2 gene, which we were involved in finding to be a contributing factor in breast cancer.”
If a woman has a BRCA1 or BRCA2 gene mutation, she has a 60–80% risk of developing breast cancer. The risk impact of PALB-2 is almost the same.
It is estimated that genome accounts for about 30 per cent of susceptibility to breast cancer, while 70 per cent is determined by environmental factors. Risk factors for breast cancer include the total amount of oestrogen during a lifetime, which depends on the number of pregnancies and children, and weight. Other factors include smoking, alcohol consumption and minor exercise. According to Mannermaa, risk factors tend to have been studied one at a time. Now the BCAC has been able to study the prevalence of breast cancer in the close relatives of patients. This includes plenty of international material that can be used for making comparisons.
“For example, studies have been done on whether there are more common factors in Finnish cancer data than in international data.”
Research material has been obtained from the Biobank of Eastern Finland and Kuopio University Hospital.
“The research team is grateful to all volunteers who participated in the study. Without their consent, this kind of work would not be possible. ”
Finnish data has been compared with data obtained through BCAC, collected from more than 100 research teams around the world. Although data has been obtained from around the world, for Mannermaa the challenge lies in the fact that the material was collected for different purposes, and is not always in the same format.
“In order to use the material, the data collected from different sources must be unified, which often takes up a large amount of the total time spent on research.”

The question Mannermaa wants to answer is which factors have contributed to the onset of breast cancer in a patient. Mannermaa’s team has created an AI model for breast cancer risk factors that is being tested with Finnish and international material.
“We also have material obtained from the Biobank. We are comparing the data of breast cancer patients and healthy individuals and trying to find the interactive combination of all variables that has the greatest influence on the onset of breast cancer.”
One of the study’s targets concerns normal genomic variation, or SNPs. The rapid development of DNA sequencing techniques has made it possible to determine single nucleotide polymorphisms (SNPs), providing a very accurate estimate of the differences between individuals. SNP is the difference in the DNA chain caused by a mutation within a population. According to some estimates, the human genome has 4–5 million SNPs, located in the DNA chain, either in the inter-gene or gene region. They can act as biomarkers, helping researchers to locate genes related to diseases. Certain SNPs can affect the operation of the gene and thus directly affect the onset of the disease.
“In practice, we focus on the differences between cancer patients and control groups. We want to learn how many SNPs are in common with these groups, and what the common SNP network is like among cancer patients compared to healthy individuals.”
Mannermaa’s team is working to identify SNPs related to breast cancer, by means of AI and learning algorithms.
“We teach our algorithm to detect SNP networks. With the help of artificial intelligence, we can identify the interactive group of SNPs with the greatest impact on disease risk.”
The results have been promising. The algorithm helped to identify genes close to SNPs, and these SNPs are probably affecting the operation of the genes. We found a gene network related to oestrogen metabolism.
“Oestrogen metabolism is a key component in the development of breast cancer, while another group that we found was related to apoptosis, or programmed cell death. Apoptosis is crucial in cancer development, because cancer cells must be able to prevent programmed cell death. That’s why we believe that the AI models helped us find the correct breast cancer factors. ”
The amount of data in Mannermaa’s team’s study is so huge that CSC’s (ELIXIR Finnish centre) supercomputing capacity is required.
“About 200,000 SNPs can be identified from one laboratory sample. Each SNP is compared with all the others. In addition, we simulate genetic variation, in other words what SNPs they have in common but remain unidentified. This means that up to another 10 million SNPs can be added to the equation. Add to this variables from imaging and the biobank, and computing capacity is definitely called for.”
The basic model of the Mannermaa team’s AI is based on genetic data. Clinical variables, i.e. breast cancer risk factors, have now been added to this model. Mannermaa believes that the models will significantly improve diagnostics.
“Artificial intelligence enhances screening and diagnostics. In the future, we can avoid overdiagnosis and use the data to differentiate those who need more accurate screening from those who don’t. This means that certain women do not need frequent mammography, due to their low risk of developing breast cancer.
Once genetic data is combined with not only known risk factors but also breast cancer diagnosis and treatment, the predictability of the disease will improve and personalised treatment plans can be drawn up.
Biobanks play a crucial role in research of this type. It is essential that all data is available.
“If the person giving a sample has consented to their data being used for biobank purposes, this data is combined with other data. The Biobank Act is the basis for secure data storage, and enables people who have given their consent to cancel it if they wish. Biobank consent is general consent based on the law. Through biobanks, everyone has the opportunity to participate in research aimed at developing health care.”
Mannermaa leads the SOTE AI Hub project, funded by the Regional Council of Pohjois-Savo. The project is seeking to improve the use of various data sources and AI in aid of decision-making. The project involves utilising and developing health data in a data lake. The Pohjois-Savo data lake consists of social and health data from the Biobank of Eastern Finland, Kuopio University Hospital and the City of Kuopio.
According to Mannermaa, the health data can be used to evaluate the impact of the research results. In addition to receiving plenty of data on the actual patient, we can see the impact of cancer patients’ treatment alternatives and solutions based on new research.
“The model and its prediction can help determine how it affects a patient’s life, and how resources should be allocated. This can make treatment more effective in the future. Patient-specific profiling and individualised treatments help to provide the right treatments for the right patients, and thereby make health care more efficient. This requires a multidisciplinary network.”
Ari Turunen
14.2.2020
Read article in PDF
Citation
Ari Turunen, Arto Mannermaa, & Tommi Nyrönen. (2020). All breast cancer risk factors evaluated with AI. https://doi.org/10.5281/zenodo.8131216
More information:
School of Medicine, University of Eastern Finland
https://www.uef.fi/en/web/laake
Institute of Clinical Medicine, University of Eastern Finland
https://www.uef.fi/en/web/kliinisenlaaketieteenyksikko
Cancer Center of Eastern Finland, CCEF
The Breast Cancer Association Consortium
http://bcac.ccge.medschl.cam.ac.uk
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Metabolomics involves the study of the body’s metabolic products, also known as metabolites, and their structure and operation in cells, the blood and secretions. The key issue is understanding the significance of metabolites and their effect on human wellbeing and health. Soile Rummukainen is using metabolomics to study canine and human cancers. Her goal is to identify olfactory molecules related to cancer.
Susanna Paavilainen, the Managing Director of the Wise Nose association, which specialises in training dogs to distinguish between various smells, found her dog Kössi sniffing a specific area of another dog’s skin. She realised that something was wrong. Eventually, it was discovered that the other dog had gum cancer. She figured out that, thanks to its acute sense of smell, a trained dog could detect cancer in other dogs.
A multidisciplinary research project was started between the University of Helsinki’s Faculty of Veterinary Medicine, Wise Nose, Aqsens Health Ltd. and the University of Eastern Finland. First, the dogs are trained to identify signs of canine mammary tumours from urine samples. According to tests, sniffer dogs’ results were good, with cancer detection of almost 100 per cent. This method will now be extended to detecting prostate and breast cancer.
Dogs have an extremely acute sense of smell. An average-size dog has up to 220 million olfactory receptors, compared to just 5 million in humans. This means that dogs’ sense of smell is thousands of times better than that of humans. A mass spectrometer used for the detection of organic matter generally needs some ten billion molecules before anything shows in the reading. A dog can smell out a disease from a much smaller number. In a test conducted at the University of Eastern Finland, Kössi only needed a sample with ten molecules.
Metabolites are compounds of low molecular weight that are involved in various cell metabolism functions. These small molecules cannot be seen or detected directly. Instead, you need measuring devices, such as mass spectrometers, which create signals for subsequent analysis.
Soile Rummukainen, an early stage researcher at the School of Pharmacy at the University of Eastern Finland in Kuopio, uses a mass spectrometer to study cancer samples sniffed out by dogs.
“We study these cancer samples and control group samples by using the non-targeted metabolomics method. The mass spectrometer will reveal tens of thousands of molecular features of metabolic products in urine samples. We use statistical methods to compare differences between groups, trying to identify the most interesting metabolites, that is, those that are different between groups.”
With a mass spectrometer and liquid chromatography, it is possible to separate the compounds from the sample and create a mass spectrum for each of them. The x-axis indicates the ion mass formed by the molecules, and the peak height (y-axis) their relative abundance. On the other hand, the molecular fragmentation product is used to determine the molecular structure. Liquid chromatography (LC) combined with mass spectrometry (MS) is an efficient analysis technique for the definition of metabolites. LC-MS methods are used extensively in pharmaceutical research and clinical diagnostics.
According to Rummukainen, the difficult part of metabolomics is the identification of molecules. You have to be able to identify the molecular structure that corresponds to the fragmentation spectrum. Fragmentation ions are compared to global databases, their fragmentation spectrum library and our own standards.
“Our own standard library consists of standards we have analysed at the university. These will enable us to identify metabolites with maximum accuracy, since they have been analysed using the same methods and analytical devices and include retention time data, which is a key identification component. However, our library is limited, so we must also use other databases.”
The retention time refers to the time it takes for the compound to travel through the chromatography equipment to the detector.
“A biological sample may contain thousands of metabolic products. When we analyse a sample with a mass spectrometer, we get data that results in tens of thousands of molecular features. These features must then be combined into molecules. By means of the exact mass, fragmentation spectra and retention time, we can perhaps identify 100 or 200 metabolites from the sample, which is quite a small number.”

That is where the dogs come in again. We move on to fractioning, that is, taking partial samples from samples. Then we use the dogs to test whether the smell is still present in the fractions. Rummukainen points out that most work is involved in making and analysing the fractions.
“In the future, we will study these partial samples and analyse the compounds they contain using mass spectrometer methods and nuclear magnetic resonance (NMR) spectroscopy. With the aid of sniffer dogs and mammary tumour samples, our goal is to create a method that can also be applied to determine metabolites related to human cancers.”
Dogs are currently trained to sniff out prostate and breast cancer.
Data processing is also important. Processing raw data obtained from a mass spectrometer requires plenty of computing power and disk space.
“A single metabolite may be linked to a dozen of intracellular signal routes. In this respect, we would benefit from computer simulation to improve our understanding of the biological significance of any changes identified. It would also be interesting to combine data obtained through genomics and proteomics with metabolomics, once the necessary software and tools become more advanced.”
Ari Turunen
6.2.2020
Read article in PDF
Citation
Ari Turunen, Soile Rummukainen, & Tommi Nyrönen. (2020). A dog can smell diseases. https://doi.org/10.5281/zenodo.8131208
More information:
LC-MS Metabolomics Center, University of Eastern Finland
http://www.uef.fi/en/web/metabolomics-center
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

ELIXIR has built researchers a versatile computing platform that includes a number of important services. Once authorised to use the platform, a researcher can make use not only of the computing features, but also various data sources, in addition to storing, transferring and analysing data. All services are combined into a seamless workflow.
The ELIXIR Compute Platform (ECP) was built for biomedical needs between 2015 and 2019. ECP is a geographically distributed platform in which ELIXIR centres act in concert to provide services for the management of biological data. The centres operate independently, but are brought together by the Authentication and Authorisation Infrastructure (AAI) with which cloud services, and computing, storage and data transfer services can be coordinated. Researchers log into the system, which checks their electronic identity and allows the appropriate level of access to biomedical data. Researchers can then create a secure analysis environment for their software. Data is stored on the European compute cloud platform. The operating environment also helps groups of researchers to create scalable services.
Thousands of research laboratories create massive amounts of data. This data is also becoming more complex, which poses a major challenge. Data must be managed so that all users understand and handle it in the same way. Effective data management requires a federation that manages the infrastructure within which the user can transfer, exchange, process and analyse data. This is why the ECP was developed in cooperation with ELIXIR centres and European research infrastructures. Services designed for ECP researchers were jointly built by four scientific user communities studying marine microbes; cultivated and forest plants; human genes; and rare diseases.
Services within the ECP are offered by various ELIXIR centres. ELIXIR’s AAI service (Authentication and Authorisation Infrastructure) enables electronic user identification and the granting of access rights. Access to data is always decided by the owner of the data or computing service, but AAI will make access to data faster, and the data use policy and analysis are clear and straightforward.
A high-capacity network is used for data transfer, and software is used to build interfaces on top of it, like data pipelines. These handle data transfer, processing and analysis. The data flows are divided into smaller parts and are processed in parallel to increase computing power, enabling the transfer to occur without bottlenecks and delays. Analyses can be performed in a distributed manner. If the data is sensitive, data security federation is required.

In 2019, the ECP had a storage capacity of 50,000 terabytes. It provided 80,000 separate computing cores, that is, processing units. Between 2017 and 2019, its storage capacity doubled and the distributed computing resources increased by 33%. In 2019, the ECP had 3,100 users.
Microbe communities affect the lives of humans and animals and play an important role in various ecosystems. However, only a small proportion of microbes have been categorised and analysed. Study of the genetics of microbe communities has created a new field in biosciences, metagenomics. A group of genes collected from the environment and then sequenced can be analysed in the same way as the genome of an individual species.
The oceans are the world’s largest single ecosystem. Plankton is at least as important to the world’s climate as the rainforests are. However, only a small number of the organisms that create this ecosystem have been categorised and analysed. Ecosystems formed by plankton contain a huge amount of life: there are more than 10 billion organisms in each litre of ocean water, consisting of viruses, procaryotes, single-cell eucaryotes and cnidarians. These unique organisms contain bioactive compounds that are useful in the pharmaceutical industry, as food, in cosmetics, and in bioenergy and nanotechnology. Between 2009 and 2013, an international research expedition called Tara Oceans collected 35,000 biological samples from 210 locations in the world’s oceans. It is the largest plankton collection in existence.
ELIXIR built a permanent, public data resource to identify and chart metagenomics samples obtained from the sea. The tools needed for identification and the pipelines for data processing were made available for transfer to different platforms. This could result in the introduction of new biochemical materials, such as enzymes and medicinal molecules. The tools and data pipelines can be used through various ELIXIR centres (Norway, EMBL-EBI, Finland, Czech Republic and France).
The European Genome-phenome Archive (EGA) is one of the world’s most extensive public data resources and holds patient data gathered from biomedical projects. The archive contains various data resources from different data producers. The EGA collects human genome and phenome data on the basis of the consent of the persons involved. ELIXIR Compute Platform enables the transfer of confidential human data via the EGA to individual users authorised to access such data.
Through the ECP, researchers can access the EGA’s sensitive data collections. First the user is identified electronically, and access is either granted or rejected on the basis of the information on the application form. If the service requires multi-factor authentication, the user is redirected to an identification service, which performs an extra authentication by means of another security feature.
After this, researchers have access to EGA data resources and can process sensitive data. Through the ECP, researchers can also store data in the EGA archive. The ECP can ensure data description, access to data and compatibility. In order to transfer data securely, an architecture was created that uses two protocols. Oauth.2.0 and OpenID Connect (OIDC) are user identification protocols used in the industry.
According to the FAO, plant diseases cause an annual loss of 20–40% in global food production. Massive sequencing of cultivated and forest plants enables the causes of plant diseases to be studied. Plant sequencing and genotyping, including pathogens and diseases, generate large amounts of genetic variation data. EURISCO (European Search Catalogue for Plant Genetic Resources) contains information about 1.9 million cultivated plants and their wild relatives. The samples have been collected by almost 400 organisations.
The ECP enables genotype-phenotype analysis of cultivated plants, based on the widest available public data resources. This data has been assembled from geographically separate research institutions. The key function is a search robot that receives searches from users, to whom it transfers integrated search results obtained from various data sources. Users can transfer the selected data into the cloud infrastructure for analysis.
The European Organisation of Rare Diseases (EURORDIS) estimates that some 30 million people in 25 EU countries have a rare disease. This translates into 6–8% of all people in the EU. The International Rare Diseases Research Consortium has set itself the target of developing 200 new forms of treatment for rare diseases by 2020.
ELIXIR has published a customised collection of tools and services to help in the development of new treatments. The collection is available through the ELIXIR biotools service (bio.tools). Researchers of rare diseases can deposit raw data, run a gene mapping and pick gvcf files (genomic variant call format) for analysis. This defines the text file used in bioinformatics when gene sequence variations are stored.
The patient’s metadata (illness, treatment, treatment results), patient samples in biobanks and all EGA can be obtained through the ECP.
Ari Turunen
2.12.2019
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2019). ELIXIR Compute Platform for life and health sciences. https://doi.org/10.5281/zenodo.8131182
Additional information:
Kataja, Teemu (2018): Designing and developing a data processing pipeline for archiving sensitive human data.
https://www.theseus.fi/handle/10024/142007
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Laboratory technologies evolve all the time. New measurement instruments are producing a huge amount of data that needs to be analysed correctly. Being in a continuous state of change and development, biomedicine needs experts who can train researchers in the latest analysis methods.
For several years, ELIXIR Finland’s training coordinator, Eija Korpelainen, has been teaching researchers to analyse data.
“Life science and medical degrees don’t include much data analysis studies. However, the use of modern measurement techniques requires solid data analysis skills, which is why we give courses at CSC. Different types of data require different analysis methods, so a number of courses are required. ELIXIR also organises courses for trainers.”
The content and teaching material of these courses are available via ELIXIR’s TeSS training portal. The portal includes bioinformatics courses offered by ELIXIR’s member organisations. Courses and course material are also provided by non-ELIXIR organisations.

Courses must be updated regularly owing to the rapid development of bioinformatics methods. This puts pressure on the development of CSC’s Chipster analysis software, because new analysis tools must be added continuously. Korpelainen says that even the name of the software should be updated:
“Chipster refers to the microarrays or “chips” used to analyse the activity of genes. Gene activity is nowadays measured with high-throughput sequencing (HTS). HTS refers to new equipment and techniques that enable faster and more efficient sequencing.”
HTS techniques can be used to study a number of topics. By sequencing exomes, i.e. the genomic areas that code proteins, it is possible to learn about gene variants related to illnesses.
The Chipster software consists of more than 400 analysis tools and an extensive collection of reference data. Chipster can be used to analyse, visualise and share data interactively.
“Measurement technology is changing all the time. In addition, because data comes in various types, it must be analysed in different ways,” says Eija Korpelainen.
As a platform, Chipster can be used for the full range of scientific disciplines, because it can be integrated with any analysis tool. For example, CSC has a separate version of Chipster for linguistic analyses.
Nature magazine selected the sequencing of single cells as the method of the year in 2013; this technique has since become available to all researchers as measurement technologies have improved. The RNA-sequencing of a single cell (scRNA-seq) can reveal complex and rare cell populations. It measures the expression levels of all the genes in each cell separately, thereby providing a more accurate picture of the differences between cells.
The sequencing of a single cell can be used when analysing changes in a transcriptome (set of messenger RNA molecules in one cell). The first analysis of this kind was published in 2009, describing cells at the early stages of development.
“Expression profiles can be used, for example, to reveal cancer cells or obtain information about drug resistance or the response to cancer treatment.”
Chipster software offers a range of scRNA-seq analysis tools.
http://bit.ly/scRNA-seq
scRNA-seq and HTS techniques are good examples of how rapidly research is progressing thanks to inventions. In order to make the most of the new techniques, researchers must be able to use the latest methods to analyse the data they produce. This means that a lot of training is needed, as indicated by the courses provided by ELIXIR.
“We must regularly update existing courses and develop new ones,” says Eija Korpelainen, who goes on to explain that keeping up with bioinformatics methods is both challenging and interesting.
“It is interesting to talk to various experts and read articles that impartially compare various methods. Data analysis in the life sciences and medicine is now so challenging that it attracts data scientists to the field.”
Ari Turunen
25.11.2019
Read article in PDF
Citation
Ari Turunen, Eija Korpelainen, & Tommi Nyrönen. (2019). New bioinformatics methods and measurement technologies call for continuously updated courses and analysis software. https://doi.org/10.5281/zenodo.8131176
More information:
https://tess.elixir-europe.org
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

In order for researchers to access the digital services of various research infrastructures, their identity and relationship with the research organisation must be verified. Until now, this has required a personal visit to a registration point where their identity document has been checked. Finland has been testing a new solution for strong electronic identity proofing that does not require a personal visit to a registration point.
Authentication ensures that the person is who he or she claims to be. At the moment, researchers can use their home organisation’s credentials when logging into infrastructure services. Logging into services containing sensitive data, for example, nevertheless requires a more reliable authentication method, but these are not available in all home organisations.
“Traditionally, identity proofing is considered reliable if a person has to visit a registration point in person, with trained personnel checking his or her passport or other identification document issued by an authority,” says Senior Application Specialist Mikael Linden of CSC.
Together with the , the ELIXIR Finnish centre, CSC, has long been developing authentication services for infrastructure. ELIXIR’s AAI service (Authentication and Authorisation Infrastructure) enables electronic user authentication and the authorisation. Access to gene data, for example, is always decided by the data owner, but the AAI will make access to data quicker.
The AAI service is effective but requires that the researcher is reliably authenticated. In general, the solution is a federated identity management, which is simple and easy to manage. A single log in and your home organisation’s user ID also provide researchers with access not only to services outside their organisation, but also to closely protected data collections. What if your home organisation cannot provide sufficiently reliable authentication?
CSC has been working together with the Sandbox of Trust project, which also involves the cybersecurity services company Nixu. The project has developed the SisuID authentication solution, the purpose of which is to create a more user-friendly alternative to passwords and strong two-factor authentication. A user uses a smartphone application to enrol their unique identity. A combination of these also enables the secure transfer of personal data between services, based on the person’s own approval.
“In a research infrastructure like ELIXIR, identity proofing in a network of registration points would be expensive and cumbersome for end users. With the SisuID concept, identity proofing is carried out by taking a photo of themselves and their passport with the SisuID mobile phone application, which will check that the two match,” says Mikael Linden.
SisuID is an open source code identification method that has been tested in five different pilot projects. According to Joonatan Henriksson, Head of Digital Business at Nixu, various ways have now been tested of reliably registering and identifying Finnish and foreign persons.
“In Finland, strong electronic authentication can currently be carried out using bank credentials, but this is not possible for foreign researchers, and not all countries have a national strong authentication method in place,” says Mr Henriksson.
He points out that, with the cross-border solution now being tested, persons performing identity proofing will use their mobile device to take a photo of their passport or ID card, and a photo of their face. These are compared algorithmically.
“The comparison can also make use of registers available in the country that issued the identity document, and, for example, Interpol databases for forged identity documents.”
But there are also more stringent identification criteria.
“If the service provider does not consider the remote identification sufficient, we can increase the identification reliability level by having the person visit a physical location for identity proofing, after which the more reliable identity will be available to all service providers using SisuID.”
According to Henriksson, the identification criteria will comply with the eIDAS EU regulation. The eIDAS regulation creates a framework for providers of identification services, on which, for example, the Finnish Act on Strong Electronic authentication and Electronic Signatures is based. By complying with the eIDAS regulation, providers of electronic authentication and signatures can apply for official approval of their services, qualifying the authentication method for cross-border central government transactions.
“We will also be able to access a facial image on a passport’s NFC chip, signed by the party that issued the passport, and take a liveness video of the face, further increasing the electronic reliability of the registered identity.”
To produce the SisuID solution, a non-profit identification cooperative is being set up that will divide the benefits, costs and risks of the identification service between the organisations that use it.
Once a person can be authenticated efficiently and reliably, the only problem that remains is that the data related to the person is located in silos. For the moment. Access to services provided by ELIXIR, for example, could be granted by combining two pieces of data: a reliably registered digital identity and assertions related to a person. A researcher can convert a university’s assertion on their affiliation or an EU grant decision into electronic format, making their research status official.
“In future, such electronic data linked to an identified person’s digital identity could be sent between actors by means of distributed, confidence-building, cross-border blockchain networks.”
As the name implies, data is stored in blocks in blockchains. A block is connected to the previous one with an algorithm that turns the data into a character string. Data entered in a single block cannot be changed afterwards, because the blockchain has been distributed to several computers.
This method enables the digital distribution of confidential data without having to reveal the interfaces of national registers, for example. The fact that blockchains cannot be changed ensures reliable data transfer by the users themselves, which means that no direct integration between the interfaces is required. Examples of this include EU-level testing in the European Blockchain Services Infrastructure (EBSI) project concerning the transfer of electronic educational certificates.
Ari Turunen
30.10.2019
Read article in PDF
Citation
Ari Turunen, Mikael Linden, Joonatan Henriksson, & Tommi Nyrönen. (2019). No need to turn up personally: SisuID improves electronic authentication. https://doi.org/10.5281/zenodo.8131086
For more information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Cardiovascular diseases are the most common cause of death in the world. More than a third of deaths in Finland are caused by cardiovascular diseases. The current objective is to create an assessment, based on health data, of each person’s risk of illness before they consult a doctor.
Andrea Ganna, Group Leader from Institute for Molecular Medicine Finland FIMM at the University of Helsinki and instructor from Harvard Medical School, wants to establish a nationwide, personalised risk assessment as foundation for planning public health interventions. The assessment is based on the health, demographic and genetic information of the citizens. The assessment, which uses artificial intelligence, improves the allocation of preventive treatments with a lower cost than today.
”Nordic countries and specifically Finland have a unique opportunity and setting, since they have been collecting health and demographic data for years. But the way they have used this data in the past is somewhat outdated. Only very specific correlations and associations in the data have been looked at. However, new methods, such as AI, are emerging, which now allow us to push for a much bigger and ambitious vision.”
Andrea Ganna and his group is developing artificial intelligence (AI) approaches to model health trajectories.
”You have a certain health trajectory and have taken certain medication. We ask if there are other people who have followed a similar path. There may be thousands out there. We leverage those people and ask: What happened to those? Let’s take that experience and bring it back to you to help you to reduce your disease risk. We can use all this data in a more comprehensive way to help public health and give more information to patients and doctors for decision-making.”
Andrea Ganna is interested in epidemiology, genetics and statistics. He has been focusing on leveraging large-scale epidemiological data sets to identify socio-demographic, metabolic and genetic markers of common, complex diseases. In Boston he worked with large-scale exome and genome sequencing data.
According to Ganna, cardiovascular diseases are ideally suited for analysis by artificial intelligence, since their treatment is preventive.
”Accurate identification of individuals at high risk is one of the cornerstones of primary prevention of cardiometabolic diseases,” he says. ”However, at the moment, risk factor assessment for cardiometabolic diseases requires patients to go to the doctor for lipid measurement. ”
Lipid is the umbrella term used for all fatty acids or their derivatives that circulate in the blood. The body stores fats from food for later use. A diet rich in fats will cause them to attach to the walls of the arteries, leading to cardiovascular and arterial diseases. Lipid measurement is effective, but some members of the population are unaware that they belong to a risk group.
Ganna wants to revolutionize primary prevention by providing risk assessment before an individual even steps into the doctor’s office.
”Some simply don’t go to the doctor’s and so lots of people are missed. However, since all the data on medication and diagnoses has already collected, we can identify high-risk patients without them going to doctor. We can make a risk map of cardiovascular diseases of the whole country by including every individual.”
Calculating the risk is done by modelling longitudinal histories of diseases and medications with the gene data, family and demographic data.
”We are trying to understand how genetics interact with data regarding medications, diagnoses, demographics, and familial risk. This can provide an unprecedented holistic view of an individual’s health status.”
Ganna gives an example.
”When you break your leg, you go to the doctor. However, today the doctor is just looking at the leg, although during the same visit other information could also be obtained. We can inform the doctor about other risks the patient has based on the collected data. We can precompute the other risks of the patient, for example if this patient has also high risk for cardiovascular diseases. Thus, during the visit, the doctor can also give advice or refer the patient to a specialist.”

Ganna chose to come to Finland because of the large genetic project, FinnGen. The FinnGen project will record the genomes of half a million Finns. The project, launched in August 2017, utilises samples collected by all Finnish biobanks. The data from genomes is combined with the information in national health care registers. FinnGen is one of the very first personalised medicine projects of this scale and the public-private collaborative nature of the project is exceptional.
“Finland also has a favourable legislation, giving access to nationwide population data. For me, this is a unique setting,” says Ganna.
Ganna and his research group integrate registry-based information with genetic information from large biobank-based studies (e.g. FinnGen) to help identify groups of individuals that can most benefit from existing pharmacological interventions.
”Probably the most important population is younger individuals who do not see a doctor very often. Current risk factors do not work well in this group. Genetics is particularly valuable because it can capture disease risk at an earlier age than other risk factors,” says Ganna.
”The first step is to understand how people perceive this information. We have to ensure that doctors use the data in a right way and what can be done with it.”

Ganna aims to integrate national and regional registries with deep and machine learning.
”Traditional methods have an advantage since they are relative simple and easy to interpret, but they simple do not scale. In the past 20 years, more than 500 million medical diagnoses have been made of Finns. We are talking about huge data sets. Every year there are millions and millions of new medication purchases and diagnoses. To scale and to leverage this massive data, deep learning methods are needed.”
Artificial neural networks are efficient machine learning algorithms, which can be used in pattern recognition. Recurrent neural networks can use their internal memory to process sequences of inputs. This makes them applicable to tasks such as unsegmented recognition. Ganna wants to expand long short-term memory recurrent neural networks to data.
”You can imagine the sequence of health events that we are trying to model as “text” in which each word is a different disease, medication, sociodemographic event etc. across the lifetime. These are naturally suited to model the sequential happening of events, for example they are used to predict the next most likely word in a text message.”
Deep learning methods need large supercomputing infrastructure.
“CSC has created a secure environment for this computation. Without a secure supercomputing environment, we could not carry out this project. To be successful, we need, on the one hand, research and development, and, on the other hand, a powerful computing environment.”
Patient data is important for research, but personal data is also protected. For example, VEIL.AI application created by FIMM anonymises patient data better and faster than traditional methods, and retains information more effectively. If necessary, the application can also produce synthetic, fully anonymous statistical data, which cannot be traced back to any individual.
“We need to guarantee individuals’ privacy but, at the same time, we need to integrate a lot of personal data to really leverage the power of artificial intelligence/deep learning approaches to better target public health interventions. Generating synthetic health trajectories will help to respect privacy and, at the same time, to combine a lot of personal information within Finland, but also across Nordic countries.”
”My hope is that personal data that is routinely collected in healthcare can help and benefit everyone. My hope is that that this information can help doctors to make better decisions, but also help patients in motivating life style changes. Thus everyone is helping everyone.”
Ari Turunen
30.9.2019
Read article in PDF
Citation
Ari Turunen, Andrea Ganna, & Tommi Nyrönen. (2019). Risk assessment of cardiovascular diseases for all citizens. https://doi.org/10.5281/zenodo.8131074
More information:
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Life Science AAI attempts to make logging in and access to available services as simple as possible.
ELIXIR has created easy-to-use and secure user ID management, enabling access to numerous data collections and services. The service is being developed and expanded in cooperation with other research infrastructures, so that researchers can also have access to biological samples offered by biobanks or compiled from test animals.
Data resources and research instruments needed by researchers are available in numerous research infrastructures. Although international cooperation within research has intensified during the past decade, researchers must nevertheless deal with various administrational processes during the course of their work. Access to material created by research often requires user identification and access permissions. If each data collection requires its own password and user ID, managing them all becomes too cumbersome for individual users. There must be a way out of the password jungle without compromising the data security of the service, or the user’s own rights covered by the General Data Protection Regulation. The ELIXIR research infrastructure has the objective of ensuring easy use of data collections without compromising data security.
ELIXIR’s AAI service (Authentication and Authorisation Infrastructure) enables electronic user identification and access rights management. Access to data is always granted by the data owner, but AAI will make access to data quicker, since the use policy is clear and straightforward.
The solution is a federated user ID management, which is simple and easy to use. A single identification with their home organisation’s login also provide researchers with secure and reliable access to closely protected data collections.
Federations allow researchers to access their home organisations’ user IDs. They provide them with access to services outside their own organisations. The idea behind federations is to manage how user IDs are transferred during login across organisational boundaries. Various user rights of different levels can be coupled with the identity in question, to ensure that the user can access the correct resources for their legitimate reasons.
Federated user ID management is by no means a new invention. It has been used successfully by organisations such as the Haka identity federation of the Finnish higher education institutions. The Haka federation consists of more than 300 services and has more than 300,000 users.
The EU-funded eduGAIN service, which combines various federations, was established in 2004. In April 2011 it became a permanent service that combines research and education identity federations around the world. eduGAIN brings together more than 50 federations, consisting of 5,000 organisations. It is open to all of the world’s academic federations, enabling reliable user login among federation members.

The ELIXIR AAI was launched in November 2016. It is part of ELIXIR’s Compute Platform, together with cloud and data transfer services. In late 2018, the ELIXIR AAI service had 2,174 users and an average of 3,200 log-ins a month.
In late 2018, researchers who had logged into the ELIXIR AAI service were able to log into another 50 services connected to the ELIXIR infrastructure. Another 44 services were tested, some of which were offered by other major European research infrastructures. One of these was the EGI (European Grid Infrastructure), which was tested to see if it could access the fedCloud computing service. The number of services is rising all the time.
The ELIXIR AAI service was developed by the Finnish and Czech nodes of the ELIXIR infrastructure. The service not only provides authentication services, but also permissions granted by the material owners. In Finland, the National Institute for Health and Welfare (THL) was the first to test the process, based on ELIXIR AAI’s federated authentication and permission management, using sensitive material from their biobank samples. THL’s biobank is part of the BBMRI infrastructure and access to the material is an example of cooperation between two European research infrastructures aimed at making researchers’ lives easier.
The idea is that users register one ELIXIR identity and continue to use it throughout their careers. All they have to do is update their contact and personal details if these change. It is not practical to maintain more than one ELIXIR identity. An ELIXIR identity does not have a password. During registration, all you need is a connection to an academic or commercial user account that is used for logging in.
ELIXIR AAI already accepts Google, LinkedIn or Orcid as part of identification. Through Orcid, researchers obtain a digital identity that enables them to distinguish themselves from colleagues by the same name. ELIXIR AAI supports also 721 institutional logins via eduGAIN.

The challenge for federations lies in the fact that there is no commonly agreed definition for various levels of assurance for identities and authentication. Privacy legislation makes some institutions wary of sharing their researchers’ personal data with other jurisdictions.
The requirements for user authentication and the management of user authorisation are tighter when dealing with controlled and sensitive data collections. Users’ access rights may have to be categorised, for example. ELIXIR experts are cooperating with other research infrastructures in the EOSC-Life project, which is assessing various user cases in the biological sciences sector to create a common and extensive federated identity management service. This service is called Life Science AAI, which utilises the eduGAIN federation for identification.
Due to the increased need for federated access between research infrastructures, many projects are helping to create common user management. The AARC (Authentication and Authorisation for Research and Collaboration) project was launched in May 2015. The project’s second phase (AARC2) was launched in May 2017 and ended in April 2019. The project piloted integrated authentication and authorisation between organisations.
The objective is that each new user will only register a single user ID that follows them throughout their career, even if they change jobs and connections. Because universities and research institutions have connections to several research infrastructures, researchers will have automatic access to them via their own organisation’s account. The objective is to create a reference model to manage not only identity (registration, identity proofing) and authentication (logging in), but also other aspects such as researcher status.
Cooperation with parties such as Federated Identity Management for Research Collaboration (FIM4R), on the other hand, is aiming for the creation of common standards in order to meet the needs of various research communities. Another key partner is GA4GH.
GA4GH (Global Alliance for Genomics and Health) is an international alliance founded in 2013 consisting of more than 500 bioindustry, healthcare and IT organisations, with the objective of creating standards for genomic data distributed for research use. ELIXIR and GA4GH decided to start a partnership in November 2017. The agreement gives the ELIXIR infrastructure a chance to contribute to the creation of international standards. The agreement is related to the project, the purpose of which is to make the data standards available for clinical patient work by 2022. Now work can be done with over 1,000 organisations not only on standards, but also common principles on how data is processed and distributed.
The challenge is to define the criteria that an organisation must fulfil to become a reliable partner in a global alliance. Registered access ensures the categorisation of various users. It also enables data reuse, but naturally only if consent has been obtained and users adhere to their ethical commitments.
The Global Alliance for Genomics and Health has created three options for accessing human data. These are:
1.No need to control access
2.Registered access based on the user’s role as a researcher
3.Controlled access based on the user’s specified access permit.
“Because data is private, federations are controlled by strict data security and regulation, such as the GDPR. ELIXIR’s member organisations adhere to European policies in terms of data protection. However, because research is global, the EU wants to share research data with Canada and the US,” says Tommi Nyrönen, Head of Node of Finland’s ELIXIR Center. Finland’s ELIXIR Center CSC has been building the ELIXIR AAI service alongside the Czech ELIXIR Centre.
“This is why we must manage user information within European and, say, alongside North American organisations. We must have common agreements on how data can be transferred for research purposes in accordance with regulations. Parties responsible for data need sufficient information on users who are requesting access. Only when the user’s identity and potential home organisation and status as a researcher have been ascertained, can the data access application be processed. We must also have a mechanism to stop and cancel access to data quickly if it is used for the wrong purposes. This can be done, for example, with a policy and technology specified by ELIXIR AAI.”

ELIXIR AAI is a service which researchers can use to request access to sensitive data collections. Users can substantiate their researcher status. To register your researcher status and personal identity in the ELIXIR infrastructure, you must first log in to your home organisation, which will submit your up-to-date user details during the log-in process. The registration may contain additional information, such as the category “bioindustry researcher”.
The person in charge of the research project fills in the application form on behalf of the other project participants, and accepts the data collection licence terms. An electronic application form is sent to the Data Access Committee, chosen by the Data Manager, that supervises data access rights. Access is either granted or rejected on the basis of the information on the application form.
If the service requires multi-factor authentication, the user is redirected to a speedy and more efficient identification service, which performs an extra step-up authentication by means of another security factor. Step-up authentication is based on a Time-based One-Time Password (TOTP) and a smartphone application that is registered in the ELIXIR AAI service. Once a user has registered, the TOTP application provides a six-character one-time password, which the user must enter in their browser.
The smartphone application is connected to the correct ELIXIR identity using a text message sent to the phone.
Data owners can cancel or review a user’s access rights.Organisations that trust ELIXIR can register their own data collections in the authorisation infrastructure and specify the application forms and the related processes.
Effective data security is based on a risk analysis of the requirements of the materials and the nature of the service. For example, researchers can be provided with access to data collections, which have a limited impact on privacy issues, through a lighter application process. This means that all researchers have to do is to prove that they are bona fide researchers and adhere to the general commitments concerning registration.
When they had completed an application, researchers could access all data collections and services available for bona fide researchers, with no extra effort. The ELIXIR Beacon service is an example of such an access process. The Beacon protocol defines an open standard. A website that offers such a service is called a beacon. Beacon is a search engine for finding the location, anywhere in the world, of genome material that contains, say, an interesting change in nucleotide such as in a gene sequence that codes protein, with cytosine (C) changing into guanine (G).
“This change may alter the structure of the protein generated by the gene. Sometimes, such changes are harmless, but they may also lead to illness. The connection between genetic modification and rare diseases is being actively studied, and results could be achieved faster by making material available through Beacon,” says Tommi Nyrönen.
The standard and technology have been developed by GA4GH’s member organisations. Data searches can be performed on the same principles in ELIXIR and the Beacon Network. Searches are federated and the number of data collections is continuously growing.
Ari Turunen
20.8.2019
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2019). Federated user ID management: a single identity giving access to numerous bioinformatics services. https://doi.org/10.5281/zenodo.8176724
More information:
Registered access: authorizing data access
European Journal of Human Genetics (26,2018)
https://www.nature.com/articles/s41431-018-0219-y
Common ELIXIR Service for Researcher Authentication and Authorisation
F1000Reserarch (7, 2018)
https://f1000research.com/articles/7-1199/v1
CSC – IT Center for Science Ltd
CSC – IT Center for Science Ltd is a state-owned, non-profit public limited company run by the Ministry of Education and Culture. CSC maintains and develops the state-owned centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR is building an infrastructure to support research and the bio sector. It combines the leading organisations of 21 European countries and the European Molecular Biology Laboratory (EMBL) into a single infrastructure for biological information. Its Finnish centre is CSC – IT Center for Science Ltd.
https://www.elixir-finland.org
http://www.elixir-europe.org

Lauri Eklund does research on disturbances in the growth and functioning of blood vessels. The aim is, among others, to develop targeted treatments for venous diseases.
Venous diseases mean chronic and progressive vascular changes for reasons that are often unknown. Disturbances in the venous circulation, for example, are common in the retina of the eye, causing impaired vision. Venous malformations, on the other hand, represent rare congenital disturbances in blood vessel formation often caused by somatic mutations in genes that are important for normal venous morphogenesis. Professor Lauri Eklund’s research group aims to identify physiological mechanisms which control the differentiation of blood vessels into veins, and cellular and molecular defects may underlie vascular abnormalities.
“The basic mechanisms of capillary formation and differentiation of arteries are known relatively well, whereas there are still a lot of open and interesting biological questions with the differentiation of veins. In addition, revealing molecular mechanism causing vascular diseases is necessary to the development of targeted therapies”
Based at Biocenter Oulu and at the Faculty of Biochemistry and Molecular Medicine, Eklund does research on the mechanisms controlling the development of veins using genetically modified mice and endothelial cell cultures to model mutations identified in patients and phenomena related to vein formation.
Disturbances in venous circulation in the retina of the eye is a relative common cause of impaired vision. Eklund’s research group has recently discovered the first growth factor that is needed for the formation of veins in the retina of a mouse. The growth factor is a protein which stimulates the growth or differentiation of cells. The growth factors also function as signal molecules between cells.
“The importance of the growth factor studied by the group, angiopoietin-4 (Angpt4), has been poorly known until this day. It controls the functioning of TIE2 receptor tyrosine kinase in endothelial cells which cover the inner surface of blood vessels.”

Identification of the TIE2/Angpt-4 cell signalling pathway in normal development of veins is, according to Eklund, a very interesting finding.
“The somatic mutations leading into gain of function of the same TIE2 receptor cause a major proportion of the venous malformations which occur in humans.” Another prevalently mutated gene is PIK3CA, which corresponds to the activating subunit of PI3K lipid kinase.
Both TIE2 and PI3K are kinase enzymes which control the functioning of their target proteins by phosphorylation. The cells use this mechanism in their normal cell signalling.
“In collaboration with Miikka Vikkula’s group (the de Duve Institute, Brussels, Belgium) we have demonstrated that the cell membrane receptor TIE2 and the downstream signal transducer PI3K are located in the same signalling pathway causing venous malformations. The switch-on in TIE2 or PI3K active kinase domains triggered by a mutation will cause the same kind of abnormalities in endothelial cells and vascular structures. The impact of mutations is that they activate cells, causing a false, growth factor independent and uncontrolled locking-on of the TIE2/ PI3K signalling activity.”
According to Eklund, these observations together refer to the Angpt-4/TIE2/PI3K signal route playing a specific role in vein formation in development and disease. Venous malformations also have a surprising connection with the cancer. In DNA certain nucleotide sequences are known as “hot spots”, where mutations concentrate in some diseases.
“The same hot spot mutations in the activating subunit gene (PIK3CA) of lipid kinase PI3K cause not only a significant proportion of venous malformations, but several cancers as well. It is very interesting, yet at the same time poorly understood, how the same PIK3CA mutations cause various malignant cancers in epithelial cells and “only” excessive growth of vein-like channels when occurring in the endothelial cells.”
“Some of the developmental defects in blood vessels are mostly cosmetic “birthmarks”, which can also heal on their own. According to Eklund, one example of problems of this kind are the congenital hemangiomas, such as strawberry naevuses in which the originally excessive endothelial cell division and capillary formation reduces as the child grows older. Hemangiomas are, in fact, the most common benign self-healing tumours, which are typically identified in children at the age of a few weeks. In many of the patients, they have spontaneously regressed by the age of five years.
“What factors cause the cessation of excessive prolifearion of endothelial cells is an interesting, yet not well understood phenomenon,” Eklund says.
Although some of the diseases under research are rare, they can help us understand the disease mechanisms and uncover typical control mechanisms related to the growth and functioning of blood vessels.
“Even though vascular developmental defects have mainly to do with rare diseases, they can also occur in veins, capillaries or lymphatic vasculature. As venous malformations, most of them may likewise originate from gene mutations, yet through other cell signalling routes. Considering the various types of vascular development defects, they are relatively common together.”
After the defective molecule has been identified, it is possible to develop targeted molecular treatments.
“These include small synthetic molecules that are aimed to only target abnormally functioning molecule and do not cause any disturbance to the rest of the events in the body. The efficacy of the drug improves, and there are fewer side effects.”
According to Eklund, identifying the disease mechanisms may also lead into repurposing the drugs for illnesses which they were not originally designed for. One example is, for example, the PIK3CA inhibitor Alpelisib.
“Although originally developed as cancer medication, it is also efficient in preventing the development of venous malformations in cell and animal models. The genetic change causing the disease is the same in both cases, but the mutation takes place in different cell types.”
Since pharmaceutical companies may not have interest in developing drugs for small patient groups, this kind of ‘repurposing’ is, as Eklund sees it, often the only opportunity for patients suffering from rare diseases to receive targeted medicinal treatment.
In rare diseases, identification of the mechanisms causing the abnormalities may also uncover yet unknown cell signalling pathways that are involved also in normal growth.
“In fact, many cell signalling pathways and mechanisms which are dysregulated in vascular diseases, are also needed with normal vascular development. One of these is the angiopoietin/TIE2/PI3K pathway that has been under investigation in our research.”

The earliest vascular structures in embryos originate from cells known as angioblasts which can differentiate into endothelial cells to form blood vessel. During embryonic development and growth, new vessels are formed from existing ones in a process known as sprouting angiogenesis. In the following maturation phase, the vascular system differentiates into a hierarchic network of functional arteries, veins and capillaries, which is needed for keeping tissues homeostasis. In healthy adults angiogenesis is needed, for example, in recovering from injuries and also occurs for example, in conjunction with menstruation. New formation and changes in the structures and functioning of blood vessels are also related to many diseases. For example, solid tumours are dependent on blood vessels which feed the tumour with oxygen and nutrients. Blood vessels which have undergone structural changes may accelerate the transition of tumor cells forming metastases into other tissues and so contribute to a disease becoming malignant.
“The neoangiogenesis of a tumour means growth of blood vessels in a solid tumour from the tissue surrounding it. Interestingly, same growth factors that are needed in development also play a role in adverse neoangiogenesis. This kind of growth factors are interesting targets for the generation of pharmaceutical agents. We have investigated one of them, angiopoietin-2, which can affect the vascular structures in many ways.”
The extracellular matrix (ECM) is a three-dimensional network of macromolecules between the cells and also supporting the vascular system. Perivascular ECM consists of a specialized, thin layer of macromolecules called basement membrane and in larger vessels fibrillar collagen matrix. Collagens are the largest group of proteins in the extracellular matrix and in the human body as a whole.

Oulu has a long tradition in ECM and collagen research, and one of Eklund’s aims is to investigate how the endothelial cells and the vascular smooth muscle cells interact with the ECM. Better understanding of the role of the perivascular ECM may help to explain developmental anomalies as well as defects of the vascular system in cancers.

The endothelium is an inner single-layer of the blood vessel and also covering inner surface in the heart (endocardium) and lymphatic vessels. Endothelial cells isolated from the organs can be utilised in in vitro studies on culture dishes in laboratories. This enables various tests which cannot be done in living animals or humans,
“The models allow us, for example, to study the effect of growth factors on cells: does the vitality of the cells improve? Do they begin to differentiate or form vascular structures?
The cell culture models are also used to study mutations found patients. For example, they can be investigated to see how cells change when they express a mutated form of a gene. In 3D cultures, endothelial cells can be used to grow structures which resemble blood vessels.
According to Eklund, the opportunities for manipulations and research methods in cell-culture models are almost limitless.
“To better understand the normal development and diseases, often very complex entities can be divided into smaller or single events. Another benefit of cell culture models is that cells of human origin can be used. In some cases, the treatments developed using non-human systems do not have the same effect in the patients. In other words, the results gained e.g. with mice are not necessarily transferred to humans as such. Cell culture models also reduce the need for animal testing, and they are not subject to similar research-ethical considerations. For example, endothelial cells of human veins are derived from umbilical cords of volunteer donors after childbirth.
In Eklund’s research group, the formation of vascular structures are also modelled in a 3D environment.
“Even though they imitate tissues better than two-dimensional cultures on plastic dishes, they still fail to have the natural flow of blood vessels”.
In addition to European partners, Lauri Eklund’s research group is collaborating with the FICAM centre at the University of Tampere and an Oulu-based start-up company FinnAdvance in a project financed by the EU.
One purpose is to develop “microfluidic” flow channels coated with human cells and perivascular ECM designed to correspond to normal blood vessels or abnormal structures found in vascular anomalies. The aim is to use the devices to study what the changes in blood flow cause in the vascular system and especially in their malformations, where the blood flow has undergone significant changes.
In many case animal models are still needed for verifying the results.
“If one uses excessively simplified models, the results may become unreliable. Therefore, animal models are still needed for verifying the findings in a more complex environment, which better corresponds to the human tissue. Currently, the mouse is the best one out of the mammalian models for this kind of research. What makes the mouse models the best is that many genetic modifications are feasible to generate, including the expression of the same gene mutation which causes diseases in humans.”
The research is financed by the Academy of Finland and the EU H2020-MSCA-ITN programme.
Ari Turunen
5.9.2019
Read article in PDF
Citation
Ari Turunen, Lauri Eklund, & Tommi Nyrönen. (2019). Targeted treatment for venous diseases with vascular system modelling. https://doi.org/10.5281/zenodo.8131049
More information:
Biocenter Oulu
Biocenter Oulu is part of Biocenter Finland, which coordinates infrastructural activities for major national research. It is also a member of various European research infrastructures. These are Infrafrontier (transgenic mice), Euro-BioImaging (biological imaging) and Instruct (protein structure research).
https://www.oulu.fi/biocenter/
CSC – IT Center for Science Ltd
CSC – IT Center for Science Ltd is a state-owned, non-profit public limited company run by the Ministry of Education and Culture. CSC maintains and develops the state-owned centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR is building an infrastructure to support research and the bio sector. It combines the leading organisations of 21 European countries and the European Molecular Biology Laboratory (EMBL) into a single infrastructure for biological information. Its Finnish centre is CSC – IT Center for Science Ltd.
https://www.elixir-finland.org
http://www.elixir-europe.org

The population in the north of Finland has a unique genetic background. This has enabled the study of rare diseases and the discovery of new genes, proteins and reaction paths.
Docent Reetta Hinttala from the University of Oulu and Johanna Uusimaa, Professor of Pediatric Neurology, have discovered mutations in the NHLRC2 gene as a cause of a severe childhood multiorgan disease. Currently, the function of NHLRC2 in cells is unknown, but it has been identified as crucial to embryonic development.
Reetta Hinttala studies hereditary neurological childhood diseases at the University of Oulu’s Faculty of Medicine. The children suffering from FINCA disease were found to have previously uncharacterised connective tissue formation in their lungs, degeneration of neuronal cells, and increased angiogenesis in the brain. The disease manifested at the age of two months, and progressed quickly.
“Since this was a new combination of symptoms, we suspected that it must indicate a new disease,” says Reetta Hinttala.
The patients’ exomes, i.e. genomic areas encoding proteins, were sequenced. Based on what we currently know, the majority of variants that cause diseases are within those areas that only cover around 1.5% of our genome.
“Exome sequencing revealed changes, variants, in the nucleotide order of the NHLRC2 gene. At that time, this gene or its variants had not been linked to any human disease or even described in scientific publications. So we started studying the protein encoded by the gene, particularly focusing on its function in cells.”

When you want to find out how a disease emerges and develops in humans, the only way of doing this is to study it in living organisms. Only then can we monitor what the protein that causes a disease is doing in the organism. In this particular case, we created a mouse model with the same mutation combination as the patients. A knock-out mouse, in which the NHLRC2 gene has been completely turned off, was obtained from the EMMA repository of the Infrafrontier infrastructure. EMMA (The European Mouse Mutant Archive) archives genetically modified mouse strains from around the world. In addition to this, Hinttala and her team used the CRISPR-Cas9 technology to create the identical point mutation that was observed in FINCA patients, to the mouse. By crossing the mouse carrying point mutation with the knock-out mouse, a model was created that has the same mutation combination as the FINCA patients.
“We are using our mouse model to determine the role of NHLRC2 in the development of the central nervous system in particular, through a project funded by the Academy of Finland. At tissue level, we have observed encouraging signs that the mouse model is exhibiting features similar to the FINCA disease, but it will still take some time to build a comprehensive picture of the phenotype.”
Cellular changes resulting from disturbances in the gene causing the disease were studied in the patients’ fibroblasts, using a transmission electron microscope, at Biocenter Oulu’s electron microscopy core facility. Fibroblasts are the main cell type present in connective tissue and they are specialized in secreting extracellular matrix proteins.
“The first indication that a mutation, or variant, is harmful is when the variant changes the protein’s amino acid code. In the patients of our study, mutations of the NHLRC2 gene indeed changed the code. Using cultured fibroblast from patients’ skin biopsies, we checked the expression of the NHLRC2 protein. Fibroblasts from healthy persons’ skin biopsies had a normal level of NHLRC2 protein whereas, owing to mutations, the protein had been almost completely eliminated in the FINCA patient derived cells. This would indicate that the mutations change the structure of the NHLRC2 protein so radically that the cell attempts to break up the harmful protein. This enabled us to ensure that these were not neutral variants.
As a result, the study discovered a gene that was necessary to normal embryonic development. The researchers drew the conclusion that the gene is vital during the first cell divisions in the mouse. Findings in FINCA patients also showed that the NHLRC2 protein plays a key role in maintaining a normal function of several organs in humans.

The study of rare diseases may also lead to discovering the causes of more common diseases, specifically proteins with previously unknown mechanisms.
“By studying a rare disease, there is a high probability that we will discover reaction paths that play a role also in more common diseases.”
Cell functions are based on biochemical reactions. Reaction paths are either turned on or off, depending on the function of the cell. Cells change their behaviour on the basis of messages they receive from their environment. For example, when a hormone attaches to a receptor on the cell surface, the receptor activates a certain molecule inside the cell, which will carry the signal further. The signal often ends up in the nucleus and controls the reading of genes . There are large numbers of signalling paths. Cancer cells, for example, do not react to many of the messages intended for them. Instead, they strengthen signalling paths that cause the cell to divide, and therefore help the tumour to grow.
The NHLRC2 protein has an effect on many cell reaction paths and events. For the first time, researchers were able to show that a dysfunction of NHLRC2 leads to changes in the cytoskeleton and the formation of vesicles within cells. The cytoskeleton plays a major role in multiple cell functions, and without it the cells would not be able to survive.
According to Hinttala, the study of rare diseases can result in discoveries that are important to basic cellular biology. It is also valuable to learn how dysfunction of genes and proteins manifests in humans.
“With FINCA, a dysfunction of the NHLRC2 protein will result in serious tissue fibrosis and degeneration of neurons. More common diseases, such as liver cirrhosis and Alzheimer’s, have similar tissue manifestations.”
According to Hinttala, the study of rare diseases involves a high probability of discovering the reaction paths that also lead to more common diseases.”
“Severe childhood diseases are rarely caused by environmental factors, but the role played by such factors complicates the study of the adult-onset disease mechanisms . The diseases we are studying are most probably hereditary and primarily caused by a dysfunction of a single gene.”
Ari Turunen
4.7.2019
Read article in PDF
Citation
Ari Turunen, Reetta Hinttala, & Tommi Nyrönen. (2019). Research on rare genetic disorders can be utilised in understanding the mechanisms behind even more common diseases. https://doi.org/10.5281/zenodo.8131030
More information:
Biocenter Oulu
Biocenter Oulu is part of Biocenter Finland, which coordinates infrastructural activities for major national research. It is also a member of various European research infrastructures. These are Infrafrontier (transgenic mice), Euro-BioImaging (biological imaging) and Instruct (protein structure research).
https://www.oulu.fi/biocenter/
CSC – IT Center for Science Ltd
CSC – IT Center for Science Ltd is a state-owned, non-profit public limited company run by the Ministry of Education and Culture. CSC maintains and develops the state-owned centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR is building an infrastructure to support research and the bio sector. It combines the leading organisations of 21 European countries and the European Molecular Biology Laboratory (EMBL) into a single infrastructure for biological information. Its Finnish centre is CSC – IT Center for Science Ltd.
https://www.elixir-finland.org
http://www.elixir-europe.org

Patient data is important for research. Personal data is protected by hiding or modifying identity attributes, while maintaining the level of statistical data significant for research purposes. This is enabled by a new, AI-based service.
VEIL.AI anonymises patient data better and faster than traditional methods, and retains information more effectively. If necessary, the application can also produce synthetic, fully anonymous statistical data, which cannot be traced back to any individual.
Developed by the Institute for Molecular Medicine Finland (FIMM), the application is now available for the ELIXIR infrastructure, alongside which a joint service is now being developed. An organisation managing data can protect it by entering the related metadata into a scalable cloud service. The service disguises any identity attributes, providing researchers with anonymised and, if necessary, synthetic data.

The VEIL.AI application employs a model based on artificial intelligence. The application creates a veil that protects the patient’s identity attributes, but can identify relevant data, which it keeps.
“Occasionally, for example when creating machine-learning models, more data is required–and more quickly–than research ethics committees tend to allow. They require justification for each variable, which is against the essence of machine-learning where maximum amount of variables is wanted without, a priori, assuming too much of their impact as far as the best mode is still being sought,” says commercialisation expert Tuomo Pentikäinen.
This is why, according to Pentikäinen, it makes sense to use synthetic data in the early stages of modelling, as this is what the VEIL.AI method can produce.
“This means data which is totally detached from the people it was derived from, but which, in terms of the desired variables, behaves like the original data. However, synthetic data is only one type of data we provide. Usually customers want anonymised data.”
VEIL.AI can find variables regarded as sensitive in terms of revealing a person’s identity, and anonymise them automatically.
“In a better and more organised way, the application can perform heavy calculations and operations concerning the calculation of data partitioning and anonymisation metrics.”
It must be possible to protect sensitive patient data, but many traditional anonymisation models also lose important data in the process. Traditionally patient data has been protected by partitioning and generalising identity attributes within the data. Anonymisation studies how variables divide/partition data into different groups. Then each group is examined separately, and if some variables are too easy to identify, they are coarsened. Generalisation means that a person’s age can be rounded off by a few years and a professional title can be changed from, say ‘nurse’ to ‘health care professional’.
“So any variables that are too easy to identify are generalised to a sufficiently general level, or perhaps even removed completely. When processing health data, deletion may have to be used quite often if a variable is unique or too easily identifiable,” says Pentikäinen.
This means that generalisation can result in the loss of important patient data.

“This tends to happen when an interesting phenomenon (such as an illness) is relatively rare and fairly evenly distributed across the data set. When such data is divided into partitions for anonymisation, it is common for the phenomenon of interest to be even rarer in each partition. In cases like this, traditional methods commonly interpret the interesting data as “outliers” in each partition, and therefore remove it. This is stupid, because with a better chosen strategy, the phenomenon of interest could have been included in the partitions, while retaining important information more effectively.”
Timo Miettinen, Information Systems Manager at the Institute for Molecular Medicine Finland, has an example: a patient with a rare type of breast cancer. Creating data that is too coarse can lead to the disappearance of data on the rare type of disease, because there are so few such patients in the data set.
“A breast cancer patient has one diagnosis, but her genetic profile indicates that she has a rare type of breast cancer. There may be a few cases like this per hospital, meaning that they may be classified as outliers and deleted. But this does not apply to the entire population. If a better view could have been gained of the big picture, this outlier would not have been deleted.”
Timo Miettinen has been involved for a long time in designing information systems that make use of and protect clinical data. Miettinen and his team have developed the VEIL.AI application, which is about to become commercially available. This micro service was created due to GDPR.
Each biobank in Finland has its own code register. The code register consists of personal identity codes and a synonym table, used to create an identifier–that is, a pseudonym–for each person.
“Certain things are difficult to change, such as height, eye colour and place of birth. They are identifiable through statistical methods. The same applies to a person’s medical history,” says Miettinen.
“We make two promises. First, we promise scalability and better performance. We are able to make use of continuously updated data from a number of sources. We can anonymise this effectively and securely. Our second promise is to try to minimise data loss. The application takes account of the data content, while fulfilling the anonymisation criteria,” says Miettinen.

The VEIL.AI application uses a neural network that can be thousands of times faster than conventional methods.
“Our system enables safer data distribution, because once the neural network has been taught what to do, each owner of confidential data can anonymise data before passing it onto its partners. Our method also produces better data, because we can test a huge number of different data partitioning strategies and pick the one that results in the smallest loss of information, while nevertheless achieving the desired level of anonymity,” says Pentikäinen.
And crucially, in terms of data security, the VEIL.AI application does not store patient data in a new location.
“We do not want to manage data. Instead, data is streamed through our service, during which we anonymise it and return it immediately to the customer,” says Tuomo Pentikäinen.
“We offer a scalable cloud service. Through the user interface, we can enter the necessary data dictionary and teach the algorithm to create the data anonymisation model using example material. The algorithm will learn to process the data, and if more data is added, it is streamed through the cloud service and anonymised,” says Timo Miettinen.
This means that organisations no longer have to share any of their sensitive data with anyone. Data arrives anonymised from the cloud service for research purposes.
The analysis of various pseudo identifiers requires plenty of processing power, which has been obtained from the ELIXIR infrastructure.
Ari Turunen
3.6.2019
Read article in PDF
Citation
Ari Turunen, Tuomo Pentikäinen, Timo Miettinen, & Tommi Nyrönen. (2019). VEIL.AI: patient data in a veil. https://doi.org/10.5281/zenodo.8119016
VEIL.AI
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Biocenter Oulu offers services for the study of proteins, cells and genes, and the generation creation of transgenic animals. One of its strengths is the light and electron microscope imaging of tissues and cells.
According to Professor Lauri Eklund, Coordinator in Light Microscopy at Biocenter Oulu, genetically modified model animals and mice in particular have helped researchers to understand more about phenomena related to the normal development and tissue operation of mammals than of any other organism. In addition to better understanding of developmental processes, these can be used as model organisms to study human diseases.
“Many imaging projects in Oulu concern the study of genetically modified mice. Mouse embryos and organs are imaged in Oulu either in whole, or by means of tissue specimens and light microscopy down to the accuracy level of individual cells, or even in higher resolution using electron microscope. We have also introduced methods to obtain images of cells and macromolecular structures in aestheticized mice, enabling us to see living tissues at high resolution. For this purpose, an intravital imaging laboratory has been set up, which enables cell examination in animals under anaesthesia. Small surgical procedures can also be performed in the laboratory.
“In addition, we use image data to create 3D models by means of optical sectioning. In addition to imaging of relatively large volumes, motorised microscopes can also be used to create tissue images of a large surface areausing tile scanning.”
The core facility service of Biocenter Oulu’s light microscopy specialises in what is known as mesoscopic imaging. Mesoscopic imaging helps to understand interaction between cells in a complex tissue environment, or even in entire organisms. Samples of mesoscopic scale are larger in volume and area than normal microscopy —ranging from a few millimetres to a couple of centimetres. These include mouse embryos, organoids resembling three-dimensional organs, and entire small model organisms, such as flies and fish emryos.
“In technical terms, mesoscopic imaging often requires a tissue culture environment suitable for microscopic imaging, specially designed 3D imaging equipment, tissue clarification methods, and advanced image analysis and processing capacity,” says Eklund.

Biocenter Oulu has a range of microscopes that enable various imaging methods and can pinpoint the location of various events in cells and tissues. A time dimension (4D imaging) can be added to three-dimensional images in living samples. Image sequences can be created, for example to show how cells differentiate and grow into, say, embryos or organoids.
“Thanks to the work done by Professor Seppo Vainio’s research group, we can grow an organoid within a few days, using various types of renal progenitor cells. This is something that has attracted international interest. Many researchers have come to Oulu to learn about the technique.”
Confocal and light sheet fluorescence microscopes are suitable for the imaging of three-dimensional and living samples. Especailly light sheet fluorescence microscope can scan the samples quickly without phototocix effect. Electron microscopY, on the other hand, can be used to find changes in the structures of cells and extracellular spaces, which are beyond the resolution of optical microscopy. However, this technology requires that the samples are fixed into place.
Although light waves cannot create magnifications in the same way as electron microscopes, innovative use of excitation laser light, fluorescently labelled molecules and image data processing, can achieve a level of resolution in optical microscopy which allows the examination of individual cells, cell organelles and macromolecular structures.
In most cases such objects must be made fluorescent in order to become visible in 3D microscopy. In the living cells or organism a fluorescent protein can beattached genetically, to the molecule to be studied. The fluorescent compounds (fluorophores) absorb energy from the excitation light and release part of this energy in the form of longer light wavelengths. This quantum mechanics phenomenon, which is visible to the human eye in certain range, is known as fluorescence.
It is also possible to search for specific proteins in cells and tissues by using antibodies to which a fluorescent marker has been attached. The antibody will identify its epitope in a certain protein and attach to it. Once it has attached, the marker can be detected with a microscope. The marker is chosen based on the kind of microscope that will be used to study the sample.
“We have microscopes equipped with spectral detectors and continuous laser light that enable us to study more than one fluorescent label simultaneously. This enables the study of complex interactions and multiple proteins.”
In fluorescence microscopy, fluorescent molecules are used as markers, while electron microscopy uses electron dense material, such as gold, for example.
”Oulu also has what is known as a label free imaging method that does not require additional markers or contrast agents, but can image endogenous molecules These include connective tissue collagen made visible with multiphoton technology, or other body’s own molecules, such as haemoglobin, which can berevealed through photoacoustic imaging. In the case of the latter technology, by combining excitation lasers, structural and functional information can be obtained from tissues, such as the structure of blood vessels and the blood’s oxygenation level.
“These technologies are very useful when imaging living tissues into which it is difficult to introduce markers.”
In terms of electron microscopy, Oulu specialises in the ultrastructural pathology in model organisms and immunoelectron microscopy of tissues and cultured cells,. These techniques provide information on the minutest details and the exact position of the proteins being studied in cell and tissue structures.
In immunoelectron microscopy, a metal labelled antibody or other reagent is related to the protein being studied, which means that the protein’s location can be determined very accurately, in a nanometer scale. This can provide new information on, for example, cell structures and protein interactions or orientation.
“The use of electron microscope methods to examine ultrastructures has been particularly fruitful in terms of the study of macromolecular structures of the extracellular matrix, which cannot be seen using optical microscopy. A fairly new area of study involves extracellular vesicles, “exomes”, which can be imaged by means of electron microscopy.

The problems with traditional imaging have been low resolution, low imaging depth, and lack of effective and quantitative analytics for the image data. In addition, special experience is required in order to extract biological data from the images.
Computer learning and machine vision methodsfor image interpretation has been developed in Oulu. In the approaches Biocenter Oulu has been collaborating with Professor Janne Heikkilä from the University of Oulu’s Center for Machine Vision and Signal Analysis.
“Data storage, transfer and analyses are challenging with respect to the 3D and 4D imaging of large samples. When data is transferred from microscope to user, one should be able to analyse it. Depending on the data, analyses may require a high amount of computing power. If the original data is stored somewhere distant, image processing may be slowed down due to insufficient data transfer speeds.”
Lauri Eklund regards the infrastructure provided by ELIXIR Finland’s CSC as the best solution, in national terms, for raw data storage and the reuse of open data.
Although metadata is linked to image data, there still can be many problems with data management.
“In order for image data to be reusable, it should conform to certain standards and be curated and annotated. Ideally, research infrastructures should provide image data “librarians” and “image information specialists”.
Ari Turunen
20.5. 2019
Read article in PDF.
Citation
Ari Turunen, Lauri Eklund, & Tommi Nyrönen. (2019). Biocenter Oulu: technology services for biomedical research. https://doi.org/10.5281/zenodo.8176718

In standard bright-field and fluorescence microscopy, light goes through the entire sample, and in doing so the light wave becomes dimmer and is diffracted in the tissues, causing the imaged object to become blurred, and resulting in poor imaging depth and resolutionin thick samples.
To overcome these shortcomings, a confocal microscope uses a narrow laser beam to scan a small part of an sample one depth layer at a time. Pinhole filters out emitted light that is not on the focal plane, thereby achieving greater resolution in samples that are too thick for traditional wide-field fluorescence microscopy.
A confocal microscope creates the final image from small aligned areas. Three-dimensional images are created by reconstructing two-dimensional optical sections from various depths of the sample. Three-dimensional modelling combines several optical layers to create visual structures that cannot be viewed with traditional optical microscopy.
“The multi-disciplinary approach of the University of Oulu has been used in the adoption of new technologies. In terms of light sheet fluorescence microscopy and photoacoustic microscopy, for example, we cooperated with Docents Matti Kinnunen and Teemu Myllyllä of the University of Oulu and the Optoelectronics and Measurement Techniques Laboratory before such technologies were commercially available. This gives researchers a competitive edge,” says Dr Eklund.

Light sheet fluorescence microscopy can be used to obtain microscopic images of light-sensitive samples or rapid biological processes within large living specimen. The sample is illuminated with excitation light one layer at a time, and the signal created by the sample is collected with another objective. The microscope has continuous optical sequencing: when the sample is moved on the light sheet, individual optical levels can be saved as 3D images. Large 3D samples can be scanned more quickly but with somewhat lower resolution than with a confocal microscope.
“Biocenter Oulu was the first laboratory in Finland to adopt this technology. The light sheet fluorescence microscope can take images of mesoscopic samples with clarified tissues, as well as living three-dimensional samples, whereby a time dimension is given for the images. This means that we can follow, for example, the genetically labelled cells during the growth of embryo or organoid within a specific time frame,” says Eklund.
Thanks to a new light sheet fluorescence microscope developed in Heidelberg, in 2015 researchers in EMBL were the first to observe how a fertilised egg developed into mouse embryo in a few days.
In 2018, the Howard Hughes Medical Institute in the United States introduced a microscope that utilises multiple angles of vision, enabling the imaging of an embryo’s growth at the level of individual cells. The researchers followed the embryos to see which genes were activated and which cells connected to each other.
Two light sheets illuminated the embryo, while two cameras recorded the early development of organs. The embryo’s location and size were tracked by algorithms. Algorithms observed how the light sheet was moving in the sample, deciding how to obtain the best images while ensuring that the embryo remained in focus. Because an embryo changes constantly, the microscope must continuously adapt andvery fast select from hundreds of images and time windows.
”The advanced mesoscopic methods of the future may be able to use non-diffractive excitation light (Bessel Beam and Airy Beam). Unlike ordinary light, this type of excitation light retains a uniform intensity in thick tissue samples. Moreover, the excitation light’s asymmetric form and ability to re-shape improve the imaging resolution in non-homogenous tissue samples with a high level of light diffraction.
By reshaping we mean that, unlike ordinary light, the excitation light can return to normal despite partially hidered by an obstacle.
According to Lauri Eklund, the rapid development of techniques for 3D imaging of living samples has led to an enormous increase in the amount of stored image data. Demand for quantitative analysis software for image data is also high.
“We can get the best out of these new imaging technologies if we also master image processing and analysis. In the case of mesoscopic imaging in particular, the large size of the samples requires effective image analytics and processing tools that can, for example, remove image noise and enable accurate 3D modelling. Smart computer software can also analyse cell behaviour and identify cell properties. In some applications you can, for instance, distinguish between cell types, determine cell division activity and analyse cells’ movement and viability.”
For more information:
Biocenter Oulu
Biocenter Oulu is part of Biocenter Finland, which coordinates infrastructural activities for major national research. It is also a member of various European research infrastructures. These are Infrafrontier (transgenic mice), Euro-BioImaging (biological imaging) and Instruct (protein structure research).
https://www.oulu.fi/biocenter/
CSC – IT Center for Science Ltd
CSC – IT Center for Science Ltd is a state-owned, non-profit public limited company run by the Ministry of Education and Culture. CSC maintains and develops the state-owned centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR is building an infrastructure to support research and the bio sector. It combines the leading organisations of 21 European countries and the European Molecular Biology Laboratory (EMBL) into a single infrastructure for biological information. Its Finnish centre is CSC – IT Center for Science Ltd.
https://www.elixir-finland.org
http://www.elixir-europe.org

Small differences in our genome distinguish us from each other. Although diversity is a good thing, many diseases are caused by genetic variants. Due to the around 80-percent similarity between the mouse and human genome, mice are of great assistance in the study of human diseases. Mouse strains can therefore be used to build models of genetic diseases that affect humans. Such models can be used to study the causal mechanisms of diseases and to design drugs.
The European INFRAFRONTIER research infrastructure offers over 6,800 mutant mouse strains, i.e. strains that model thousands of different diseases.The infrastructure has 23 member organisations. Finland is represented by Biocenter Oulu at the University of Oulu.
The Nobel Prize in Physiology or Medicine 2007 was jointly awarded to Mario R. Capecchi, Sir Martin J. Evans and Oliver Smithies for discoveries, which enable stem cells to be used to create germline modifications and additions into a mouse’s genome. Evans developed a technique for growing mouse-embryo stem cells. He implanted the stem cells into mouse embryos, which then developed into mice composed of cells with two distinct genotypes. In other words, stem cells could be used to include genetic material from one mouse to another. The hybrid mice develop normally at cellular level.
Capecchi and Smithies developed a procedure for silencing the expression of a certain mouse gene by transferring foreign DNA to a precisely targeted part of the mouse’s genome. The foreign DNA either knocks out or extensively decreases the expression of the gene, enabling scientists to identify which gene affects which mouse characteristic. Stem cells from a mouse embryo were used to create a mouse model with targeted silencing of genes. In this way, a hybrid mouse was created whose offspring were full ‘knockout’ mice, i.e. a gene function had been knocked out from their genome. This technique has now largely been replaced by the CRISPR-Cas9 method.
Being able to silence a single gene in a mouse enables studying the effect of switching off the gene in question. This has allowed the identification of genes that determine phenomena such as the development of mammalian bone structure and certain internal organs.

Genome editing has enabled the creation of mouse models of several human diseases. Such models can be used to study the causal mechanisms, progression and, of course, the treatment of diseases.
“If a new mutation or disease is found in a patient, we see only the clinically defined symptoms of the disease. If a disease has reached the late stage or has multiple symptoms, it is difficult to distinguish the primary cause from secondary effects,” says Adjunct Professor Reetta Hinttala from Biocenter Oulu.
Hinttala is the Coordinator of the Transgenic Core Facility at Biocenter Oulu. The facility is part of the European INFRAFRONTIER infrastructure. INFRAFRONTIER provides researchers around the world with access to mouse models for the study of genomes and diseases.
Within the Faculty of Medicine’s PEDEGO Research Unit at the University of Oulu, Reetta Hinttala uses mouse models to study rare inherited diseases. Mouse models help to identify the genes causing illnesses.
“Mouse models are a key element in gene research. They enable scientists to study disease genes and mechanisms at organism level. A model provides basic information on how a disease progresses. By studying mice of various ages, we can discover what happens in different tissues during each stage of a disease.”
Hinttala explains that it would be difficult to perform similar tissue-level analyses on samples from human patients, particularly for diseases of the central nervous system.
”An animal model provides valuable information on events at tissue level during the earliest stages of a disease. They may be of a kind that would go unnoticed in humans. Targeting research at early changes of this kind enables the discovery of early-stage treatments, which can be used in future pharmaceutical development.”
Hinttala points out that mouse models are particularly important when studying poorly characterized proteins and disease mechanisms. They enable the study of pathological changes at the tissue level during a disease’s progression in their proper environment. The similarity between the mouse and human genome allows observation of the same fundamental disease-causing mechanisms in mice and humans.
Tissue is composed of multiple cell types and their surrounding extracellular matrix. Researchers in Oulu are investigating histopathological structures of tissues and how they are organised to form organs. A microscope capable of imaging structural features of the tissue in detail is a key tool in the collection of data. New information is gathered on tissue samples by using various staining techniques to visualise chosen structures or molecules.
Digital images of tissue samples are tagged with metadata and archived. However, the storage and sharing of the imaging data is challenging. An image file of a tissue section scanned using a Slide Scanner can be several dozen gigabytes in size. Ensuring the future management of files of this size is challenging.
While researchers have an increasing need to store image material, they also need to be able to describe the data, to enable its distribution to the scientific community.
”Both INFRAFONTIER and the ELIXIR infrastructures are working to promote access to open research data. To make data derived from mice maximally available, it must be processed and analysed to become suitable for research purposes. I also think that describing imaging data in line with international standards – which enables the further use of images for purposes such as combining data with supplementary data sources – is an important line of work,” says Tommi Nyrönen of CSC, Head of Node of the Finnish ELIXIR centre.
Europe’s scientifically important mouse strains are kept in INFRAFRONTIER’s EMMA repository, one of the world’s leading mouse strain repositories. EMMA (The European Mutant Mouse Archive) archives genetically modified mouse strains from all over the world, free of charge. EMMA currently holds 6,800 mutated mouse strains, i.e. mouse models, many of which have been connected to diseases. The operations of Infrafrontier’s various nodes are organised via a centre in Munich. Located in 12 different countries, the nodes engage in the cryopreservation, archiving and distribution of mouse strains. Some of the nodes also phenotype mouse strains.
”Researchers can preserve their own mouse strains, or mouse models, in the repository, if the strain is sufficiently well characterised and a certain mutation has been reliably verified. When a new mouse strain is accepted for the EMMA repository, the researcher sends the mice to a selected node. INFRAFRONTIER’s website includes a search tool for exploring which mouse strains are currently archived. A cryopreserved strain can later be rederivated, resulting in a live mouse.”
A total of 226 mouse strains are cryopreserved at Finland’s EMMA unit in Oulu. They include a mouse model of the rare FINCA disease (discovered by Reetta Hinttala and Johanna Uusimaa, Professor of Paediatric Neurology). The NHLRC2 gene has been silenced in this model. The NHLRC2 protein was discovered to be vital to normal foetal development and several organ functions.

Phenotype classification is done on the basis of the mouse’s symptoms and other characteristics. Systematic analyses are used to determine the effects of a genetic modification on a mouse. Such analyses thereby describe the mouse model in question.
”Analyses of this kind are performed in mouse clinics, in which advanced analysis and diagnostics techniques are used to study genotype-phenotype interactions,” explains Hinttala.
Through Infrafrontier, researchers have the opportunity to use the services of the German Mouse Clinic and the worldwide International Mouse Phenotyping Consortium (IMPC). The IMPC generates phenotyping data from mice, in which one of the 20,000 or so genes has been knocked out, to aid studies on disease mechanisms.
Phenotyping is also needed at national level. Reetta Hinttala is the chair of the FinGMice platform, which is part of Biocenter Finland. Together, Biocenter Finland’s four locations in Helsinki, Turku, Kuopio and Oulu provide a comprehensive and diverse mouse phenotyping network.

”The aim is to guarantee researchers access to services, equipment and analysis support in both primary and secondary phenotyping. For this reason, it is very important to be able to transfer for example large image files of tissue sections between universities, so that we can consult experts all over Finland when needed.”
Ari Turunen
23.4.2019
Read article in PDF
Citation
Ari Turunen, Reetta Hintala, & Tommi Nyrönen. (2019). Mouse models provide insights into the causal mechanisms of diseases. https://doi.org/10.5281/zenodo.8118927
More information:
Biocenter Oulu
https://www.oulu.fi/biocenter/core-facilities
INFRAFRONTIER
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

An infrastructure has been created in Europe to enable European researchers and companies to utilise various organisations’ technologies and services related to bioimaging. Euro-BioImaging, headquartered in Turku, Finland, currently consists of 29 service units, called nodes, and involves 12 member countries and the European Molecular Biology Laboratory EMBL.
Researchers are offered an opportunity to utilise, for example, cell and tissue imaging technologies and, animal and preclinical imaging services. The services of the Euro-BioImaging infrastructure consists of the best selection of top-quality imaging centres in Europe. Users can access dozens of biological and medical imaging technologies. These include positron emission tomography (PET), magnetic resonance imaging (MRI) and various super resolution techniques based on light microscopy. More than 200 research instruments are available.
The infrastructure headquarters are in Turku. Turku BioImaging (TBI) shares administrative responsibility with EMBL’s Heidelberg unit and the University of Turin. EMBL focuses on biological imaging and Turin on medical imaging. TBI is responsible for the coordination of the entire infrastructure and the development of the Euro-BioImaging Web Portal. Through the portal, researchers can utilise technologies, data collections and educational opportunities within the infrastructure. The simple protype of the portal is operating and the actual portal will be launched during the year 2019.
“For the time being, Euro-BioImaging operates on the basis of voluntary resources, but an official infrastructure should be established, under a European Commission decision, in 2019. The infrastructure will be headquartered in Finland and take form of an European Research Infrastructure Consortium (ERIC),” says TBI’s Administrative Director Pasi Kankaanpää.

Euro-BioImaging offers not only research instruments but also image analysis tools and general services, such as image collections, related to image storage and data. The collections contain images by the terabyte, which can be used as reference data, for example.
“Our objective is that in future all published image data will be stored in the cloud and centralised services. We already have a fair amount of material, ranging from plankton imaging to cancer cells,” says Kankaanpää.
“Image analysis services and methods will also increasingly be used in various technologies and data collections. Researchers can, for example, look at how the images stored in the collections have been analysed previously.”
Image Data Resource (IDR) already shows the biological significance of the stored image sample. The data available on an image sample may, for example, reveal what aspects of the sample’s genes have been studied, and how.
According to Pasi Kankaanpää, IDR and other Euro-BioImaging data services are under active development.
“It is important to ensure that the services are compatible with the Euro-BioImaging Web Portal. How can researchers access the data and how has the data been stored and linked to various imaging technologies? This is where metadata comes in.”
Data collections and research instruments needed by researchers are available in a number of European research infrastructures. Access to such resources often requires an authorisation process. Federal user identity management is a simple and administratively agile solution. With a single password and their own organisation’s user ID, researchers will gain secure and reliable access to infrastructures.
The Euro-BioImaging portal uses the ELIXIR AAI service for identification and authorisation. In the future, its name may change to the Life Science Authentication and Authorisation Infrastructure service. This is the next step in the authentication service developed within the ELIXIR infrastructure. Infrastructure from various bioindustries, such as ELIXIR and Euro-BioImaging, are included in the EOSC Life project, which involves the implementation of this new and more comprehensive identification service. TBI and Finland’s ELIXIR have, for example, been working on the technical requirement specifications of Life Science AAI. TBI is also one of the pilot testers.
“Euro-BioImaging’s current data services are entirely public, requiring no permit process or login. This may change if we use central storage for other than public data and link such data to user applications created by users in our portal, for example.”
The Finnish service centre of the Euro-BioImaging infrastructure operates from three locations: Turku, Oulu and Helsinki. It has built high-quality and comprehensive light microscopy services. Finland’s ELIXIR has been developing cloud-based computing and user management for a long time.
According to Pasi Kankaanpää, these two infrastructures have many aspects in common, especially in terms of data management principles and authentication solutions concerning users.
“Cooperation between various infrastructures can also be possible when sharing educational material or developing other online solutions. I think we should also be able to develop solutions further, offering users services in cooperation. For example, a user could produce image data with Euro-BioImaging and process it further through ELIXIR.”
Data management and its further processing require more and more capacity. According to Pasi Kankaanpää, the current amount of data in the Finnish service centres alone can total around 100 terabytes a year. On the other hand, the forthcoming imaging equipment can produce similar data volumes in just a week.
“This means that we will probably need plenty of data processing, as well as the computing capacity offered by the ELIXIR infrastructure.”

Ari Turunen
4.3.2019
Read article in PDF-format
Citation
Ari Turunen, Pasi Kankaanpää, & Tommi Nyrönen. (2019). Euro-BioImaging: imaging infrastructure. https://doi.org/10.5281/zenodo.8118919
More information:
Turku BioImaging (TBI)
Euro-BioImaging
https://www.eurobioimaging-interim.eu
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.

Bioinformatics has been used to crack the human genome. New imaging techniques will now provide us with a direct view of how genes interact with each other and their environment.
Modern imaging methods can be used to obtain accurate structural images of the body. They can be used to diagnose diseases, and plan treatments and monitor their effects. Imaging has taken huge leaps ahead. Nowadays, we can examine and analyse even living cells to the accuracy of individual molecules.
Turku BioImaging (TBI), provided jointly by the University of Turku and Åbo Akademi University, offers top-quality imaging technology to researchers. TBI also trains researchers in the use of modern biomedical imaging techniques and develops international infrastructures in the field. Computer modelling and software development is needed for the processing and analysis of image data.
“Some of the methods were already invented in the 1950s, but technology enabling their use by researchers took a long time to arrive. Now, the situation is different. Use of these methods has simply snowballed. This has been made possible by better lasers and computers, and the discovery of certain self-illuminating molecules and super-resolution techniques. Thanks to these advanced methods and techniques, we can see things that used to be in the realm of science fiction,” says TBI’s Administrative Director Pasi Kankaanpää.
Various techniques are used in imaging. Turku offers at least the following imaging services: light microscopy, electron microscopy, atomic force microscopy, positron emission tomography (PET) and magnetic resonance imaging (MRI). Turku also offers facilities for analysing thousands of cells and their properties on the basis of flow cytometry. Plenty of open-source software is available for analysing any data obtained.
“Imaging is important for research. You can comfortably say that it is now one of the most important areas in biological and medical research of all kinds,” says Kankaanpää.
He refers to surveys carried out in Finland to determine key biomedical research methods and their use. Although bioinformatics has helped to determine many aspects of the human genome and other species, and biomedical research has made huge progress, Kankaanpää says that this is not enough. Research also requires imaging.
“Now we need to find out what genes do and how they interact with other genes and their environment. What would be a better way to do this than simply seeing what is going on.”
According to Kankaanpää, image material in itself is not enough to guarantee results. In recent years, a new field of science, known as bioimage informatics, has emerged. It means methods that are used to manage images and specifically analyse them quantitatively. The size of a single three-dimensional model can run up to several gigabytes. A huge amount of image data is processed to extract genuine information and to understand what is going on in the images. Analysis can also be automated through machine learning, for instance.
“Bioimage informatics has been forecast to have the same kind of revolutionary potential as gene technology had a few decades ago. Now we can use imaging to analyse the birth of illnesses and the operation of cells.”
TBI offers, for example, cell imaging services that utilise the best light microscopy available. Such equipment can be used to take images of individual molecules or small living creatures in their entirety.

Microscopy is based on the wave movement of light or electrons. As the name implies, the light source of electron microscopes is a ray of electron particles with which the sample is bombarded. An electron microscope has a much better resolution than a light microscope. The resolution is up to thousands of times better, capable of an accuracy of 0.2 nanometres. Although on electron microscope enables us to obtain images of structures and cell organelles, it cannot be used on living cells, because the sample preparation will destroy the sample in practice.
The Turku unit has hosted some major achievements in terms of microscopy. Stefan Hell, who used to work at the Biophysics Laboratory of the University of Turku, received the Nobel prize in chemistry in 2014 together with Eric Betzig and William Moerner for the development of extremely accurate light microscopy. He conducted the crucial experiments in Turku in 1993–1996.
Light waves cannot directly create as high a resolution as an electron microscope can, but you can get around this with clever use of lasers and fluorescent molecules. These methods make use of fluorescence, that is, a molecule’s ability to absorb light at certain wavelengths and to send back light at a higher wavelength.
The fluorescent protein, or fluorophore, is attached to the molecule being studied within the cell for instance by means of genetic engineering or antibodies. In a manner of speaking, the fluorophore is used to “dye” the object being studied.
By using fluorescent markers by, so to speak, switching the light “on” or “off” in various ways, the latest light microscopes can see structures that were once only visible to electron microscopes. One such method is stimulated emission depletion (STED) microscopy. This can reach an accuracy of just of few nanometres, that is, millionths of a millimetre. The wavelengths of visible light are several hundred nanometres.
STED microscopes can show the cell organelle structures and even individual molecules and their functions in tissue. STED microscopes can also create three-dimensional image data, and can be used for living samples.
“We have used advanced light microscopes to examine flu viruses to see how they invade a cell. The analysis software we have developed has enabled us to calculate the percentage of the viruses that has entered the host cell and how many have remained outside. We can also follow where the viruses go within a cell, how fast they move and when they break down.”
Imaging has been used as a model to develop nanoparticles that can deliver drugs with precision inside a cell, by imitating the operating mechanism of the viruses. In cancer treatment, for example, small particles have been inserted into a metastasis with a catheter to target radiation treatment at a tumour.
“We can now obtain 3D image data of living cancer cells and see how the particles move. In the same way as with flu viruses, we imaged how particles enter a cell and how they break down.”
The objective is to obtain a silver bullet that does not affect healthy cells.
“Our goal is to target the drug directly at the cancer we want to kill and no other cells. This would reduce the side effects of cancer drugs. Imaging enables this type of development work. It is extremely difficult to imagine how this type of work could be done without modern imaging,” says Pasi Kankaanpää.
Ari Turunen
28.2.2019
Read article in PDF
Citation
Ari Turunen, Pasi Kankaanpää, & Tommi Nyrönen. (2019). Imaging helps to highlight significance of data. https://doi.org/10.5281/zenodo.8118822
More information:
Turku BioImaging
Euro BioImaging
https://www.eurobioimaging-interim.eu
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

A lot of genetic data and clinical data is easily generated from measurement instruments. It is important, however, to decide in good time how and in which format the raw data is stored and how the postprocessed data is classified and described, the measurement event included.
It is important to determine metadata, i.e. the information that describes the data, by the exact same means in all research institutions and laboratories around the world. Otherwise, we cannot obtain the maximum benefit of the data for research, as it cannot be linked to data produced elsewhere.
“Even within our own research group it can become chaotic if, for example, the same files have not been used, meaning that the data is not comparable”, says professor Aarno Palotie at the Institute for Molecular Medicine Finland (FIMM), University of Helsinki.
Owing to international cooperation, some standards have already been created. The VCF (Variant Call Format) format determines the text file used in bioinformatics when storing genetic sequence variation data. BAM (Binary Alignment/Map) is a format that can be converted to a readable text format.
Founded in 2013, GA4GH (Global Alliance for Genomics and Health) is an international alliance of more than 500 organisations representing the bioinformatics, health care and IT industries with the aim to create standards for data that is distributed for research purposes. In November 2017, ELIXIR and GA4GH decided to launch cooperation. The agreement gives the ELIXIR infrastructure an opportunity to influence the creation of international standards. The agreement is related to the GA4GH Connect project, the purpose of which is to introduce data standards in clinical patient work by 2022.
Aarno Palotie finds cooperation between ELIXIR centres and GA4GH important because, in addition to standards, it allows more than 1,000 organisations to work together in creating common principles for data processing and distribution.
“The ELIXIR centres have extensive networks in their respective countries and can influence the local practices.”
There is still work to do to achieve uniform data processing, analysis and principles of use. In Finland, the aim is to enable exploitation of genome data in patient health care. The purpose is to create a national genome data resource which will be maintained by Genome Centre Finland.
“In Finland, the aim is to create progressive laws for the linking and utilisation of clinical and genome data of the population.”
According to Palotie, the various uses of data must, however, be considered in the linking of clinical data and genome data used in research.
“No errors are allowed in patient-oriented analysis. Messing up with samples cannot be tolerated. The data must be available in identical form and easily accessible if it is used for clinical decision-making. Scientific genome data must, in turn, be flexible, quickly accessible and available in various formats. Research will advance only by flexible means.”
The researchers involved in the FinnGen project have had to deal with many kinds of agreements. The data protection regulations require extremely stringent protocols agreed upon in advance which, according to Palotie, is in contradiction with the research ideology.
“Basic research just fails to progress if it follows protocols determined from the outside. In research, the processes are modified and applied according to how the data is produced. The aims are different than when utilising clinical genome data.”
As the requirements vary, Palotie says that parallel routes are needed for the exploitation of these two different types of data. Legislation should further specify how genome data created for clinical purposes can also be utilised in research.
“A consensus should be reached also on how the data generated in research could, in some situations, be sensibly used for clinical decision-making. The situation is unclear at the moment in the Finnish Biobank Act.”
Naturally, it is important to take care of the necessary data protection, but excessive or tangled regulation of data use is troublesome from research point of view. The European regulatory environment is partially broken. Palotie mentions the mining of metadata as an example.
“A researcher wants to know, for example, how many individuals there are in the Finnish biobanks with a certain genotype, illness and age. Ideally, we would have a portal, which would give this piece of information in real time. A researcher cannot access the individual data which the computer is processing in the depths of the system. When utilising personal data, which is subject to stringent protocols in the EU, the starting point is that the data must not be processed. Research makes an exception here, but when strictly interpreted, it may even require a separate permit process each time.”
Utilisation of data is challenging because the regulatory environment has mainly been interpreted from the viewpoint of data protection, and not from that of the health benefit to an individual.
“The regulatory environment should be developed in such a way that all authorities interpret the legislation in the same way, which would also allow using the portal I have proposed. The use of this kind of portal, if constructed in an appropriate manner, does not pose a threat to data protection by any means whatsoever”, says Palotie and points out that the metadata regulations are looser, for example, in the United States than in Europe.
“The European permit processes have been characterised as a bureaucratic farce. It may take years to receive a permit”, Palotie sighs.
Palotie wishes that the new legislation created in Finland that relates to using genome data and secondary use of register data would accelerate the permit processes and clarify the regulations concerning the use of data.
“The data policy needs to be clear and the data available to everyone. At the moment, various authorities interpret the law in different ways, which is the weakest point from the viewpoint of a researcher’s legal protection. Hopefully the new laws will remedy this situation. I also hope that the updating of the Finnish Biobank Act due to the General Data Protection Regulation of the EU (GDPR) is aligned with other new legislation.”
Ari Turunen
14.1.2019
Read article in PDF
Citation
Ari Turunen, Aarno Palotie, & Tommi Nyrönen. (2019). Data harmony and standards: data must be processed, described and stored by uniform means. https://doi.org/10.5281/zenodo.8118815
More information:
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Professor Aarno Palotie from the Institute for Molecular Medicine Finland (FIMM) focuses on genetic analysis of diseases by utilising large quantities of data gathered from subjects. Together with his research team, he has been able to use data analysis to demonstrate that the underlying causes of various neurological diseases consist of numerous genes, instead of a single genetic mutation. For example, there are hundreds of different genes that affect a person’s predisposition to migraine, epilepsy, Parkinson’s disease or Alzheimer’s disease.
Aarno Palotie’s research requires an enormous number of samples. In 1998, while working as a professor at The University of California, Los Angeles (UCLA), he had access to the most extensive research data on migraines, available at the time: the data had been collected from 400 Finnish families suffering from migraines. Over the years, the research data has grown and now covers 1,600 families. Between 2007 and 2013, he carried out research related to migraines, schizophrenia and epilepsy in Cambridge, UK, using extensive research material.
Clinical and research data collected on people is constantly being produced and recorded. The more data there is available for research use, the easier it is to find statistical variables. Extensive amounts of data have the potential of revealing new information if the data is mined and analysed well.
“Studies should be able to utilise sample sizes, that are no longer measured in thousands but in millions,” Aarno Palotie notes.
“The large sample sizes have been collected from the diverse donor base of different biobanks. Data from different sources is combined in order to really increase the numbers. This way we can increase the signal and reduce the noise.”
By increasing the signal, Palotie refers to making the data statistically significant. For example, mining rare diseases from the research material requires large amounts of data.
It is valuable for researchers if the data has been collected using the same methods. Different laboratories and research facilities may have different practices for collecting, processing and classifying raw data from measuring instruments. The more consistent the data is, the easier it is to analyse.
“However, in real life, absolute harmonisations in disease research is very challenging to achieve. That is why it is important to harmonise the aspects that can be harmonised, in order to make real discoveries and correct interpretations using vast quantities of data.”
Genome data has conventionally been collected using sequencing techniques that are used to investigate the base sequence of genes in test tubes. However, sequencing costs are very high if a lot of data is required for the study, such as research into common or chronic diseases. Genotyping has become established as a cost efficient and reliable method. In genotyping, a DNA microarray technique is used to collect genetic data from DNA samples. The samples are studied with a microarray scanner and the collected raw data is then processed. Only the sections of chromosomes known to feature genetic variants related to the studied disease are studied in genotyping. After collecting the data, computational methods are put into use. A reference genome (a reference assembly created using DNA sequences from various donors) can be used to predict the variants that were not examined earlier.
The genetic variants that are studied in genome wide association studies (GWAS) are measured from sample sizes varying from hundreds of thousands to millions of samples. The GWAS method is most commonly used when the genetic background of a disease is multifactorial, or polygenic, meaning that hundreds or thousands of genetic variants affect the disease risk. Multifactorial diseases include cardiovascular diseases, allergies, diabetes and mental disorders, for example. An extensive amount of research data is required for a reliable GWAS analysis. The computing power of super computers is needed to analyse the data.
“Sequencing requires even more extensive amounts of data than GWAS techniques do. Sequencing is also expensive compared to the GWAS method. Producing data using the GWAS methods costs a few dozen euros. The method can be used with extensive enough sample materials. Data is standardised, and it preserves well. Data genotyped at different locations can be easily combined.”

Migraine is a disorder that causes headaches and is usually considered to stem from a disturbance in the brainstem caused by external factors. One in ten adults suffer from migraines and it is three times more common in females than males. Palotie has studied migraine for a long time. One study utilised samples collected from 375,000 people worldwide. 60,000 of them suffered from migraines. In 2016, his research team identified 30 new hereditary risk factors related to migraine. Many of them are located in genes that regulate vascular function.
In 2018, an article that provided new information on the causes of migraine, written by Palotie and other researchers, was published in the Neuron scientific journal. A significant observation was that even in migraine families migraine is not only affected by certain genes, but a vast number of genes. Palotie talks about gene load.
“For decades, the genetics of diseases have been thought of the way Mendel described them. It is actually far more complex,” Palotie says.
Gregor Mendel, who has been called the father of genetics, demonstrated that certain characteristics of a person are inherited by the succeeding generation. Genes can be dominant or recessive. According to Palotie, new research results have shown that it is not that simple. For example, a disease may be affected by a group of genetic mutations, not necessarily only one genetic variant.
“The assumption has been that if there is migraine, heart attacks, cancer or another common condition in the family, the genetic variants that cause it are strong and transferred to children from their parents. The migraine study together with other research actually show that the cause of certain diseases is likely to be an accumulation of very common genetic variants. These are some of the same variants that can be found in the entire population. Sometimes however, a person and their spouse simply happen to both have a heavy genetic variant load. When these two gene loads, which contain thousands of genetic variants, come together the risk of disease in their offspring increases.
Palotie and his team also search for similar gene loads related to other neurological diseases. Palotie is currently working on an extensive international study on the genetic background of psychotic disorders. Research material for the study is being collected from over 100,000 people worldwide. Genetic findings are believed to play a significant role in understanding diseases and forming a basis for developing new treatments.
“If there is a sufficient amount of data available on patients it is possible to provide more specific treatment, this is referred to as targeted treatment or more individualised treatment.”

Palotie conducts his research from two different perspectives: he looks for both accumulations of common genetic mutations and rare genetic variants.
“Rare variants may provide a short cut to biology,” he says.
As an example, Palotie mentions a patient with schizophrenia who had a few genes with a strong mutation connected to the disorder. This is very rare with schizophrenia, because predisposition to the disorder is usually the result of a combined effect of thousands of genes.
“However, such a rare occurrence may reveal something about the mechanisms of the disorder. It may be easier to identify a cell’s biological signaling pathway by studying rare mutations in the genes of patients with schizophrenia than it is by studying accumulations of common variants.”
Studying the signaling pathway, or understanding how a cell reacts to communication, plays a key role in understanding disease mechanisms. Cells react to messages they get from their surrounding environment. Often the signal goes all the way to the nucleus and starts to regulate what the gene does. Sometimes cells contain special proteins that are there to stop the signal. For example, cancer cells do not react to many signals intended for them. Instead, cancer cells enhance the signaling pathway, which makes the cell divide and grow.
Finns have a number of genetic variants that are rare in other parts of the world but rich in Finland because of our demographic history. When data collected from Finnish people is combined with data from other populations, we can get more information on signaling pathways. This means that a Japanese patient may benefit from data collected from Finnish patients and vice versa.
“Even if a variant identified in Finland is unknown in Japan the physiology and biology of people is still very similar. Hopefully, the identified variant helps steer toward the correct cell signaling pathway. When we identify a new cell signaling pathway another genetic variant, that is in-fact connected to the same signaling pathway, may be discovered from another population base. In such case, the finding helps confirm that the signaling pathway is significant in relation to this disease.”
The location of the genetic variant matters. Palotie has an example of this. When visiting Iceland, Palotie talked about Leif Groop’s study, as a result of which a rare genetic variant, that protects from type II diabetes, was discovered in the population on the west coast of Finland. Leif Groop’s colleague asked Icelandic researchers whether a similar variant had been identified in the Icelandic population. The Icelandic researches checked their databases. The same genetic variant had not been found but there was another variant of the same gene.
“This Icelandic discovery confirmed that the gene in question protects against type 2 diabetes. Such a protective genetic variant is obviously very interesting from the perspective of molecular drug design.”

The FinnGen project was launched in autumn 2017. The aim of the project is to record the genomes of 0.5 million Finnish people. The project utilises samples collected by all Finnish biobanks. The data collected on Finnish heritage will be combined with clinical data from national healthcare registers. The goal is to gain a better understanding of diseases by combining genome and healthcare data. Patient healthcare can only be significantly improved by analysing large quantities of samples.
FinnGen is centred around Finnish phenotype data collected from healthcare registers. In Palotie’s opinion, FinnGen can be considered a prime example of how data from biobanks and healthcare registers can be combined for genome data analysis.
The project has partners from all over world. The goal is to combine data from Finnish biobank samples with biobank data from other countries and carry out a meta-analysis.
“Combining data from the different sources is a great challenge and meta-analysis is often a more functional solution. FinnGen meta-analysis has been carried out with biobanks located in Great Britain and Japan. The aim is also to get other countries involved.”
According to Palotie, it is crucial to be able to combine the research data collected on Finnish people with data from other countries and populations. New methods for processing and cultivating data are constantly being developed. New methods are useless if researchers do not have access to enough data that can be analysed and inspected.
“Even artificial intelligence has difficulty functioning without enough data.”
Ari Turunen
10.12.2018
Read article in PDF
Citation
Ari Turunen, Aarno Palotie, & Tommi Nyrönen. (2018). Hundreds of genes could lie behind a single disease. https://doi.org/10.5281/zenodo.8118783
Read also:
Help from the Finnish genome for the prevention of cardiovascular diseases
Massive data management project: Finns’ heredity is collected and safeguarded
More information:
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Professor Samuli Ripatti’s group at the Institute for Molecular Medicine Finland and the Faculty of Medicine of the University of Helsinki studies the underlying mechanisms of cardiovascular diseases through genetic variation. The genetic heritage of the Finnish population provides a good opportunity for this.
In Finland, there are about 40 hereditary diseases that are characteristic of the Finnish population. The illnesses included in the disease heritage are caused by certain mutations that are more common in Finland than elsewhere in the world due to the so-called bottleneck effect. In the last 10,000 years, a relatively small number of settlers have moved to the Finnish territory. The individuals of this new population represented small and narrow genetic material, resulting in the regional enrichment of certain disease gene variants.
The bottleneck effect is beneficial today for determining the genetic heritage of diseases.
“Finland is the largest bottleneck population on the European scale. Few people have moved here over millennia. The genetic variants that have come to Finland with these immigrants can be a hundred times more common in Finland than elsewhere. This is not often the case in more admixed populations. In Finland, permanent settlements were first established in closer to the coastal areas and only much later in the east and in the north”, says Samuli Ripatti.
According to Ripatti, the internal migration that took place in the 16th century was a second bottleneck thanks to which large differences are discernible in the Finnish population between the east and the west. Although population isolates also exist elsewhere, defining the genetic background of the Finnish disease heritage has also inspired investigations of other diseases.
“We are now able to understand the onset principles of many genetically inherited diseases, allowing the research data to be applied elsewhere. Although the diseases are different, they have genetic mechanisms that operate in the same way. Understanding this dynamic is a big thing and may provide new opportunities for developing new treatments.”
For example, when studying Parkinson’s disease, it is possible to determine whether the disease is more common in some parts of Finland. If this is the case, research into these parts of the country may produce new genetic information.
Ripatti is interested in population genetics and genetic variation in Finns. He has been a Professor of Biometry at the Faculty of Medicine of the University of Helsinki since 2013. Biometry is a field of statistics that focuses on the analysis of biological data. Ripatti’s research group combines statistical methods with sequence-level measurements of the human genome.
“Sequencing provides information on genetic variation, which may be rare in a population. Based on variation, it is possible to see the prevalence of certain genetic changes that modify disease risk in some areas of Finland or demonstrate predisposition for a certain illness.”
This allows the screening of valuable information about the health effects from the population’s genetic data. People at a high risk of getting sick can be found while also looking for ways to prevent diseases.
“We look at this from the point of view of diseases which touch many individuals. We study illnesses that are common in Finland, such as cardiovascular diseases and diabetes.”
Even though these diseases are affected by, for example, diet and other lifestyle choices, hereditary factors are also significant. That is why these conditions are referred to as common complex diseases. Thanks to the Finnish bottleneck effect, Ripatti’s group has identified genetic variants, genes that predispose you to cardiovascular diseases, in particular, and predictive and marker-controlling genes measured in blood. Ripatti uses high cholesterol levels as an example.
“Those with high cholesterol levels can be examined and their genomes sequenced in an efficient and easy manner.”
Cardiovascular diseases are the cause of one in three deaths in the world. The most affected areas are Central Asia and Eastern Europe. In the 1960s, Finland was the world leader in coronary artery disease mortality of middle-aged men. Upon entering the 2000s, men’s mortality had fallen to about one fifth of the highest level.
However, regional disparities in cardiovascular disease morbidity and mortality are high in Finland. The incidence of the diseases is much lower in Western and Southern Finland than elsewhere in the country. This large regional difference is of interest to researchers. So-called silenced genes that protect against diabetes and cardiovascular diseases have been found in the population of Western Finland.
One interesting area of study is a gene being turned or becoming inoperative. Such genes are referred to as silenced genes. A study led by Professor Aarno Palotie analysed over 80 mutations silencing an entire gene that are rare but more common in Finland than elsewhere in the world. The material was obtained from the genomes of more than 30,000 Finns.
In fact, Finns have more genes that silence the function of an individual gene than other populations.
“Gene variants that disrupt protein production are generally quite rare in human populations. However, the gene variants that disrupt protein production brought to Finland with settlers are more common here than in the rest of Europe and that is why studying the health effects of those variants that arrived here at that time is much easier in Finland than elsewhere.”
Gene knockouts whose inoperability does not cause health problems have been found in the population of Western Finland. On the contrary, they protect their carrier against diabetes or cardiovascular diseases.
“A gene variant that protects against diabetes has been found in Finland. Carriers of the variant have less incidence of diabetes compared to others. There are more carriers of the gene variant in Ostrobothnia than elsewhere in the world. This may benefit the pharmaceutical industry if it is possible to mimic the function of such a gene through molecular preparations.”
Another example is a gene that prevents the function of lipoprotein(a). Heart disease risk can be assessed by measuring lipoprotein(a) in the blood. Lipoprotein(a) or LPA is a member of the lipoprotein family that carries LDL cholesterol. The data contained, for example, genetic variants whose carriers lacked lipoprotein(a) produced by the LPA gene almost completely. People lacking lipoprotein(a) develop cardiovascular diseases less frequently than others.
“There are a few variants of the LPA gene that shut off its function. In such cases, there is less lipoprotein(a) in the blood. This results in less vascular disease. Lowering the protein level by pharmacological means would be possible and this could be beneficial in preventing coronary artery disease.”
The function of the USF-1 gene has also been studied in Finland. In humans, the gene affects blood fat levels and cholesterol. When the function of the gene was knocked out in mice, the level of the good HDL cholesterol in the blood increased.

Thanks to the SISu (Sequencing Initiative Suomi) project, the data on genetic variation in Finns has been compiled into one database.
“The sequenced sample material has been collected from Finnish patients and volunteers. The material has been used to calculate statistics on how prevalent each genetic variant is in Finland. Once a large enough database has been compiled, it is possible to determine what Finnish genomic variation is like in general.”
The SISu database currently has the protein-encoding variants of the genomes of some 10,000 Finns, and even the full genome of several thousand Finns has already been sequenced.
“The sequence data of the SISu database gives us the opportunity to supplement our material measured with more favourable genome microarrays than others with statistical imputation algorithms. We can now tell quite accurately what kinds of gene variants Finns have. For example, if one in a thousand carries a specific gene variant in Finland, it means that at least 20 people on average should have the variant in the current database.”
The database is already helping patient diagnostics.
“The variation data is in the database and the data is utilised all the time, especially in clinical genetics. The starting point is that the database provides further clarification for the treatment of a hospital patient. Therefore, if it is suspected that a variant in a gene may be the cause of a disease, the doctor will check the database to see how often this variant occurs in Finns. If it is common, it is unlikely that it would be the cause of a rare disease. If it is rare and its effect on gene function is significant, then the likelihood of the variant’s significance in the onset of a disease is also increased. This is a very concrete clinical application of the database.”

The SISu project data has been collected from research projects and patients. However, the project has focused solely on genomic data, meaning that the potential for utilising the data in health research is limited.
“A biobank sample should be collected from all of us”, says Ripatti. “Those with a predisposition for illness should be screened more closely.”
In Samuli Ripatti’s view, it is a great shortcoming that genomic data is not yet available in connection with medical examinations. It should be part of everyone’s routine check-up to allow concrete decisions for treatment.
“Finland would have excellent setting for this. We have well-functioning occupational and basic health care as well as good expertise in genetic research.”
As a continuation of SISu, the FinnGen project that will record the genomes of half a million Finns launched in August 2017. The project utilises samples collected by all Finnish biobanks. The data from genomes is combined with the information in national health care registers.
“Tools for risk assessments exist and statistical models have been developed for several diseases. Interpreting the data obtained from the genome as part of routine health care is the goal of the next few years. FinnGen contributes to this.”
Ripatti and his team participate in the development, implementation and testing of statistical algorithms.
“We develop prediction algorithms that evaluate, for example, the risk of cardiovascular diseases in a patient. We combine genomic data and lifestyle factors, based on which a prediction is made. Therefore, we are looking for ways to motivate the patients to change their lifestyles.”
Ripatti’s group also supplements genomic data with statistical algorithms. Due to Finland’s population history, it is possible to predict the missing genotypes in microarray data better and more precisely here than almost anywhere else in the world. The algorithm works well in Finnish data because Finnish genomes are on average more alike than elsewhere. Sequence data is used to computationally supplement data collected using gene microarrays.
“If a gene microarray scanning the essential variation points has 500,000 genetic markers and we know 30 million genome variants of gene sequences in addition to these measurements, we can supplement the measurement done using the gene microarray into a full genome sequence with good statistical quality indicators. This allows the creation of more sufficiently reliable full genomes at a lower cost. For the time being, sequencing the entire genome is considerably more expensive.”
Preserving genomic data in the future and designing its analysis environment are big issues in which the ELIXIR infrastructure plays a key role.
“We have an existing pool of data that can be used by authorised researchers. With large databases, it is necessary to provide researchers with a secure and efficient data analysis environment where the data can be analysed.”
Due to data protection, the best solution would be, for example, a remote desktop. At present, there are hundreds of copies of the population data of different countries on the disk servers of various research groups around the world. It is an enormous amount of data.
“On the other hand, in the future, we must have solutions suitable for analysing genomic data that enable the efficient and decentralised storage and analysis of enormous data sets. This challenge will not be resolved by the closed remote desktop solutions developed previously for much more modest data volumes. Instead, what is required are open computing environments that efficiently utilise cloud services and international cooperation. Considering this is quite essential.”
Ari Turunen
5.11.2018
Read the article in PDF
Citation
Ari Turunen, Samuli Ripatti, & Tommi Nyrönen. (2018). Help from the Finnish genome for the prevention of cardiovascular diseases. https://doi.org/10.5281/zenodo.8118771
Read also:
Massive data management project: Finns’ heredity is collected and safeguarded
More information:
Institute for Molecular Medicine Finland (FIMM)
The mission of the Institute is to advance new fundamental understanding of the molecular, cellular and etiological basis of human diseases. This understanding will lead to improved means of diagnostics and the treatment and prevention of common health problems. Finnish clinical and epidemiological study materials will be used in the research.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Computational methods can now be used to deduce from data sets as to who is at risk of developing, for example, diabetes or cancer. Laura Elo and her research group develop methods which are used to find different predictive markers for diseases. Combining clinical data with molecular data can also provide valuable information about suitable drug treatment.
Research conducted on human biology produces a lot of new data for researchers to study. DNA sequencing generates an individual’s genetic profile. RNA sequencing, in turn, provides measurement data on the activity of genes. It tells which genes are expressed and possibly produce proteins in the cells at any given time.
Thousands of different molecules and their interactions can be measured from a tissue sample. For example, it is possible to study different active forms, or transcripts, of a gene. When the goal is to determine the function of proteins or their deviations in connection with diseases, it is called proteomics. Mass spectrometers are used as aids to measure molecular mass.
Laura Elo, Research Director of Bioinformatics at the Turku Centre for Biotechnology, and her research group develop modelling methods that allow the measurement data collected in follow-up studies to be utilised to determine disease risk on an individual basis.
“I started my career as a mathematician at a time when bioinformatics was still a marginal field. I became excited about biology and medicine back then”, Elo says.

One important material for the researchers is the data collected from different populations. The studies use data stored in Auria Biobank in the Turku region as well as data obtained from elsewhere in Finland and from other countries. The electronic medical records also have a lot of data collected from patient care that can be used in research, subject to consent. However, the data alone is not enough to determine the emergence and development of diseases. Computational methods and models are required to make comprehensible interpretations from data masses. The aim is to develop functional models for use by doctors.
“Almost all of our research is related to medicine and the needs of doctors. One of our major goals is to provide practical tools for doctors. The data alone is not useful, unless it can be modelled and interpreted. In the future, our work will hopefully allow patients to be offered treatments that are increasingly individually targeted.”
Effective treatment is always personal because drugs and treatment methods work in different ways for different individuals. A patient’s treatment response is affected by a number of factors, the information about which is obtained, for example, through laboratory measurements. In addition to clinical variables related to the patient’s health, there are many factors at the gene and protein level, which affect the efficacy of treatment methods. Mathematics helps with the analysis of data obtained about an individual.

“Biology is complex. One disease can actually present itself in many different ways at the molecular level, and different treatments can be effective for different people. A specific drug can cause serious adverse effects for some while being ineffective for others. Computational methods allow us to predict, who will suffer from the adverse effects and who are likely to benefit from the treatment. We mathematicians can help medical scientists to identify the key predictive factors”, Elo says.
Development of mathematical models requires large volumes of data as their raw material. For example, some of the predictive models have been developed using clinical patient data from the US, but they are also suitable for the patient data of the Turku University Hospital.
“When a sufficiently large amount of genomic and clinical data is obtained, they can be combined and the modelling phase can start. This is only possible if the description of the data, metadata, is in order.”
Many things have to be taken into account in the development of models. It is important to assess the prediction ability of the model in advance. Models easily become overfitted for the data that is used to create them. This means that the model is too well-suited for the data. Therefore, the predictive model works with one data, but the prediction is no longer good with new data. Validation is required to verify the model. This can be achieved, for example, by using patient cohorts from another hospital or country. Checking the model by using other patient data is important to allow for the general adoption of the model. Data from different biobanks helps with this.
“If the model is built and tested using the same data, you may get it to work almost perfectly in that data. However, it may not work on new individuals. Therefore, we strive to build models which predict the outcome as closely as possible but can still be generalised to new data.”
The work of Laura Elo and her research group with modelling involves continuous experimentation and change.
“After developing a model and showing that it works with certain data, the validation process is continued. We aim to find as many new data sets as possible to test the accuracy of the predictions produced by the model. You can always develop a model that works in one data. However, it is only after it has been verified in several data sets that the predictive model can be considered reliable enough to be given to doctors to support decision-making. The more widely the model can be tested, the better we can assess whether it only works for a specific population or if it is more universally applicable.”
New factors are added to models and their effect on predictions is analysed. For example, linear, simplifying models are easy to understand and interpret in hospitals. However, sometimes the interactions between molecules are so complex that linear models do not work and, therefore, other solutions are needed.
“The more new variables are added to a model, the more critical its validation becomes. An important question is understanding which variables are most significant for prediction and how their combinations provide the best predictions. You need to find balance for the model: it must be complex enough for prediction, but the model must not be overfitted to the data.”

Laura Elo and her team have been involved in the development of predictive models for renal cell carcinoma. Renal cell carcinoma originates in the epithelial cells of the renal cortex. The prognosis of renal cell carcinoma is poor as 40% of patients die within five years.
A new computational method can be used to find predictive markers from patient samples. The study found that the expression of 152 genes can predict the life expectancy of patients with renal cell carcinoma after surgery.
“The prognosis of renal cell carcinoma is usually good if the cancer is localized. On average, however, 50% of patients develop metastases after surgery. The goal is to predict as early as possible whether the patient’s prognosis is good or bad in order to select the best treatment strategy.”
Two different sets of data were utilised in the development of the predictive model. The gene expression data of more than 400 renal cell carcinoma patients were obtained from the international Cancer Genome Atlas (TCGA) database. The model was then validated using an independent Japanese data set of 100 patients.

Laura Elo studies patient data to search for different biomarkers that can predict disease onset or treatment responses. A biomarker is a factor or characteristic that indicates a change in biological status, for example, in genes or proteins. In Finland, researchers have aimed at determining the underlying mechanisms of type 1 diabetes for a long time. Type 1 diabetes is caused by the destruction of insulin-producing cells. The pancreas does not produce the insulin hormone needed by the body, thereby causing blood sugar to rise.
“Finland has the highest incidence of type 1 diabetes in the world relative to the size of the population. Both genetic and environmental factors play a role in the development of the disease. We look for biomarkers that could predict the development of the disease as early as possible.”
Because Finland has the highest levels of type 1 diabetes relative to the population in the world, diabetes research here is also significant. As early as in 1994, the ambitious and extensive research project DIPP (Diabetes Prediction and Prevention) was started in Finland. Genes that predispose you to type 1 diabetes are being sought in the blood samples of newborns. Children who are found to have a genetic risk to develop diabetes are invited to a follow-up study. Samples are taken every three months and, from age 2 onwards, every six or twelve months. The screening participants include the university hospitals in Turku, Tampere and Oulu.
“The children with a genetic risk of developing type 1 diabetes have been monitored until the age of 15. The goal is to identify the factors affecting the onset of the disease at the cellular level even before it can be diagnosed with the current methods.”
Laura Elo collaborates with Professor Riitta Lahesmaa, whose research group studies leucocytes and aims to understand what factors make cells cause diabetes. In the future, this could lead to preventing the onset of diabetes and curing current patients.

Going forward, Laura Elo wants to focus on the underlying mechanisms of diseases and the risk factors for falling ill. The statistical modelling of the complex interactions between different factors requires many new methods and measurement technologies developed and tested by researchers.
In addition to statistical modelling, Elo and her team applies different machine learning techniques to create predictive models. The machine is taught to learn the essential factors from the data. For example, the machine can learn to provide binary predictions of the consequences of treating an illness with medication: good response/bad response.
“New tools and methods must be brought as close to the patient as possible. We are constantly thinking what is required so that the model can be used in treating patients. What should be measured and how? Is there anything that could be done better? The model must be sufficiently simple and easy to use in order to end up at a clinic to be used by a doctor in their everyday work. It is important to know how doctors use them.”
According to Elo, the essential thing about this work is that it is interdisciplinary.
“Just how much more information can be obtained using computational methods than sieving through the data only manually? Computation has become part of medicine.”
The Turku Centre for Biotechnology has its own computer cluster whose computing capacity is supplemented by a connection to the ePouta cloud service of the Finnish ELIXIR node CSC.
“The computing capacity and tools provided by ELIXIR facilitate the utilisation of data produced by other organisations. Utilising European data is important, but the data should be standardised. Making data compatible is a job for a large infrastructure.”
Ari Turunen
8.10.2018
Read article in PDF
Citation
Ari Turunen, Laura Elo, & Tommi Nyrönen. (2018). Disease prediction models are becoming more accurate thanks to the computational methods. https://doi.org/10.5281/zenodo.8118762
More information:
Medical Bioinformatics Centre, University of Turku:
Bioinformatics services provided by the Finnish ELIXIR node CSC:
https://research.csc.fi/biosciences
Biotools, a range of bioinformatics tools provided by ELIXIR:
https://www.elixir-europe.org/services/tools/biotools
ELIXIR collaborates with the US-based GA4GH (Global Alliance for Genomics and Health) to utilise genomic data.
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Biotechnology industry research produces a huge volume of data, and the amount doubles every few months. That is why data management requires sophisticated tools. This can be implemented in cooperation between public biological data infrastructures, such as ELIXIR, and companies, such as BC Platforms.
BC Platforms offers information systems for the management of genomic data. The two heavy-duty databases developed by the company are also used in the ELIXIR infrastructure through the Finnish ELIXIR node CSC. BC Platforms is now in the process of creating an ecosystem where the data sets of biobanks from different countries can be searched by using a common user interface.
BC Platforms has more than 20 years of experience in handling large data sets. The company’s information management systems can be set up in a local computing environment or the cloud. A virtual file system operates in the background. Users log in to the database and retrieve the material from the server. The changes made by users are then saved back into the database, so that huge amount of files are exported and imported by the secured network. This so-called object-based storage is particularly suitable in cases where data needs to be stored for a long time while also taking into account information security.
The items analysed by the customers of BC Platforms range from the data of a single person or animal to cohorts consisting of millions of individuals. The clientele also includes research organisations that produce up to 10,000 genomes a day.
BC Platforms wants to create an open ecosystem between researchers, pharmaceutical companies and biobanks. The BC I RQUEST service provides information about the data in different biobanks. Via the service’s user interface, researchers and drug developers have centralised access to the material of the biobanks belonging to the cooperation network.
Each biobank that has joined the ecosystem has a module developed by BC Platforms that transmits biobank data to the service. According to Timo Kanninen, Chief Architect at BC Platforms, a common biobank user interface benefits everyone.
“We help pharmaceutical companies find the right biobanks with data significant for them stored. For example, using the search term “asthma” allows you to see how many asthma patients have their data stored in the biobanks of different countries. In the past, it was necessary to e-mail the operator of an individual biobank, ask how many asthma patients they had and then wait for a reply.”
The software automatically generates aggregate data, i.e. data collected from multiple sources. As it does not contain personal information, the data can be transferred outside national borders. Identified data of biobanks can be combined in a system once authorisation has been obtained.
“It is possible to conduct smart searches on existing data. The service and ecosystem bring together the data owners, providers and users. Because the users are companies developing drugs, they often want to define the data they need. Our analysis tools are well-suited for this purpose.”
According to Timo Kanninen, the goal is to have the clinical and genomic data of five million patients under the search functions of the common interface by 2020.
“Now we have a broad view of what kind of data is available. We are constantly recruiting biobanks with genomic data in addition to clinical data into the ecosystem. This benefits drug designers as it allows findings to be verified in another population.”
BC Platforms’ application automatically generates metadata, improving the search results from biobank materials. BC Platforms classifies the metadata based on the existing standards. However, the harmonisation of metadata is still a challenge for efficient data processing. Recording practices vary depending on the country and hospital.
“Age, gender and diagnosis are generally known, but surgeries, operations and laboratory values are often recorded in a non-uniform manner. Different information systems add even more challenges”, says Kanninen.
Companies in the bio-industry will not wait for the results of standardisation if it takes years. They have to come up with their own solutions. However, the harmonisation and standardisation of metadata as well as the provision of public databases in standard format would be a big relief and resource. ELIXIR aims to this.
Genetic data is increasingly being used in patient care and the industry. The clientele of BC Platforms includes one of the largest companies providing genetic tests in the world, for which BC Platforms produces the genetic data. Finnish research groups utilise the systems of BC Platforms in analysing plant, animal and human genomes. The University of Helsinki conducts, for example, research related to animal breeding and the researchers need tools to manage genomic data. The data analysed with the BC Platforms system is also used to look for new target areas for drugs and to study the efficacy and safety of drug ingredients.
“We digitise the genetic data into a format that researchers can use in their analyses. It can then be combined with other data, such as clinical or patient data”, says Anita Eliasson, Director of Administration and Development at BC Platforms.
Genomic data can be utilised in cancer research when determining the patient’s cancer type. Based on genomic data, it is possible to know what the drug response is like and what kind of treatment should be recommended.
“We use public databases with information on what kind of genome finding typically has specific treatment responses or what type of cancer it is when a person has a certain genome. This is combined with other information. The patient can be treated in the correct way from the start, saving time and money. Being able to select the right medication saves lives.”
Even though the main database system was developed by BC Platforms, Eliasson stresses that BC Platforms is an ecosystem company that places great importance on a partner network.
“We have developed our information systems together with researchers for a long time. Genetic research is now entering a new phase as information is also needed for uses other than research. We are not aiming to provide analysis services for every purpose. That is why our information system has open interfaces. It can then easily be connected with other analysis methods, such as artificial intelligence.”
BC Platforms’ two information systems, BC I Genome and BC I Insight, are available in the ELIXIR infrastructure through the Finnish ELIXIR node CSC. Research groups have their own virtual server with BC Platforms’ databases and tools. Virtual servers operate on CSC’s computing platform and, if necessary, the ePouta cloud service with increased level of information security.
“Researchers can use these to store genomic and other research data while also being able to perform a very wide range of different genome analyses in the same environment by combining data in different ways.”
The research environment is currently being used by groups from the University of Helsinki studying animal genes.
“It is possible to connect more applications to this environment because BC I Genome and BC I Insight have open interfaces. When analysing human data, if necessary, the material could be stored in an environment with stricter information security, such as CSC.”

Because the processing and combination of data are automated, the research group does not have to perform data conversions or worry about data formats.
“Maintenance is efficient because the environment is consistent. Few research organisations can afford to acquire such a heavy-duty solution and its maintenance for a single research group. This is now possible for bioscientists through the ELIXIR infrastructure.”
According to Anita Eliasson, companies like BC Platforms are in great need to utilize replicated public databases, automatically extracting local copies of the database. Bits do not travel quickly enough from the EMBL databases. Physical distance is a factor when it comes to transferring really large data masses.
“Transferring all the data is not sensible. That is why databases should be replicated at the nodes of Finnish ELIXIR. Companies that want to analyse large data masses with artificial intelligence seek out locations that are physically close to the databases due to data transfer costs.”
Ari Turunen
11.9.2018
Read article in PDF-format
Citation
Ari Turunen, Timo Kanninen, Anita Eliasson, & Tommi Nyrönen. (2018). Genetic data under control and in the desired format. https://doi.org/10.5281/zenodo.8113213
More information:
BC Platforms
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Over the past three years, over 500 TB of sequencing data from Finnish research samples was transferred over fibre optic cable from the United States to Finland. The licensed and protected data were transferred to Finnish biobanks and will significantly contribute to the study of genetic diseases.
Until 2015, there were no resources to return Finnish genomic data generated by international research projects back to Finland. As a result, the Academy of Finland funded a project in which the Aarno Palotie‘s and Samuli Ripatti’s research teams from the University of Helsinki’s Finnish Institute of Molecular Medicine (FIMM) and Finnish IT Center for Science, CSC, started transferring data back to Finland from the genome sequencing centres in St. Louis and Boston.
“We created a good process that included license tracking, data transfer, reliability and security. Not many have transferred such a large amount of material from the United States to Europe. Thanks to Finnish universities’ core network, FUNET, the data transfer rate was sufficient. In addition, CSC has experience in recording massive data files, such as storing all Finnish TV and film production on tape,” says CSC’s Ilkka Lappalainen, Head of Service Development for Health and Life Sciences.
The eSISu project (e-Infrastructure for Sequencing Initiative Finland) securely stores the details in Finnish genomes, that is, gene variations. By analysing variations, new information on hereditary diseases can be discovered. The purpose of the SISu project (Sequencing Initiative Finland) is to aggregate genomics into a form that benefits best Finnish doctors and researchers. To date, the full genome of thousands of Finns and the protein-encoding parts of the genomes of almost 30,000 Finns have been determined.
The data collected from Finns is largely similar to that of other European countries, but certain parts of the Finnish genome have either been refined for this northern condition or they only existed in a few families that inhabited small villages in the north.
“For this reason, certain genetic variations occur in Finns, affecting, for example, cardiovascular diseases. If we do not have data on our genetic composition, how can we investigate the genetic effects of different diseases?”, asks Ilkka Lappalainen.
Genomic data is part of an integrative entity that combines lifestyles, medications, treatments and individual health data. It will allow for accurate statistical studies where the combined health effects of individual differences in genetic makeup and varying response to medication can be assessed.
“In certain cases, e.g. cancer treatments at the Helsinki University Hospital (HUS), this is already in use. Huge amounts of statistical data are being collected from cancer causing genes to redefine procedures and medical therapies and recommendations.If standardised data can be obtained from the entire population of Finland, people can, if necessary, be called in for cancer screening and decisions can be made about the appropriate treatment. Future forms of treatment are not possible with the data only collected from Finns. Cancer treatments are largely developed through international cooperation.”
The hope is that this knowledge will be passed on to general health care that notifies the person who gave the sample. It is then up each person to decide if they want to know more about their risks.
One of the most important research topics in bioinformatics is to understand the mechanisms causing diseases. Part of the data collected during the eSISu project came from migraine patients. In summer 2016, the project reached a major milestone when the first set of data for the migraine genome was transferred to Finland. The data transfer was executed without any technical or security problems.
FIMM’s scientists have been using SISu data to verify that inherited susceptibility to migraine does exist, and showed that genetic starting points for migraine sensitivity can be traced to 38 regions in the genome. The finding is a first step in understanding the mechanisms of migraines and opens up the possibility of more accurate diagnostics and treatments.
Finnish researchers have also found new genetics variations that affect the susceptibility to coronary heart disease. Thanks to this new information, risk groups for coronary heart disease can identified earlier and instructed to start preventive measures that might include lifestyle changes or preventive medication.
There is still plenty work left to do with the collected eSISu data. There are cases where several samples have been taken from one individual and sent to different places for analysis. Now these all will be traced back to the source.
“We are now working with metadata to find out what was collected earlier and add value to future research projects.”
According to Lappalainen, a lot of valuable experience in data management was gained through the project. This will be useful for the new FinnGen project.
Started in December 2017, the FinnGen project will store the genomes of half a million Finns. The project utilises samples collected by Finnish bio banks. The data from the genome is combined with the information in national health registers. This makes it possible to better understand the mechanisms by which diseases are born and to then develop new treatment methods.

SISu has already been identified as a significant data resource for ELIXIR and BBMRI infrastructures. In the next phase, the organisation and management of data will move to a scalable and secure platform (ePouta cloud service). This will make data computationally available. Finnish biobanks, such as the THL biobank, will continue to oversee the use the material and grant permissions for the use of it.
“Currently, data transfer is being tested and it will continue to work once the metadata has been updated.”
eSISu is creating Finnish capability to manage controlled access to genome data between the Finnish ELIXIR Centre and other ELIXIR Centres. It enables CSC, with consent from the data owners, to integrate these data with other registers and databases in Finland.
“This way, we can combine Finnish data with European EGA (European Genome-phenome Archive) data.”
The European Genomic Archive is one of the world’s largest public data repositories with patient data from biomedical research projects. EGA shares human genetic and phenotypic data through a consent process that allows the reuse of data for research purposes. Thanks to EGA, many ELIXIR research projects are possible.
www.sisuproject.fi is a search service where you can find more about genetic variants of the Finnish population. The KITE search engine, on the other hand, searches the data on the basis of the metadata. These are examples of services that are being developed for both national and international use. Data management and licensing practices are handled using the new ELIXIR REMS (Resource Entitlement Management System) software.
“Technically, the data management works well. A significant part of SiSu’s material will be available during 2018.”
Ari Turunen
23.8.2018
Read article in PDF
Citation
Ari Turunen, Ilkka Lappalainen, & Tommi Nyrönen. (2018). Massive data management project. Finns’ heredity is collected and safeguarded. https://doi.org/10.5281/zenodo.8113203
More information:
Tommi Nyrönen
Head of Node, ELIXIR Finland
tommi.nyronen@csc.fi
+358-50-3819511
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Up to 50% of all the approved drugs affect only three protein families: nuclear receptors, G protein receptors and ion channels.
Drugs usually affect the cell receptors or enzymes in the body, both of which are proteins. Many drug molecules also bind themselves to enzyme receptors and transport proteins on the cell membrane. The drug may, for example, bind itself to the active site of an enzyme, thereby inhibiting the chemical reaction controlled by the enzyme. Most often the enzymes that catalyse the chemical reaction caused by the drug are metabolised by cytochrome P450 enzymes before they are in active form.
Most of the target proteins of drugs belong to only ten protein families, and up to half to only three families. Proteins belonging to a specific family have a similarly folded three-dimensional structure, function and significant similarity in amino acid sequences, which usually indicates a common ancient history. Proteins of the same family are derived from a single original form which, through evolution, has adapted and specialised due to environmental pressures as well as functional roles that differ from their original role in cellular processes.

Protein families were discovered when the structure and amino acid sequences of a few proteins began to be known. It was then found that proteins consist of several independent, structurally distinct areas with a special task. These became known as domains.
New protein families have been discovered when studying the underlying mechanisms of various diseases. Nuclear receptors, for example, were discovered while studying breast cancer. It has been known for a long time that tumour growth ceased in one third of women with breast cancer whose ovaries or adrenal glands had been removed. However, the molecular basis of breast cancer was still a mystery. In 1947, medical researcher Elwood Jensen began to investigate this. Jensen discovered the estrogen receptor and found that the estrogen receptor is activated when its natural estrogen, estradiol, binds itself to it. After this, the activated estrogen receptor travels to the nucleus of the cell where it participates in regulating the function of genes.
The estrogen receptor, a protein molecule belonging to the nuclear receptor family, is very important to humans. If changes occur in its function, they have a great significance for cell health. The estrogen receptor plays an important role in the emergence of breast cancer. Normally, estrogens regulate the activity of the estrogen receptor in the cell. The changed shape of the estrogen receptor is active all the time, meaning that the normal regulation mechanisms of the cell based on the level of estrogen do not function correctly. This can lead to cancer, i.e. uncontrolled growth of abnormal cells.
Elwood Jensen proved that breast cancer patients with a low estrogen receptor concentration in their cancer cells did not benefit from the removal of the ovaries. The ovaries produce a large proportion of women’s active oestrogen. The receptor concentration indicates who should have surgery and who should skip it. In the mid-1970s, Jensen and his colleague Craig Jordan discovered that cancer patients whose mutated tumour cells had a large number of oestrogen receptors were also likely to benefit from tamoxifen. It is an antioestrogen, meaning that it overrides the effect of estrogen in cells. The patients with low numbers of receptors, in turn, could immediately be transferred to other treatments. By 1980, the test developed by Jensen, which was used to measure the number of receptors in breast cancer samples, had become a standard test for breast cancer patients.


The discoveries revealed a protein superfamily functioning in the cells, the nuclear receptors, which includes the oestrogen receptor. The nuclear receptor family includes, among others, the estrogen receptors alpha and beta, androgen receptor, progesterone receptor and vitamin D receptor. What nuclear receptors have in common is that they are activated when a cell membrane-penetrating the signal molecule, i.e. a ligand, a nuclear receptor hormone, binds itself to them, after which they travel to the nucleus to influence cellular processes. Hormones that activate the members of the nuclear receptor superfamily include, among others, testosterone, estradiol, progesterone, glucocorticoids, mineralocorticoids and vitamin D as well as molecules created through drug design that mimic the structure of natural ligands. For example, the lip balm of Therese Johaug of Norway’s 2016 national skiing team may have contained clostebol, a ligand of the androgen receptor. Clostebol functions as an anabolic factor, meaning that it promotes muscle cell protein growth.
Small molecules introduced to the human body through drugs or other routes can thus affect nuclear receptors by activating or deactivating them, thereby affecting the functioning of the cell’s genes. The discovery of nuclear receptors has revolutionised biochemical endocrinology research. Endocrinology is a speciality that studies and treats diseases of hormone-producing organs. The diseases can result from the excessive production of hormones or lack thereof; furthermore, both benign and malignant tumours can occur in endocrine tissues. Prior to the discovery of nuclear receptors, the functioning of the hormones in the human body was a complete mystery. Now it can already be slightly modified.
In order for an organism to function, signals must be transmitted in the body’s cells and the organs they form. The body as a whole sends and receives signals through electrical currents and certain molecules. Martin Rodbell and Alfred Gilman determined how signal transduction occurs through the cell membrane via cooperation of molecules. In 1970, Martin Rodbell proved that the signal transmission takes place in three stages: signal reception, transmission and amplification. The transmission takes place so that a cell surface protein transmits a command to exchange the guanosine diphosphate (GDP) bound to the protein located on the other side of the cell membrane for guanosine triphosphate (GTP). This phenomenon is data transfer at the molecular level.
In 1980, Alfred Gilman studied leukaemia cells and found that they did not respond to the external signals transmitted by hormones. This was due to a mutation of the receptor protein which caused the signal transduction of hormones to be inhibited. Gilman isolated the protein from normal cells, and, with these proteins, he was able to repair the damaged cell. The molecules involved in the signal transduction are a large family of proteins that bind themselves to guanosine triphosphate. When they are bound to GTP, they are ‘on’, and, when they are bound to GDP, they are ‘off’. He called them G proteins (also known as guanine nucleotide-binding proteins).
G proteins are perhaps the most important molecules involved in signal transduction. In addition to some forms of cancer, they are associated with diabetes, alcoholism and the underlying molecular mechanisms of many other diseases.
The protein family of the G protein-coupled receptors in the cell membrane conveys signals to the G proteins inside the cell membrane. G proteins, in turn, respond by exchanging GDP for GTP. The consequence of this activation is, for example, an enzyme released for the breakdown work in the cytoplasm inside the cell, opening or closing of an ion channel located in the cell membrane.
This mechanism is how rhodopsin, with which we detect light, or see, with our eyes, works in the eye, for example. One third of all the known drug ingredients affect G protein-coupled receptors. Catecholamines (e.g. adrenaline, noradrenaline and dopamine), peptides, glycoprotein hormones and rhodopsin are examples of ligands that bind themselves to these receptors. Alfred Gilman and Martin Rodbell received the 1994 Nobel Prize in Medicine for the discovery of G proteins. In 2012, the Nobel Prize for Chemistry was awarded to Robert Lefkowitz and Brian Kobilka for explaining the workings of G protein-coupled receptors.

Proteins perform their work cyclically and accurately by exchanging shapes and molecules based on signals. The shapes of the G protein and G protein-coupled receptors are in a continuous, dynamically changing biochemical equilibrium reaction with each other. Changes in the sensitive balance of G proteins may result in illness. For example, the toxin of the cholera bacterium locks G proteins into one shape and affects the nerves that control the absorption of salt and liquid in the intestines.
Beta-amyloid (Abeta) peptide, 3D rendering. Major component of plaques found in Alzheimer’s disease. Stick representation combined with semi-transparent molecular surface.
Some ion channels are complex, multi-part structures that are huge in terms of molecule size. Huge ion channel-coupled receptors react directly to tiny ligand molecules, such as the ionotropic receptor located in the brain that reacts to the amino acid glutamate. The protein, consisting of four domains, readily changes its shape as the thousands of times smaller glutamate binds itself to the domain for signal transduction and opens the ion channel permeating the cell membrane. Memantine is used for Alzheimer’s disease. It protects brain neurons from destruction by blocking the excessive glutamate transmitter effect.
Receptor-gated ion channels open when a particular chemical compound binds itself to them. The chemical compound may be an extracellular molecule, such as a hormone, a neurotransmitter, a drug ingredient or a toxin, or an intracellular molecule.
By understanding the functioning of ion channel receptors, researchers can develop, for example, treatments for addiction by changing the activity of the receptors.
Ari Turunen
Tommi Nyrönen
14.6.2018
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2018). Half of all drug ingredients affect only three protein families. https://doi.org/10.5281/zenodo.8113184
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

A good drug molecule will not be created unless it is known which proteins it affects in our body. That is why, in drug design, it is important to utilise massive databases with all the discovered protein structures and protein families as well as knowledge about how proteins function in our cells.
The majority of the drugs in use are designed so that their target molecules are the body’s biomolecules, i.e. proteins.
Most drugs take effect in the body by binding themselves to these targets like receptors of signal molecules. Receptors are natural targets for example for signal molecules such as neurotransmitters and hormones. They are specialised triggers of the cell associated with cellular signalling mechanisms.
The idea for drug design is to build small synthetic molecules that selectively affect the desired proteins. Most of the target proteins of drugs belong to only ten protein families, and up to half belong to only three families. Small molecules are able to absorb well into the bloodstream, allowing the drug to take effect. Depending on the location of the protein, the drug molecule has to penetrate the cell or transmit a signal outside the cell that affects the processes within the cell. The aim is to design the molecules, for example, in such a way that they slow down or accelerate the functioning of a particular protein.
In the past, little was known about in which part of the cell the drug takes effect. In 1980, 150 of these target areas of effect were known. However, that number has grown enormously with the determination of the genomes of organisms since, currently, already more than 5,000 possible target areas of effect are known. Approximately 2,500 drug molecules are available for medicine. The function of the human genome is being investigated more and more closely and, in the next few years, the number of known possible target areas of effect for drug ingredients may rise to 10,000. According to current estimates, our body has 2,000–3,000 proteins that are possible target proteins for a drug. Existing drugs have been shown to work through only about 450 drug targets on a limited number of diseases. Thus, drug designers have two major goals – to build new safe molecules that can be used to safely affect known targets and, on the other hand, to study the use of known, safe drugs for new illnesses for which there is currently no drug approved by the authorities.
The goal of researchers is, among other things, to understand which structural and chemical characteristics of a drug molecule play a key role as they modify the function of proteins at the cellular level.
An effective drug can be developed once a three-dimensional structure of the target protein, which allows interaction with the drug molecule, is found. Chemical counterparts that recognise the amino acids at the protein’s binding site are built into the drug molecule. When this kind of molecule encounters the target protein in the body, it automatically finds its way to the binding site of the protein because attaching itself there is energetically advantageous for it.
The binding of a well-designed drug molecule to the target protein could be compared with putting on a wool glove. It fits firmly on the hand with precisely five fingers: it would be very uncomfortable for one with six or seven fingers. In addition, a left-hand glove fits poorly on the right hand.
The shape of proteins tells more about the function of the molecule than the amino acid sequence. Proteins with the same shape can function similarly biochemically even if their amino acid sequences differ from each other by more than 80%.
Once the structure of one member of a protein family has been determined, the structure of other proteins belonging to the same family can be predicted by modelling. Modelling which is carried out using a computer speeds up research because hundreds of times more protein amino acid sequences are known than protein structures that have already been determined by testing. It can roughly be said that the task of genomics is to determine the sequence of nucleotides. This sequence is translated into an amino acid polymer in the cell, but it starts to function only after the protein folds up into its three-dimensional shape. This function is investigated through proteomics. Thus, cooperation between experts in genomics, proteomics and drug molecule modelling supports one another.

Even though there is much information, the development of new drugs is quite challenging. Only 5% of drug ingredient candidates progress through laboratory testing even to treatment tests on animals. Of those, only a few per cent will ultimately be suitable as medications. It has been estimated that up to 75% of the price of drugs is due to the costs of failed pharmaceutical development projects.
One major challenge is minimising side effects. With the development of genomics, drug molecule have been found to have an individual effect. Historically, drugs have been developed assuming that people are similar in terms of their biochemistry but, in reality, we are unique at the cellular level in the same way as people are slightly different physically. When small drug molecules are used to try and influence the situation of a diseased body in a healing way, these individual differences at the molecular level may affect the performance of the drug.
By collecting and storing human biological data, it will be possible in the future to target drug molecules for treatment purposes that do exactly what they should in that exact situation, and tailored to the person who needs the medication. This is called personalised medicine.
A particular gene produces a specific protein affected by the drugs. When the DNA base sequence of a person’s genome is known, it is also possible to deduce the basic structure of the corresponding protein in that person. Like DNA, a protein is also a string consisting of successive building blocks, and a specific block of a gene always corresponds to a specific block of a protein.
One person may have – inherited or caused by the environment – a change in one DNA nucleotide that is reflected in a protein through this chain. That change may be just where the protein should receive signals from elsewhere in the body or interact with a drug molecule. By storing protein structures and sharing them to be used by researchers, this phenomenon can be controlled and understood. The shapes of the drug molecule and the protein molecule can be matched to each other so that the drug is adapted to the situation, allowing the drug to adhere and take effect as effectively as possible. Many cancer treatments are based on this. The genome of a tumour changes over time. Tumours at different stages can be affected through drugs, but the shape of the drug molecules must take into account the changes in the shape of growth-stimulating proteins.
That is why especially proteins whose three-dimensional structure can be determined by tests or predicted through modelling are studied in drug design. The adherence of the drug molecule can be studied using modern computer modelling software in which the three-dimensional protein and drug models are matched to each other. This also enables the tailoring of the ideal drug shape.
Usually, a drug takes effect by adhering to a defective protein in the body and altering its function. An ideal drug does only this; it does not interfere with healthy proteins or cause other side effects. Up to the present, we have been happy to find one protein affecting a disease and a drug molecule that is moderately effective against it.
Now, the entire arsenal of proteins and drug molecules can be screened and the best candidates selected. This is due to the advancement of molecular biology, computer computing power and databases. It is now possible to screen the entire protein range of the body.
The Protein Data Bank, i.e. the PDB protein database, includes more than 100,000 protein structures divided into protein families. The members of a protein family are usually similar in terms of their three-dimensional structure, which is why they also function in a similar manner.
The PDB database is maintained by the international consortium Worldwide Protein Data Bank (wwPDB). It is tasked with maintaining individual macromolecular structural data that is freely available to researchers.
The Human Protein Atlas is a Swedish-based programme started in 2003 with the aim to map all the human proteins in cells, tissues and organs. Various omics technologies are used in the mapping, meaning technologies in which all genes or the proteins produced by them are studied simultaneously. These include antibody-based imaging, mass spectrometry-based proteomics, transcriptomics and systems biology. All the data collected is open to researchers.
In January 2015, the Human Protein Atlas published a map showing the locations of 17,000 different proteins in the human body, providing valuable information for drug design. The map included the locations of proteins that were the target proteins of approved drugs. Researchers can view proteins in 32 different tissues, representing all of the most significant tissues and organs in the body.
In December 2017, the Human Protein Atlas released version 18. At that time, the database contained 26,000 antibodies targeting proteins encoded by almost 17,000 genes. It corresponded to 87% of protein-encoding human genes.
Tommi Nyrönen
Ari Turunen
12.6.2018
Article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2018). Looking for a good drug. https://doi.org/10.5281/zenodo.8113165
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org
The majority of the microbes in humans are located in the intestines. The intestinal microbes form their own ecosystem. With metagenomics, it is possible to also study them better in their natural habitat, be it soil or the intestines. Anu Kantele, Professor of Infectious Diseases from the University of Helsinki, has been studying bacteria and diarrhoeal diseases for a long time. “I am interested in knowing how new bacteria that arrive in the intestines manage to settle into the ecosystem formed by the native intestinal bacteria and how antibiotic treatment affects this”, says Kantele.
Enterotoxigenic coliform bacteria (ETEC) are one of the most common causes of severe diarrhoeal disease. Anu Kantele and her research group participates the vaccine study which is being conducted in collaboration between the universities of Helsinki, Gothenburg and Johns Hopkins and the vaccine manufacturer, Scandinavian Biopharma AB. This so-called ETVAX vaccine has been estimated to provide protection against moderate to severe ETEC diarrhoea in 60–80% of those vaccinated. For tourists, diarrhoea is usually just an unpleasant experience, but it is life-threatening for the children in developing countries. That is why the developing an ETEC vaccine is also one the goals of the WHO. Read the article here
Biobank operators convened at CSC to discuss national cooperation at the IT day for biobanks on 2 May 2018. Finnish biobanking operations have become significantly more active. BBMRI.fi National Coordinator and Docent Anu Jalanko stated that, in 2017, there were already more than 300 ongoing biobank studies in Finland.
BBMRI.fi is an infrastructure that provides strategic support for biomedical research, health care as well as drug and product development for nursing. There are 10 biobanks in Finland, 9 of which are closely involved in the BBMRI.fi cooperation.
The Finnish Biobank Cooperative – FINBB is a cooperative established by six hospital districts and six universities to serve and act as a service coordinator for its biobank customers.
In her opening remarks, Anu Jalanko said that the services and functions of BBMRI.fi and FINBB will be fully integrated, meaning that ICT services will also be harmonised.
Professor of Pharmacogenetics and Chief Physician Mikko Niemi is leading a research group at the University of Helsinki studying how genes influence the effects, safety and efficacy of drug ingredients. He is also investigating when genetic tests should be considered in drug selection.
“If the genomes of patients were tested systematically, drug treatments could be better tailored and their dosages measured more individually”, says Niemi. Around a dozen genetic tests related to drug treatments are currently available in Finland. The goal of Professor Mikko Niemi is to devise an interpretation algorithm that helps doctors determine the appropriate drug and correct dosage for a patient. Treatments become more effective and side effects are reduced, thus also decreasing the costs. Read more in the article.

The human genome contains millions of genetic variants that make each individual unique. Some variants affect eye colour or blood type and others affect hereditary diseases. The DNA sequence may also include a pathogenic sequence variant that causes various disruptions in the function of the gene. The disruptions manifest themselves as hereditary diseases. Blueprint Genetics from Finland classifies genetic variants found in the genome from patient samples and analyses their connection to the described symptoms of the patients.
Blueprint Genetics started its operations focusing on the diagnostics of cardiovascular diseases. The company is now able to analyse majority of hereditary diseases based on the patient samples it receives. More than 6,000 disorders resulting from a defect in a single gene are known in humans. On average, one in two hundred will inherit a genetic defect from their parents. There are also many multi-factor disorders in which the combination of multiple genetic variants causes the disease or increases the risk of illness. These include, for example, Alzheimer’s, diabetes, rheumatoid arthritis or cancer.
Jussi Paananen, Director of Data Science at Blueprint Genetics and researcher at the University of Eastern Finland, has a background in computer science with data science as his field of specialisation. Paananen became interested in biomedicine at an early stage because it utilises technologies that produce a lot of data. In recent years, he has been interested in machine learning and artificial intelligence, which are on their way to becoming research methods in bioinformatics thanks to increasing computing power.
“I am interested in how artificial intelligence can help geneticists in decision-making as well as processing large amounts of data.”
Research into artificial intelligence is on the rise and the methods are changing. In machine learning, the computer learns to arrive at a particular outcome independently. Machine learning algorithms find patterns that people are not able to detect from large data sets. Machine learning utilises neural network research, which has a long tradition in Finland. The neural network learns the non-linear dependencies of the variables directly from the observation data. It is able to classify the ears from animal-themed images, for example.
“Neural networks are at their best in solving classification problems”, says Paananen.
“In image analysis, images or parts of images are identified and classified. A machine can identify objects and things: this is a human, this is a car, this is a cancerous tumour. What we do is classify DNA variants. From patient samples, we try to find which DNA variants cause diseases as well as which genetic variants are a part of our normal genome.”

The customers of Blueprint Genetics are doctors treating patients. The doctors want to find out whether the illnesses of their patients are due to hereditary factors or not. Doctors from around the world send Blueprint Genetics their patients’ blood or saliva samples, the isolated DNA of which is then sequenced. Sequencing generates a huge amount of data from which the interesting variants are drawn. In practice, this means that the patient’s genetic variants are compared to the average human reference DNA.
Blueprint Genetics employs top professionals, geneticists and doctors who classify the variants. They go over the data mass that has already been processed and divided into smaller parts. The experts practically sieve through existing scientific literature and databases.
“We are trying to figure out which of these variants explains the disease or its symptoms.”
Since similar information has been collected around the world, a single DNA variant that explains the disease can often be found in scientific articles and databases.
“We issue a clinical statement based on the results. The clinical statement is typically a few pages long document, that is delivered to the customer physician. The physician uses the statement as an aid in diagnosis and planning of treatment.”
Blueprint Genetics utilises a variety of data sources. Where possible, the analysis of the data is automated. Software analyses the data and performs complex data processing. The field is under constant development. Software is updated several times a year, data volumes and computing power are increasing. Methods evolve and change rapidly.
Initially, Blueprint Genetics focused on certain interesting genes, or gene panels, based on the patient’s symptoms. A panel typically includes about a hundred known genes associated with a particular disease. A team of geneticists sieves through the approx. 2,000 variants studied using the panel. The company has now shifted to exome sequencing, meaning that it sequences all protein-encoding genes, of which there are approx. 21,000 in our genome.
The human exome is the part of DNA with which all human proteins are produced. The part of the gene that encodes and directly guides protein production is called the exon. All the human exons in our genome together are called the exome. The human exome is approx. 1.5% of the entire genome.
“When our analysis focused on gene panels, we obtained, for example, 2,000 variants that a team of geneticists went through. Now, there may be 200,000 variants. As we advance to sequencing the entire genome, the number of variants will be 5 million. This amount of data cannot be sieved through manually.”
External databases are important in interpreting the data collected from patient samples. Genomic variants have been catalogued in various international databases, the most important of which are located in the organisations of EMBL-EBI in Europe and NCBI (The National Center for Biotechnology Information) in the US. In addition, ELIXIR coordinates the public biomedical infrastructure in Europe, enabling genetic variants to be mined from these international databases.
Variant databases provide useful lists that can be used to find correlations between genetic variants and phenotypic data. EMBL-EBI classifies, stores and distributes information on genetic variants. The most important databases include the European Genome-phenome Archive (EGA) where patient data from biomedical research is stored, the European Variation Archive (EVA) that includes genetic variants, Ensembl that provides interpretation for these variants, the gnomAD service for population-level variant occurrence data and the ClinVar archive for clinically significant variants. Therefore, the doctor often needs information from more than one service in order to produce the correct interpretation of the genomic variant for the patient. For this reason, European and American services regularly exchange information on the latest research results so that the services would always provide the latest information on our genome for research and medicine.
“Genetic variant databases are important because they have information on the prevalence of the variants in healthy people. This information can be utilised, for example, when it is known that only 1% of people have a certain rare hereditary disease. When we see that there is a variant that 5% of people have, it can be concluded that this cannot be the variant causing the disease. Thus, it is possible to filter out major, common DNA variants that cannot be associated with the rare disease.”
Public sector data services offered by ELIXIR are important.
“We utilise our own local copies of different data sources. Physical distance and communications links require the sources to be in the same place. From public services, I would like to see more measures related to the versioning of databases. Old versions should not be discarded. Long-term storage should be available for different versions.”

A major challenge in both public research organisations and the private sector is the standardisation of the data used for interpretation. Data notations can vary greatly. The big challenge for Blueprint Genetics is the so-called phenotypic data.
“In one sense, it is metadata in itself, i.e. information accompanying a patient sample: symptoms, diagnosis and other background information. A sample may be accompanied by a lot of metadata or none at all.”
The standardisation of phenotypic data has the same problem as patient data in health care, where the challenge is different notations.
“We obtain information from different countries that has been recorded in different ways. The background information varies.”
Jussi Paananen thinks that firms like Blueprint Genetics find it difficult to utilise data produced and managed by publicly funded and research-focused organisations.
“Research organisations and joint infrastructures are interested in large population cohorts, in which case we are talking about a huge amount of data being collected and harmonised. We process information in different ways than cohorts which, for example, compile the information of tens of thousands of people living in the same geographical area. We, however, always deal with individuals.”
Blueprint Genetics seeks to use internationally consistent classification, terminology and standards in its operations.
“We produce the DNA data ourselves and can decide what form it is in and which standards it conforms to. However, we utilise guidelines provided by others when interpreting the results.”
The first attempt at such a standard was made a few years ago. The American College of Medical Genetics and Genomics (ACMG) has issued guidelines on how sequence variants should be classified. ACMG has proposed the following common terminology for single-gene disorders: pathogenic, likely pathogenic, uncertain significance, likely benign and benign.
“We have our own modified version of ACMG’s classification.”
The challenge for companies like Blueprint Genetics is the ability to utilise data. There is a lot of information in peer-reviewed publications, and the aim is to develop good text mining tools in order to automate the screening of articles.
“There should be centralised access to all publications. We have now long been negotiating with academic publishers about licence fees, which are high.”
“We have our own software production combining different data sources and facilitating literature searches. However, the final interpretation is always carried out by a geneticist.”
Analysis and interpretation of patient data is demanding work because it involves a lot of legislation and regulation. Blueprint Genetics provides medical doctors with processed information, but the doctors always make the actual decision.
Blueprint Genetics is also interested in cooperation between the public and private sectors.
“The utilisation of genetic data is an enormous challenge that concerns the whole human race. The solution requires cooperation from companies, academic research groups as well as publicly funded organisations. Blueprint Genetics strives to contribute to the development of open science solutions and is constantly looking for new partners.”
Ari Turunen
29.5.2018
Read article in PDF
More information:
https://www.elixir-europe.org/platforms/data/core-data-resources
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org
Bank of million patient samples is getting bigger. More than a million tissue samples and tens of thousands of blood samples are stored in Auria, the first hospital biobank in Finland.
The sample collections of Auria Biobank, which operates in connection with the Turku University Hospital and the University of Turku, are physically located in hospitals in southwest and west Finland. Samples are collected and combined with the necessary metadata, indicating the clinical information on the sample donor, quantity, date and how the sample has been processed. The samples of Auria Biobank include tissue, blood and DNA isolated from cells.
Read the interview of Perttu Terho, Vice Director of Auria Biobank.
Mapping the genomes of all organisms enables the new vaccines and medicines. A formidable task is to understand the information contained in the genome of various organisms and humans. It will require cooperation between various research organisations and well organised databases. To do this, also the new kind of linking and analysis of the sources of data will be needed. Through the Finnish ELIXIR centre the researchers have had access to a DNA bank of dogs and cats, the data of which has allowed us to discover the gene of a nerve degeneration disease, for example.
Read the rest of the article here.
A data management software program provides security and only grants access to authorised biomedical material. CSC, the Finnish ELIXIR node, develops and maintains the open source REMS tool that can be used to manage access to datasets containing confidential material. REMS (Resource Entitlement Management System) is an access management tool that, where necessary, prevents the illegal use of data. With the REMS tool, it is possible to order a specific file from a large amount of data and have it delivered to the ordering party locked in a secure manner. The researcher creates a secure analysis environment for the data to be analysed. This is made possible by the REMS tool.
Read the interview of REMS Product Owner Tommi Jalkanen from CSC.

In recent years, intestinal bacteria have become more and more resistant to antibiotics. The bacteria that cause diarrhoea are also increasingly resistant. Antibiotic-resistant bacteria are a global threat. Professor Anu Kantele is interested in knowing what happens in the intestines as people travel to the tropics and back home again.
Regular Finns have been sent to Benin as subjects to test a new diarrhoea vaccine. The ETVAX vaccine is administered as an oral solution, similarly to heartburn medicine. Once the vaccine is on the market, it is planned to be sold to developing countries at a low price.

For tourists, diarrhoea is usually just an unpleasant experience, but it is life-threatening for the children in developing countries. Diarrhoeal disease is the second largest cause of death in children under five years old in the world. Every year, more than 1.7 billion children fall ill with diarrhoea. Of them, more than half a million under 5-year-olds die according to the World Health Organization (WHO). Diarrhoea is also the main cause of malnutrition, short stature and impaired learning ability in small children.
Diarrhoea is a symptom brought on by disease-causing bacteria, viruses and parasites that have reached the intestines. They generally spread through water and food contaminated by faeces. Diarrhoea is transmitted especially when there is a lack of adequate hygiene and clean water for drinking and household consumption. According to UNICEF, in 2010, up to one fifth of the world’s population had to relieve themselves outdoors. Almost 900 million people suffer from a lack of clean drinking water.

On trips to the tropics, the share of those who fall ill with diarrhoea may be up to 60%. Bacteria cause 80% of the cases of traveller’s diarrhoea. Enterotoxigenic coliform bacteria (ETEC) are one of the most common causes of severe diarrhoeal disease. That is why there is demand for a vaccine that works against diarrhoea. The pathogens have been studied extensively both in developing countries and with tourists. The information has been used to develop a new vaccine designed to train the human immune system to identify and obliterate pathogens before they can cause symptoms.
The Escherichia coli bacteria, or coliform bacteria, normally live in the intestines of humans and animals. More than 700 types of E. coli have been identified. They are part of the normal microflora of everyone’s intestines and are mainly useful. For instance, they protect us against many disease-causing microbes.
“E. coli is the most studied bacterium in the world. It is usually not dangerous, but some are disease-causing. There are several known so-called diarrhoea coliform organisms that cause diarrhoea.
ETEC is one of them. It causes severe watery diarrhoea”, says Anu Kantele, Professor of Infectious Diseases from the University of Helsinki.
ETEC differs from other types of coliform bacteria in that it produces two toxins that cause significant fluid secretion from the small intestine, that is, watery diarrhoea.
“The tolerability of the vaccine and the immune defence it elicits are now being studied, while also investigating its efficacy against traveller’s diarrhoea. Developing an ETEC vaccine is also one the goals of the WHO. This so-called ETVAX vaccine generates a good immune response and is the most promising of the current ETEC vaccine candidates.”

ETEC strains of bacteria were among the first pathogenic organisms for which molecular diagnostics were developed. Vaccine researchers are currently interested in several diarrhoea-causing microbes, such as ETEC bacteria, Shigella bacteria and norovirus. Vaccines already exist against other major causes of diarrhoea, such as cholera and typhoid bacteria and rotavirus.
The study is being conducted in collaboration between the universities of Helsinki, Gothenburg and Johns Hopkins and the vaccine manufacturer, Scandinavian Biopharma AB. United Medix Laboratories Ltd is also involved. The safety of the vaccine has previously been tested on 140 Swedish adults and 450 Bangladeshi children. In both studies, the vaccine and placebo groups had an equal amount of side effects, so the vaccine is considered to be very safe.
The results of the Swedish research group have demonstrated that the ETVAX vaccine elicits a strong immune response not seen in those who receive the placebo.
“Among the Bangladeshi children, the vaccine was well-tolerated and the response was good. Its safety was studied previously, and now we are studying the effectiveness of this oral vaccine for the first time”, says Anu Kantele.
The vaccine has been estimated to provide protection against moderate to severe ETEC diarrhoea in 60–80% of those vaccinated. Studies on the cholera vaccine have been a great help in the development work for the ETEC vaccine. The pathogenic mechanisms of the ETEC bacteria and cholera bacteria (Vibrio cholerae) are very similar. They cause illness by attaching to the surface of the small intestine and producing enterotoxins, or intestinal poison, that are responsible for the symptoms of the disease. The toxin kills cells by preventing their protein synthesis. The toxin makes the mucous membrane of the intestine permeable, whereupon a lot of water passes from the tissues to the intestine. This causes very severe watery diarrhoea. The cholera toxin and ETEC bacteria toxins are structurally, functionally and immunologically similar.
ETEC bacteria produce both heat-labile (LT) and heat-stable (ST) toxin. Both have a protein structure and are toxic to humans. ETEC carries a plasmid, a circular DNA molecule that guides the production of the toxin.
“The ETVAX vaccine contains a number of components: killed ETEC bacteria, so-called colonisation factors that allow the bacteria to reproduce in the intestine, and detoxified LT toxin and an adjuvant produced from it”, says Anu Kantele.
Once the samples of the 800 test subjects who travelled to Benin have been analysed, it will be possible to evaluate, in particular, how an immune system trained by the vaccine works against the ETEC bacteria contracted on the trip and its LT toxin. Half of the subjects have received the vaccine and the other half a placebo.

Anu Kantele and her team study the microflora, pathogens and resistant bacteria in the human intestines from stool samples. The participants provide various samples for the study. The analysis of ETEC bacteria and other pathogens requires stool samples. Blood and saliva samples are used to study, for example, the immune response to the ETEC vaccine. Data is also collected on the possible adverse effects of the vaccine. Diarrhoea samples are collected and processed immediately on site in a laboratory in Benin. The number of samples obtained is essentially huge. One gram of human faeces can contain up to one million bacteria.
The researchers compare cultivation-based and molecular laboratory methods used to identify ETEC and other causes of intestinal infections from the stool samples. They analyse the antibodies and genes involved in immune defence.
Anu Kantele has been studying diarrhoeal diseases for a long time.
“I am interested in knowing how new bacteria that arrive in the intestines manage to settle into the ecosystem formed by the native intestinal bacteria and how antibiotic treatment affects this”, says Kantele.
Antibiotic-resistant strains of bacteria are most likely to develop in the poor countries of the world. The reason is the excessive use of antibiotics. If the sanitary conditions are inadequate, resistant bacteria spread easily and even from one country to another. According to the WHO, for example, the bacterial strain resistant to fluoroquinolone, which is commonly used to treat urinary tract infections caused by coliform bacteria, is widespread.
“The Benin test subjects will provide a lot of data that can be used to analyse the efficacy of the vaccine. In addition, the intention is to use genetic engineering techniques to analyse the microbes in the stool samples and to investigate the presence of antibiotic-resistant strains. The amount of data is enormous but, by combining data, it is possible to gain new insight into, for example, the spread of antibiotic resistance.”
According to Kantele, the majority of the antibiotics used by tourists are taken for traveller’s diarrhoea. The antibiotic shortens the duration of the disease, but it would almost always go away by itself also without antibiotics. Kantele emphasises that the symptoms can be alleviated with drugs that affect the functioning of the intestines without increasing the risk of contracting resistant bacteria.
“Antibiotics facilitate the settlement of resistant bacteria into the intestines, and so one of the best ways to avoid such colonisation is to not take antibiotics. Nowadays, antibiotic treatment is usually recommended only for severe diarrhoea; less severe cases are treated with fluid therapy and possibly drugs that affect the functioning of the intestines. The carriers of antibiotic-resistant bacteria may carry the bacteria to their home country and possibly spread them further there. To reduce the flow of resistant bacteria into the home country, antibiotics should be used with caution in the treatment of traveller’s diarrhoea.”
Antibiotic-resistant strains can now be quickly identified through genetic engineering techniques, particularly polymerase chain reaction (PCR). Sequencing can be used for even more accurate analysis. The rapid identification of the infection-causing bacteria prior to drug selection is one way to control the use of antibiotics.
One of the studies conducted under Kantele’s leadership demonstrated that 80% of the tourists to high-risk areas who fell ill with diarrhoea and took antibiotics brought the ESBL super bacteria with them. ESBL (Extended Spectrum Beta-lactamases) is a special enzyme that breaks down antibiotics and makes the bacteria resistant to many common antibiotics.
The diarrhoea-causing ETEC coliform bacteria may also have the ESBL characteristic.
The premise of Anu Kantele is that we can learn from travel. What interests her is how the microbial activity of human intestines changes while travelling.
“I would like to find out what happens in the intestines of the 800 test subjects during the trip. We are talking about an ecosystem where the strongest bacteria win. It is exciting to combine the data on the changes in the microflora to how the body responds to them, what genes are activated, etc. We learn more about the intestines every day.”

Diarrhoea-causing bacteria can be detected in a stool sample through cultivation or PCR examination, or a combination thereof. PCR, or polymerase chain reaction, is one of the most important techniques used in molecular biology. It can be used, for example, to amplify a single gene or any segment of DNA multiple times. PCR is performed outside of living cells in a laboratory (in vitro) using a special PCR device. With PCR, a very small amount of DNA can be amplified to produce a billion times the amount of the same DNA in a few hours.
PCR technology is used for many purposes, including finding hereditary diseases, identifying individuals using genetic fingerprints, diagnosing infectious diseases and cloning genes. Microbial DNA is isolated from stool samples and amplified. By amplifying different gene areas, it is possible to quickly and efficiently identify pathogens from the stool sample. Microbes are identified by the base pair sequence. Cultivation is necessary alongside PCR because it allows the detection of antibiotic sensitivity.
The majority of the microbes in humans are located in the intestines. More than a thousand different species of bacteria live in the intestines of an adult human. The microflora in the intestines has up to a hundred times more genes than the human genome. 99% of intestinal bacteria are anaerobic, meaning that they grow in the absence of oxygen. Of the remainder, the most common are E. coli bacteria.
The intestinal microbes form their own ecosystem. Microbes have traditionally been studied and cultivated in laboratories. Now, with metagenomics, it is possible to also study them better in their natural habitat, be it soil or the intestines. DNA sequencing is used to try and ascertain the genome of an entire ecosystem. The human genome includes 20,000 genes. However, in addition to these genes, the intestinal bacteria of a single human being encode up to a million genes that affect the regulation of bodily functions. Almost 10 million genes originating from various bacteria have been identified in samples from human intestines. There is great genetic diversity and the amount of data is enormous. Knowledge of the functions of the genes of the intestinal microflora is still in its infancy.
The metagenomics service of EMBL-EBI is an automated data transfer service (EBI Metagenomics Pipeline) for the analysis and archiving of metagenomic data. There are samples from the human digestive system, soil, water, animals and plants. The data to be studied can be submitted to the service for analysis and comparison.
The service can be used to gain additional information on the evolutionary history of different microbial species as well as the functioning and metabolism of microbes. The data archived by EMBL-EBI is publicly available. EMBL-EBI is part of the ELIXIR infrastructure.
https://www.ebi.ac.uk/metagenomics/
Ari Turunen
7.5.2018
Article in PDF
Citation
Ari Turunen, Anu Kantele, & Tommi Nyrönen. (2018). Secret of the intestines. https://doi.org/10.5281/zenodo.8112378
More information:
EMBL-EBI
https://www.ebi.ac.uk/metagenomics/
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

The goal of Professor Mikko Niemi is to devise an interpretation algorithm that helps doctors determine the appropriate drug and correct dosage for a patient. Treatments become more effective and side effects are reduced, thereby decreasing the costs.
People react differently to medications; the efficacy of drug treatment remains insufficient for some, while others suffer from adverse effects. The reason for the atypical responses may be our physical characteristics, other medication and each person’s genetic makeup. An algorithm could be used to help predict the necessary dose or adverse effects of a drug when data on the patient’s genome is also available in addition to physiological information from the patient. A genetic test can be performed through a simple blood sample.
New information about the human genome is obtained all the time. At the same time, the costs of genetic research and bioinformatics have fallen significantly. Data is accumulated and there are many new opportunities for utilising it. Pharmacogenetics is the study of the effect of genes on the efficacy and safety of drug ingredients. If the data on patient genomes was available to doctors, medication costs and significant adverse effects would often be reduced. The number of days in hospital care would also decrease.
“If the genomes of patients were tested systematically, drug treatments could be better tailored and their dosages measured more individually”, says Professor of Pharmacogenetics and Chief Physician Mikko Niemi.
Niemi is leading a research group at the University of Helsinki studying how genes affect the concentrations, safety and efficacy of drug ingredients. He is also investigating when genetic tests should be considered in drug selection.
“The information on the results of the genetic test should be available when a medication is prescribed, but generally you have to wait a week or two for the result. It could, therefore, be sensible to proactively test for the most important genetic variants affecting drug treatments. Through our research, we seek to identify those patients who would benefit the most from such proactive testing.”
Niemi’s research group is also developing decision-making support systems related to pharmacogenetics. The aim is to devise an interpretation algorithm for doctors treating patients with cardiovascular disease to help find the most effective and safe cholesterol medication for each patient. The algorithm uses data on the patient’s characteristics, illnesses, other medications and genome.
Statin drugs intended for cardiovascular disease reduce the level of LDL cholesterol and increase the level of good HDL cholesterol in the blood. However, they cause muscle pain in some patients. The predisposition for muscle symptoms is partly hereditary.

The dosage requirement of individual drug ingredients may vary by more than tenfold between different individuals. This may result from how rapidly or slowly the drug leaves the body. Cytochrome enzymes (CYP) are central to the breakdown and removal from the body of many foreign substances, such as drugs. CYP enzymes are present especially in the liver.
When Mikko Niemi was working on his doctoral dissertation on the synergistic effects of diabetes drugs, he suspected that the variation in drug metabolism in different individuals was hereditary. Of particular interest are the three CYP enzymes CYP2D6, CYP2C9 and CYP2C19, as they affect up to one third of all drug ingredients in clinical use. Genetic variation in the activity of the CYP enzymes is high. This variation may lead to manifold differences in the concentrations of different drug ingredients and the responses to them in different individuals.

Genetic tests allow people to be classified into up to four different groups, depending on the drug, based on how quickly the body eliminates certain drug ingredients: very fast, normal, slowed down and slow. This so-called metabolic rate can affect the dosage requirement, efficacy and adverse effect risk of a drug.
In very fast metabolisers, the drug ingredient leaves the body faster than normal and its effect can be insufficient. In slow metabolisers, the drug exits slower than normal and its effects may be intensified. Consequently, the same drug dose may be too low for some and too high for others.
Some drugs become active by means of CYP enzymes. With such drugs, the effect of the hereditary metabolic rate is reversed. For example, in one third of the population, the effect of clopidogrel, a drug that inhibits blood coagulation, is weaker than normal due to hereditarily slowed down CYP2C19 metabolism. It is, therefore, advisable to opt for alternative medication with such patients.
Variation in the CYP2D6 enzyme, in turn, has a significant effect on, for example, codeine. Codeine is a common prescription painkiller, part of which usually turns into morphine in the liver via the CYP2D6 enzyme. In slow metabolisers, the effect of codeine may be inadequate. In very fast metabolisers, the amount of morphine in the body may run too high.
“Were the doctor to already know at the start of treatment that the patient’s CYP2D6 metabolism is slow, the patient would not need to suffer from inadequate pain management.”
Other enzymes besides CYPs are also relevant. TPMT, for example, is an enzyme that affects the metabolism of thiopurine drugs. Thiopurines are used to treat, for instance, autoimmune diseases, inflammatory bowel diseases and leukaemia.
“A hereditary TPMT deficiency predisposes you to the severe adverse effects of thiopurine drugs on blood cells. A genetic test to identify this hereditary deficiency has been in clinical use in Finland already since 2005”, says Mikko Niemi.
Around a dozen genetic tests related to drug treatments are currently available in Finland.
The suitability of a drug ingredient for each individual depends on many factors. It is not solely affected by enzymes that break down drugs. The transport proteins of the cell membrane affect the delivery of drug ingredients to their site of action. In the target tissue, the drug ingredient interacts with its target of effect.
“This results in a chain of events that brings about the desired drug effect. There are individual, partly hereditary differences in all these factors. It would be important to consider all these individual factors, including the genome, when selecting medication.”
In 2017, Mikko Niemi was granted substantial funding by the European Research Council for a project to develop an algorithm facilitating the selection of cholesterol medication. For this purpose, Niemi’s research group is building a so-called system pharmacological model.
“It is a kind of virtual patient that can be used to individually predict the effects of each alternative cholesterol drug.”
No similar algorithm has been attempted to date.
“If the algorithm works in the selection of cholesterol medication, a similar way of thinking could also be extended to other drug treatments.”
Of course, the algorithm cannot be built if there is not enough reliable research data available. Niemi’s research group has been compiling such data for years in their research projects. The biobanks established in Finland and the future genome centre will also speed up the collection of data needed for such research.
Better utilisation of genetic information is also desired by the Finnish state. Due to Finland’s exceptional settlement history, the genetic structure of the population provides special opportunities to combine genomic and health data. Pharmacogenetics is one of the four leading projects of the national genome strategy. The goal of the strategy is to have genetic data in efficient, health-promoting use already in 2020.
At present, the number of genes with significant effects on the efficacy and safety of drug treatment is relatively low: less than 20 of the total of about 20,000 human genes.
Since the group of genes is so small, according to Mikko Niemi, it would be technically possible to test even large numbers of patients.
“The next step is to proactively test for all genetic variants affecting drug treatment.”
The National Institute for Health and Welfare (THL), HUSLAB’s Department of Clinical Pharmacology and CSC have launched a pilot project that will be implemented by combining the genetic data of THL Biobank and the patient document information of HUS. The materials will be used to map the prevalence of the genetic variants affecting drug treatments in Finns. In addition, the project will look at how many patients in the sample receive drug treatment during or after the treatment period wherein genetic data could have affected the selection or dosage.
For the study, HUS and THL will have their own private and secure network connections to CSC’s data centre. This will allow HUS and THL to process data quickly and efficiently.
Sufficient long-term storage and data transfer at a speed of at least 10 Gbit/s to the systems of HUS and THL are prepared for in the project, and the necessary number of virtual servers for processing information is provided for the pharmacogenetics software environment.
Ari Turunen
4.4.2018
Read article in PDF
Citation
Ari Turunen, Mikko Niemi, & Tommi Nyrönen. (2018). Algorithm determines the appropriate drug. https://doi.org/10.5281/zenodo.8082229
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

More than a million tissue samples and tens of thousands of blood samples are stored in Auria, the first hospital biobank in Finland. The biobank is also able to combine donor-related data to the collections, providing significant assistance for research. The data can be requested from the donor of the sample, patient records or national registers.
An electronic health record accessed with your identity number has been in use in Finland for a long time. Registers requiring an identity number create good conditions for the efficient future utilisation of sample collections from people and the data linked to them. This is a great advantage over many countries.
The sample collections of Auria Biobank, which operates in connection with the Turku University Hospital and the University of Turku, are physically located in hospitals in southwest and west Finland. Samples are collected and combined with the necessary metadata, indicating the clinical information on the sample donor, quantity, date and how the sample has been processed. The samples of Auria Biobank include tissue, blood and DNA isolated from cells.
The Finnish legislation relating to biobanks is progressive. Consent provided once from the donor of the samples is sufficient for the stored samples to be utilised in various studies and in the future too. The law allows the biobank to contact the sample donors who have given their consent, for example, to enquire about the willingness of the donor to participate in a study not covered by the consent or to donate additional samples.
“In most cases, the contact has to do with drug research. If the patient is interested, they will contact the author of the study directly, and then they will form a separate agreement with the research organisation, after which the matter no longer involves the biobank”, says Perttu Terho, Vice Director of Auria Biobank.
The Personal Data Act and the Biobank Act are complied with in the transfer of data, safeguarding the privacy and confidentiality of patient information.
Consent for the donation of samples can be given in hospitals or online through an electronic form.

New samples are collected from consenting patients in connection with normal diagnostics and treatment. Tissue samples filed in hospitals are scanned, digitised and transferred to databases. Before transfer to the biobank, personal data is removed from the samples and replaced with a code. This ensures the efficient protection of personal data.
Auria collects tissue samples taken in connection with surgery that are left over from a diagnostic procedure, such as cancer tissue, and biobank blood samples taken in connection with laboratory visits.
“After surgery, the tissue sample is taken to a pathologist for examination. Typically, the sample is cast into paraffin and cut into slices with a thickness of a few micrometres which are stained with the colours needed for diagnostic purposes. The pathologist examines the stained tissue sections to establish whether there is a tumour present in the sample, for example. If some of the sample remains, it can be utilised in biobank studies. The sample must not run out, so there should be enough for the hospital to use. Once this has been confirmed, the tissue sample can be used for other research”, says Terho.
Auria Biobank digitises the samples that are needed for research projects.
“The purpose of digitisation is that we can, for example, ask a pathologist to assess the samples and mark the spots where cancerous tissue is found and where there is healthy tissue. The pathologist can do this from anywhere on their own computer, and the samples themselves do not need to be transferred. The digitised images can also be analysed in an automated way using pattern recognition algorithms and methods based on artificial intelligence.”
Auria has previously isolated DNA only from those blood and tissue samples that were needed in projects. Now, DNA isolation is to be done from every blood sample stored.
“Isolating DNA from every sample enhances research. Samples are received and stored, but nothing is yet studied. The samples are left to wait for future research as it is not yet known what the samples may be needed for.”
DNA will be isolated from 16,000 blood samples this year. Going forward, more than 20,000 samples will be taken every year.
The blood sample is taken in conjunction with a normal diagnostic or clinical blood sample.
“We are talking about one extra 10 ml blood sample for the biobank. The blood plasma and white blood cells from the sample are placed in different tubes before being frozen.”
Perttu Terho emphasises that the donated sample is valuable when it can be combined with patient data.
“Researchers may want data on patients with a specific diagnosis, medication and blood count. In this case, it is possible to quickly check the biobank and see whether there are samples that meet these criteria and the associated data exist.”
Biobank material can be used to identify the special characteristics of diseases and drugs. For example, it is possible to learn more about why some patients have side effects from medication and others do not.
“It is important to collect a sensible amount of relevant patient data from as large a number of people as possible. This allows samples from patients who are of interest to research to be obtained for the biobank.”
Sample-related requests from researchers are received every week.
“Based on an analysis, we map the quantity of samples and data in the biobank that the researcher is interested in. If the researcher is satisfied with the outcome of the pre-analysis, they will submit a request for data and samples describing the study and defining the required samples and data.”
The requests for data and samples are processed by the biobank’s Scientific Steering Committee which convenes once a month. The steering committee evaluates the requests. If the steering committee decides in favour of the study, the applicant can proceed to the preparation of a Material Transfer Agreement.
In principle, the operations of the biobanks operating in connection with Finnish hospitals are the same. They collect samples from their own hospital districts and store associated data. It would, of course, be a tempting idea to be able to search all the available sample collections in one go. The challenge is that, over the years, the different hospitals have stored and classified the samples in different ways. Different systems have different information registered, meaning that there is variation in the data provided on patient samples. Data should flow smoothly between the different biobanks.
“Hospital data is difficult to analyse. The expertise of a clinician is required to interpret what has been recorded. The data available is not directly commensurable. It would be important to create an availability service that is able to combine the data of the different biobanks so that at least the basic data would be available.”
The Finnish Biobank Cooperative was established in 2017. Its members include hospital districts and universities with faculties of medicine. The purpose of the biobank cooperative is to provide the material in the sample and the data collections of Finnish biobanks to be used by researchers under a one-stop principle. It would provide customers with a unified view and a centralised channel to the materials of Finnish biobanks. The biobank cooperative is responsible for the development of information systems, among other things.
According to Terho, it is possible to combine the associated clinical data relevant to research to the samples. Biobanks will utilise the sensitivive data platforms developed by CSC – IT Center for Science when designing the information services of their own.

Auria Biobank is involved in the establishment of the future genome centre. According to Lila Kallio, Acting Director of Auria Biobank, the way in which the transfer and storage of research and diagnostic sequences will be organised is, so far, only at the consideration stage.
“Genome legislation is being drafted and reform of the Biobank Act is underway. In addition, the new data protection regulation of the EU will also clarify the operations of biobanks.”
According to preliminary plans, the Finnish Genome Center will start its operations in 2019.
Ari Turunen
19.3.2018
Read article in PDF
Citation
Ari Turunen, Tommi Nyrönen, Perttu Terho, & Lila Kallio. (2018). Bank of million patient samples. https://doi.org/10.5281/zenodo.8081169
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

With the development of the methods used in bioinformatics, also the costs have lowered. It has become faster and cheaper to find out the genome of various organisms. However, we have a formidable task to be done to understand the information contained in the genome of various organisms and humans. It will require cooperation between various research organisations and well organised databases.
The mapping of the whole human genome was completed in 2003. Owing to the Internet, the Human Genome Project was completed earlier than anticipated, since it enabled efficient cooperation between various laboratories. The entire human DNA was sequenced. The human genes have been packed into three billion base pairs. Now, the next step is to find out how these genes work. Through the analysis of the base pairs of the genome we will begin to understand the pathogenetic mechanism of various illnesses and effective forms of treatment.
Today, research is generating quite versatile genome data. The aim is, for example, to use the information to evaluate the status of the environment and effects on health by analysing microbes, to cultivate edible plants into plants that will better withstand draught to alleviate the crises caused by climate change, or to develop drugs against diseases for which there is no cure at the moment. To do this, new kind of linking and analysis of the sources of data will be needed.

It is becoming faster and cheaper to find out the genome of various organisms. Now, as part of the Earth Bio-Genome Project (EBP), the aim is to map the genome of all eukaryotic organisms. Eukaryotic archaea and eubacteria, i.e. prokaryotes, are cells the DNA of which is constituted of only one chromosome. The group of eukaryotes consists of unicellular protozoans and three groups of multicellular organisms: plants, fungi and animals.
By means of bioinformatics, we can map the remaining 80 to 90 per cent of those organisms whose genome still remains unknown. In 2011, Census of Marine Life estimated the number of animal species to be approximately 8.7 million, 6.5 million of which are terrestrial and 2.2 million are marine animals. According to the estimates based on high-performance sequencing methods, there may be as many as 5.1 million species of fungi. There are approximately 400,000 plant species.
For the first time in human history, we will have the opportunity to efficiently sequence the genome of all known eukaryotic organisms. EPB’s aim is to sequence all of the known 1.5 million eukaryotes. Samples are being gathered all around the world. Part of them, probably around half a million, will be derived from botanical gardens. The rest will need to be directly collected from the nature. One of the most significant collection sites is the Amazon. In January 2018, EPB launched cooperation with a Brazilian gene bank project which concentrates on the organisms of the Amazon area.
The Amazon area has a richer variety of plant and animal species than anywhere else in the world. Probably one third of all species are found there. Rain forests are the home of a huge potential of new drugs.
For example, ACE inhibitor, i.e. the angiotensin-converting enzyme, was discovered from the venom of the jacaraca viper in the Amazon. The enzyme generates angiotensin, which helps lower blood pressure and lighten the pumping of the heart. In the 1970s, researches developed a synthetic version of the venom of this snake.

The oceans are the largest continuous ecosystem in the world. The significance of planktons for the global climate is at least as important as that of the rain forests. However, only a fraction of those organisms which create this ecosystem, have been classified and analysed. The ecosystems constituted by planktons contain a vast amount of life: in every litre of ocean water there are more than 10 million organisms, containing viruses, prokaryotes, unicellular eukaryotes and cnidarians. These genuine organisms contain bioactive compounds which can be used in the pharmaceutical industry, foods, cosmetics, bioenergy and nanotechnology. In 2009-2013, the researchers of Tara Oceans, an international expedition, collected 35,000 biological samples in 210 different measurement locations from oceans around the world. This is the largest plankton collection until this day. Another campaign in which samples were collected from the sea, was Ocean Sampling Day. In that campaign, research stations were asked to collect samples and to generate data. BioSamples collects descriptions and metadata from biological samples that have been used in research. The samples are references or have been used in various databases.

Analysing genomes and the proteins that determine their operation is a huge task, which would not be possible without cooperation. The European life science infrastructure for biological information ELIXIR provides an efficient platform for cooperation with members from nearly 200 research organisations, and an infrastructure which is used by almost half a million researchers. ELIXIR enables access to various data archives.
Massive sequencing of cultivated plants and forest vegetation allows us to do research on what is causing plant diseases. EURISCO (European Search Catalogue for Plant Genetic Resources) contains information on 1.9 million cultivated plants and their wild cousins. The samples have been collected by nearly 400 different organisations. A total of 43 countries are involved, and the aim is to preserve the agrobiological diversity of the world.
Uniprot (Universal Protein Resource) is collecting protein sequences and annotation data. An annotation means the determination of the functioning of the protein on the basis of the sequence. Owing to Uniprot’s data, we can learn more about the functioning of proteins and their interaction with other molecules as well as their location in cells and organisms. The aim is to collect all publicly available protein sequence data. Uniprot is the largest publicly available protein sequence database.
The European Nucleotide Archive ENA is a collection which offers free access to all published nucleotide sequences and annotated DNA and RNA sequences. The International Nucleotide Sequence Database is a collaboration forum between DNA Data Bank of Japan (Japan), GenBank (USA) and ENA. New data is synchronised between these three databases every day. Already in 2012, these databases contained the entire genomes of 5,682 organisms. The amount of data is doubled every ten months.
The European Genome Archive EGA is one of the largest public data storages in the world with patient data from biomedical projects. EGA stores the genotype and phenotype data collected from humans on the basis of a separate consent for research use of the sample and the data. Thanks to EGA, many of the ELIXIR research projects have become possible.
The ELIXIR infrastructure has more than 20 member states. Biomedical data is offered for use by researchers through the national centres in the member states. The benefits are indisputable. The genes of dogs and cats have proven useful in the analysis of rare human diseases. Through the Finnish centre the researchers have had access to a DNA bank of dogs and cats, the data of which has allowed us to discover the gene of a nerve degeneration disease, for example. The aim is now to develop a drug for the disease. Canine genes have proven useful in the research of human diseases, because the canine and human genomes are 95 per cent identical. The canine gene bank contains more than 70,000 samples from 60,000 dogs and 300 breeds of dogs. It is probably the largest of its kind in the world.
According to the estimates, by 2025 we will be able to sequence 100 million to two billion human genomes. To receive the best benefit from the data, genotype data should be linked to other health data. ELIXIR will be able to do this. The research infrastructure consists of nearly 200 organisations which form a federation, a network of trusted parties, which enables secure processing of human data. By 2016, with the help of the ELIXIR infrastructure, 21,000 scientific articles had been drawn up and 8,500 patents had been granted. The patents had been applied for vaccines, biomarkers, enzymes and prevention of the Ebola virus.
The order of magnitude of a single atom of a living biological molecules is one tenth of a nanometre. Should one carbon atom of that biomolecule be of human size, it would mean that its functioning would have a crucial impact on events that take place dozens of millions of kilometres away.
The diameter of our solar system is of the same order of magnitude.
If only one carbon in the biological molecule is replaced with another atom, say nitrogen, it could serve as a decisive property for the drug, whether it works or not, for example. Just this particular atom could be the one with which the drug molecule is making an attempt to attach to a protein, but fails to get a strong enough grip on it as a result of this change.
The protein, which the drug was supposed to influence, again, forwards orders to other proteins in our cells. If influencing the order is left undone, influencing on the biological message chain is left undone, too.
We could also ask whether all parts of the message chain located in the cell are flawless. All these factors will have an impact on whether researchers will be able to design a drug molecule correctly so that it can help the cells heal.
Unlike in space, there is no vacuum in a cell. The cells are full of constantly interacting biomolecules.
Our chances to have an impact on the fusion reaction of the sun, for example, are much more limited than the impact of the atom level digital information stored in the living molecules on people falling ill, even though the difference in the order of magnitude is the same.
Tommi Nyrönen
Ari Turunen
20.2.2018
Article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2018). Mapping the genomes of all organisms enables the development of new vaccines and medicines. https://doi.org/10.5281/zenodo.8070219
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

To start with, personal medical data is private and strictly protected. However, progress cannot be made in medicine without human data. The solution is a data management software program that provides security and only grants access to authorised material.
Data on the human genome should be treated with the utmost care and complying with information security protocols. In order to ensure information security, ELIXIR provides a service in which researchers log into a system that identifies their electronic identity while also distributing access rights to the biomedical data stored in the cloud. In this way, the researcher creates a secure analysis environment for the data to be analysed. This is made possible by the REMS tool.
ELIXIR strictly adheres to the EU law on information security. When researchers utilise data, the REMS tool can be used to ensure that the shared data is subject to authorisation.
CSC, the Finnish ELIXIR node, develops and maintains the open source REMS tool that can be used to manage access to datasets containing confidential material. REMS (Resource Entitlement Management System) is an access management tool that, where necessary, prevents the illegal use of data. With the REMS tool, it is possible to order a specific file from a large amount of data and have it delivered to the ordering party locked in a secure manner.
“There may be various tools within an organisation handling similar things. Although there are many ready-made tools and services available for identity and role management, I have not heard of any other general resource entitlement software like REMS”, says the REMS tool’s product owner Tommi Jalkanen from CSC.
REMS is part of a federated system formed by the ELIXIR community comprising nearly 200 organisations. Becoming a federation has required agreements between the different organisations regarding information security, personal data law, rights and obligations. This has resulted in ELIXIR’s own trust network, ELIXIR AAI, the rules of which each member organisation has committed themselves to follow.
In practice, ELIXIR AAI is a community that uses federated authentication and identity management. This federation has been developed based on the trust network of Finnish universities and research institutes (HAKA). The ELIXIR federation enables Single Sign-On (SSO) to joint services.
ELIXIR’s member organisations maintain basic user information that shows the role of the user in addition to the name and contact details. Determining the role is important because the REMS tool distributes access rights based on it. That is to say, REMS decides what kind of a view opens for the user in the service on the basis of personal details. This is so-called entitlement-based REMS.
Despite the high level of information security, REMS is still easy to use. No separate sign-on is required to use the tool. Logging in to the service is done with the user name and password of the ELIXIR home organisation. So no service-specific user name/password pair is required. It is this federated management that ensures the use of data resources can be monitored. At the same time, it is possible to ensure that the materials are not used for wrongful purposes. The use of the service can be monitored and reported.
The way the service works is that a researcher applies for permission to use the data with the REMS tool. The researcher logs in to REMS with their federated identity and then fills in an application for data use and agrees to comply with the terms of use. ELIXIR’s Data Access Committee (DAC) receives the application through REMS and approves or prohibits the use of the data. The applicant is notified of this by e-mail. If approval is granted, the applicant is provided with instructions on what happens next. REMS directs the data request to CSC’s Data Access Service. It provides the researcher with a view of the entitled data in the ePouta cloud service.
A federated user ID can be easily closed by the responsible organisation if the user switches workplaces, for example. The use of strong identification facilitates traceability and reporting. Fumbling with user name/password pairs is also reduced, as are password resets. Single sign-on reduces the need for separate user IDs and saves time, effort and money. Overlapping data management is reduced and data quality is improved. The service owner can focus on the service as the data administration of the ELIXIR organisation manages the IDs. These new practices support, for example, the use of ELIXIR’s many software services.

A new feature of the REMS software is a programming interface support for utility programs. A modern and widely-used web technology that enables the joint use of services, such as databases, is now available for researchers. This makes it possible to easily and safely build ecosystems and grant third-party access to the service. REST (Representational State Transfer) is a well-known and frequently used application architecture for decentralised systems. The REST interface allows different software programs from different platforms to use the same resource.
“Creating an all-encompassing interface is currently in the works, providing extensive opportunities for the building of third-party utilities”, says Tommi Jalkanen.
Using statistical methods, it is possible to identify a person with sufficient probability from anonymised material if genomic information is available on the subject. Therefore, this issue must be approached through information security, the usage agreements of the service providing genomic data as well as national and international legislation.
If the anonymised material is linked with additional information, such as year of birth or the name of the disease, the researcher must be reliably authenticated in the service and accept the service’s terms of use, which prohibit the identification of the persons included in the materials. It is also possible to profile users, in which case each profile can be provided with an appropriate view of the material. The access rights and legislation define how the materials should be, for example, stored and analysed.
Ari Turunen
7.2.2018
Article in PDF
Citation
Ari Turunen, Tommi Jalkanen, & Tommi Nyrönen. (2018). Ordered and secured. https://doi.org/10.5281/zenodo.8070212
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

The data obtained from the human genome will become part of health care decision-making. Combining a patient’s genomic data with the information available on the current state of the patient’s health enables the development of new algorithms, making it possible for a doctor to quickly select the best possible treatment and medication for the patient.
Medications have different effects due to the individual nature of a person’s genome. For example, some antibiotics cause drug allergies. The body may break down the medication before it has time to take effect, or the patient may experience harmful side effects. That is why utilising genomic data in pharmacotherapy will reduce the number of incorrect prescriptions. On the other hand, if a person knows that he/she has a digestion-related genetic trait that augments or weakens, for example, the breaking down of caffeine into energy and building materials, that knowledge may have a positive effect on his/her lifestyle. In the future, the algorithms of genetic databases linked to electronic patient record systems could automatically warn against possible adverse drug reactions and provide advice on the most effective alternative.
In Finland, CSC – IT Center for Science, the National Institute for Health and Welfare (THL) and the Institute for Molecular Medicine of the University of Helsinki are creating a secure framework for storing the genomic data produced on Finns and interpreting the data for health care purposes. The aim of the Helsinki University Hospital (HUS), which is involved in the cooperation, is to investigate the benefits of digital health data on humans for research and care. The six-month pilot project is part of an assignment given to the Genome Center to be established in Finland, coordinated by the Ministry of Social Affairs and Health.
Storing data becomes cheaper and capacity grows year after year. An exemplary file on the data collected on the health of Finnish people is the FINRISK cohort of THL. The analyses of the data collected for decades on Finns have been developed further in the GeneRISK project studying the hereditary risk factors for cardiovascular diseases. An algorithm that calculates the risk points for an individual to suffer from cardiovascular diseases is tested at the same time. A tool called Cardio Compass provides people with their current risk level and the development of the risk over the next few decades.
Cardio Compass is tested in practice by recruiting and testing 10,000 people from the Kotka region, the customer base of Mehiläinen and blood donors in Helsinki. The people participating in the project receive important feedback on their own health, more accurately than ever before, through the combination of genomic data. The information is collected in Cardio Compass. The participants may also talk directly with experts on the interpretations made based on the data.
In April 2016, the Finnish Government decided to establish a Genome Center in Finland with the aim of introducing genomic data as part of health care. In order to build the functions of the Genome Center, the data already collected and stored from the Finnish population will be utilised and combined in research which, if successful, will improve the accuracy of prescriptions. It would be possible to determine suitable medications or rule out the poor ones based on the patient’s genomic data. Algorithms can be developed to select a suitable drug ingredient and to optimise the amount of medication with standardised software methods. This is called pharmacogenetics.
In 2016, Professor Mikko Niemi from Biomedicum Helsinki was granted substantial funding by the European Research Council for a project to develop an algorithm for finding a suitable cholesterol drug for a patient. The mathematical model takes into account the patient’s genome, other medication, gender, age and weight.
However, effective utilisation of algorithms requires that there is enough different data available on patients. It is important to know the quality and purpose of the data. Sufficient metadata describes the quality of the data, based on which decisions on the utilisation of the data can be made. The interpretation of the data will become easier once a functional technical distribution platform is provided for reference data, making it possible to design better interpretation algorithms for the data.
Creating interpretation algorithms for genetic data for clinical use is the long-term goal. In addition to algorithms helping doctors to, for example, determine the appropriate medication, they can even be suitable for predicting changes in the function of proteins. The goal is that once the interpretation algorithms are ready for clinical use, they would be available in patient information systems automatically instead of as a request for information to be ordered separately.
Most of the technologies exist; we just have to be able to connect them. Expertise is attracted to Finland, for example, as part of European cooperation. The Finnish node of the ELIXIR infrastructure, which operates in connection with CSC, is building the secure infrastructure necessary for the management and storage of genomic data.
In the project, information technology is applied to the sample and data files of THL’s biobank. The aim of the project is to adapt genomic data so that it can be used by Finnish doctors and researchers in the best way possible. The full genome of about 9,000 Finns (www.sisuproject.fi) has already been determined through the digitalisation of this THL resource and other important Finnish sample collections, but the genomic data of up to half a million Finns has been discussed.
The project brings together the technological expertise of THL, HUS and CSC in Finland. The future goal is that this type of data would be analysed by a large group of Finnish bio-industry experts from universities, the public sector and companies in the bio-industry. Just storing the data is not enough; a service that covers the utilisation of all biological data must be created. At present, the expertise of the parties storing and providing the data is not sufficient for all possible health applications. Thus, the implementation of the pilot will provide important guidelines on how the efficient storage and secure distribution of genomic data can be carried out in cooperation between organisations so that the data can be fully utilised in health care, research and future innovations.
The question is largely whether Finland wants a specialised infrastructure the size of a small factory and expertise on the management and further processing of common genomic data based on which the data interpretation system would be built or if we want to outsource the data infrastructure services elsewhere.
In many countries, genomic data covering the entire country is a challenging goal. The services of the Finnish Genome Center are taking shape, and they will be created in cooperation with the parties managing the data, such as biobanks and licensing authorities. The data resources coordinated by the Genome Center are securely available for utilisation. In the future, all Finns could thus have their own health and welfare profiles that would include the data on their own genome.
Tommi Nyrönen
Ari Turunen
2.11.2017
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2017). Striving for a national service to utilise genomic data in health care. https://doi.org/10.5281/zenodo.8070200
More information:
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 20 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Plant growth and physiology are analysed with imaging methods, generating enormous amounts of data on the genomic and environmental response of plants. The aim of this is to improve the productivity of crops, allowing food and raw materials to be produced for the growing human race in an ecologically sustainable manner.
In NaPPI, a joint infrastructure of the universities of Helsinki and Eastern Finland, plants are measured and analysed automatically. The operation of the infrastructure and the data produced by it can be organised from the outset so that it is also compatible for the use of other European research organisations. This is a good goal because, until now, every laboratory around the world has collected data on the genome, phenotypes and environmental factors of plants in their own way.
The Viikki Plant Science Center (ViPS) of the University of Helsinki is a research cluster with 36 groups studying plants. The research topics range from adaptation to a particular habitat and climate change to plant stress tolerance and plant breeding.
The activities of NaPPI (National Plant Phenotyping Infrastructure) focus on plant research and breeding. The aim is to produce comprehensive phenotypic data from a large number of plants. NaPPI provides the technical possibilities to combine the information on plant genomes to phenotypic data.
The phenotype of a plant is jointly produced by genes and the environment. The phenotype can take a very different shape due to the impact of the environment. Plants have a much wider capacity for non-hereditary variation than animals. Plant growth, for example, can be effectively influenced in various ways, including nutrients and light.
People have been cultivating plants for thousands of years due to a desire for better food. This has been done locally, and the information collected on plants has not been recorded systematically. A good example is the numerous varieties of grape, with more than a thousand in Europe alone. The origin of all the varieties is no longer known and that is why the origin is being investigated through genetic engineering.
“The data on plant phenotypes has not yet been standardised. Various research groups have been producing and categorising it in their own laboratories”, says Kristiina Himanen, Research Coordinator of the NaPPI infrastructure from the University of Helsinki.

The aim of the NaPPI infrastructure is to enhance and specify the collection and analysis of the data from plants with new imaging technologies. The infrastructure uses imaging devices that analyse plant growth and physiology. The plants are measured and images are taken of them automatically, after which the computer calculates the height, width and, for example, the surface area and shape of the rosette based on the images.
“The size, growth and form of a plant, i.e. plant architecture, are important in agricultural production,” Himanen emphasises.
“Plant architecture can affect the yield or cultivation characteristics. As dwarf varieties of rice have been produced, they do not become lodged as easily anymore, and this affects the harvest. Genes can influence plant architecture and hence the quantity and quality of the harvest.”
What happens when a dwarf gene is fed into the genome of turnip rape is being studied in Viikki. Tarja Niemelä, PhD (Agriculture and Forestry), and partners are investigating whether the dwarf gene can increase the productivity of turnip rape by reducing the biomass of the stem in relation to the seed yield produced by the plant.
“There is a huge amount of genomic data available, but you have to be able to combine it with other data. We want to link the phenotypic data that we produce with imaging devices to genomic data. Ultimately, of course, we are interested in how the information obtained from genomes and phenotypes can be transferred to plant breeding.”
According to Himanen, the volume of plant research will increase thanks to new imaging methods.

In addition to plant forms, the NaPPI infrastructure equipment is also used to analyse the physiological state of plants. The Spectromics Laboratory located at the Joensuu Campus of the University of Eastern Finland is the first research environment in Finland that focuses on the spectral imaging of plants and other biological samples. Spectral imaging consists of images taken at different wavelengths of light with their own colour channels. The Spectromics Laboratory is developing optical methods especially for the study of plant stress responses.
The human eye or a conventional camera sees colours as combinations of three wavelength bands (red, green and blue). With a spectral camera, however, it is possible to detect up to hundreds of different wavelength bands. It is also not limited to visible light, but is capable of taking images in the ultraviolet and infrared ranges. A separate image may be formed of each band and each pixel contains a complete spectrum.
“Spectral imaging enables very precise separation of colours, but it also multiplies the amount of data produced”, says Professor Markku Keinänen from the University of Eastern Finland.
“This, in turn, requires complex computational approaches in image analysis. So spectral imaging is, to a large extent, computation, and the images illustrating the results are not produced until the final stages of the analysis.”
When plants are analysed with thermal and fluorescence cameras, you can see things that are not visible in ordinary light. Fluorescence is visible light of a certain colour that is generated when the atoms of a plant are excited due to, for example, invisible ultraviolet radiation. Thermal and fluorescence cameras can be used to calculate, one pixel at a time, the size of an area of a different colour in the plant and to study, for example, infections in the plant.
The Finnish ELIXIR node offers efficient capacity for the processing and storage of data. Since the data collection of phenotypes has been automated and digitalised, according to Kristiina Himanen, it is now possible to also start the standardisation of data.
“Data must have the same format. The Excelerate project is developing standards for phenotypic data and metadata. There are 22 countries involved. Although everyone has their own infrastructures, their operations are now being harmonised.”
In practice, researchers have access to information about the plant’s genome and phenotypic data on growth conditions and other environmental factors. Once both data sources have been combined, it becomes possible to create comprehensive databases and the laboratories across Europe can avoid doing redundant work and divide data collection in a sensible way.
“The introduction of a single gene in plant breeding will become easier because the amount of work involved in the analysis of a single plant will become more reasonable.”
Going forward, the Viikki research groups will produce image-based data to which genomic data is linked. The Finnish ELIXIR node, in turn, is figuring out how to analyse and standardise the data and how to hand over the metadata to ELIXIR for a cloud database. The division of labour between the NaPPI infrastructure and the Finnish ELIXIR node CSC is a good example of how genotype and phenotype data on plants should be produced for research.
Ari Turunen
11.8.2017
Read article in PDF
Citation
Ari Turunen, Kristiina Himanen, Markku Keinänen, & Tommi Nyrönen. (2017). Better harvests on the horizon? Data will also be harvested. https://doi.org/10.5281/zenodo.8070177
More information:
NaPPI
NaPPI is part of a cooperation network with the Spectromics Unit of the University of Eastern Finland (www.spectromics.org) and several other Finnish plant research institutes. Partners from the universities of Turku and Oulu and the Natural Resources Institute Finland are also involved in the cooperation.
Viikki Plant Science Center
https://www.helsinki.fi/en/researchgroups/viikki-plant-science-centre/about-vips
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 20 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org
In NaPPI, a joint infrastructure of the universities of Helsinki and Eastern Finland, plants are measured and analysed automatically. The operation of the infrastructure and the data produced by it can be organised from the outset so that it is also compatible for the use of other European research organisations. This is a good goal because, until now, every laboratory around the world has collected data on the genome, phenotypes and environmental factors of plants in their own way.
Read the rest of the article here: https://www.elixir-finland.org/en/better-harvests-on-the-horizon-data-will-also-be-collected-in-the-future/
Utilising genomic data in pharmacotherapy will reduce the number of incorrect prescriptions. On the other hand, if a person knows that he/she has a digestion-related genetic trait that augments or weakens, for example, the breaking down of caffeine into energy and building materials, that knowledge may have a positive effect on his/her lifestyle. In the future, the algorithms of genetic databases linked to electronic patient record systems could automatically warn against possible adverse drug reactions and provide advice on the most effective alternative.
Read rest of the article here: https://www.elixir-finland.org/en/striving-for-a-national-service-to-utilise-genomic-data-in-health-care/

Genetic research has revealed that there are a lot more microbes with much more diverse communities than we even knew about. Studying the genetics of microbial communities gave rise to a new branch of life sciences, metagenomics. Jenni Hultman studies the significance of the microflora in Arctic regions on climate change.
Microbes, or microorganisms, are what single-celled organisms or life forms consisting of a few cells are commonly referred to as. These include bacteria, protozoa, viruses and unicellular algae. Although microbes exist everywhere in our environment and even in extreme conditions, their genetic origin and function remain poorly understood. The vast majority of microbes are unknown.
The term metagenome references the idea that a collection of genes picked up and sequenced from the environment could be analysed in a way analogous to the study of the genome of a single species. With metagenomics, it is possible to investigate changes in microflora during the course of various diseases and, after treatment, find new pathogens and obtain information about their function during medication, for example. Metagenomics can also be used to study how microbes affect our environment.

In metagenomics, DNA is isolated from the microbial community. This has been relatively easy when microbes have been studied in the intestines and bodies of water, for example.
The examination of soil is considerably more challenging due to the large number of microbes in a single sample. One sample may include up to 10,000 different species. Since new technologies allow the DNA of different microbes to be isolated from the soil, microbial research is under constant change. New information about organisms as well as the origin of life on Earth is obtained all the time. However, microbial communities are challenging as research subjects. Microbial diversity is very high and microbes also affect each other in ways that are not well known yet.
“Microbes have traditionally been grown in petri dishes. But we are now talking about a huge number to be studied because the research subjects are microbial communities where the different microbes are dependent on other microbes or nutrients. Such communities cannot be grown in dishes. Now, the aim is to sequence the majority of the genes in the soil sample. Even if you find out what the species is, it is also important to know what the genes do. Since up to millions of genes are sequenced from a microbial community, this requires computing capacity”, says Academy Research Fellow Jenni Hultman.
Hultman is particularly interested in the microflora of Arctic regions. As microbes act as decomposers in nature, they may play a significant role in the formation of greenhouse gases, such as carbon dioxide and methane. In the short term, the impact of methane on the greenhouse effect is several dozen times that of carbon dioxide.
“The microbes in the Arctic environment are not well known. They can have an effect on how the climate and conditions change. There are many questions. How is nature adapting to climate change? What do species do when the climate changes?”
The melting of peat bogs under the permafrost especially generates methane emissions. But what is the significance of microbes in this process? That is what Hultman wants to find out.
Hultman, who works at the Department of Food and Environmental Sciences of the University of Helsinki, collects research data on microbes in different parts of the Northern hemisphere. In her research, Hultman analyses soil samples in Kilpisjärvi, Alaska and Greenland. She is now looking for a survey site in Siberia, after which the samples she has collected would well represent the entire Northern hemisphere.
“20% of the Earth’s land surface is covered by permafrost. Within the permafrost are huge stores of carbon dioxide. The melting of permafrost may release the highest amounts of carbon dioxide ever measured into the atmosphere. This process is dependent on a microbial response but, at present, we know rather little about the activity of microbes under permafrost.”

Hultman is interested in the activity of microbial communities and especially in what the genes of the microbial communities do (metagenomics) and how active the genes of the communities are at a given time (metatranscriptomics).
Hultman isolates the total DNA and RNA from the soil samples of the field area in Kilpisjärvi, divides them into smaller pieces and sequences them. She isolates the DNA and RNA from samples of 0.5 grams. The number of sampling points is over a hundred. The area has a microclimate, allowing Hultman to take into account various factors, such as humidity, pH and temperature. This makes it possible to study the significance of the activity of microbial communities on climate change on the scale of “mini climate change.”
“A high number of parallel samples weighing half a gram are needed because the microflora of the soil is diverse and because the soil itself varies greatly. Microbes can be present in stone, a dead worm, the root of a plant or just in a place that is more humid than another. So there is a lot to dig up and isolate.”
The essential thing is to know what the genes of the microbes are actively doing and how they affect climate change.
“I am studying what is happening in the soil sample at this moment. Which genes are active? Are some microbes accelerating climate change and some slowing it down? Do microbes just produce methane or do they utilise it?”
One important goal of Hultman’s research is to produce data obtained from metagenomics also for climate models. This may potentially improve the reliability of climate models.
One gram of soil may contain up to ten billion different microbes. When microbial ecology research truly started in the late 1970s and microbial samples from the environment were compared with cultured microbial samples, it was found that the samples from the environment contained up to 99% more new and unknown microbes than the cultured samples.
Traditionally, the sequencing of genes is started by growing cells in a Petri dish. When DNA from the cells is placed in a DNA sequencer, it determines the order of the DNA base pairs: adenine, guanine, cytosine and thymine. However, early metagenomic studies revealed that there are large groups of microorganisms that cannot be grown in laboratories and that, therefore, cannot be sequenced.
The early studies focused on the sequences produced by the 16S rRNA gene. The function of the 16S rRNA gene, which is found in all living creatures, is to produce the ribosomes in which protein synthesis occurs. In 1977, microbiologist Carl Woese started the sequencing of this gene when studying microbes. Because the gene is always slightly different in different microbes, Woese noticed that it can be used to study the development history of the microflora in the samples. However, Woese and his colleague George E. Fox were surprised when many of the isolated 16S rRNA sequences did not belong to any known species. The discoveries made with the 16S rRNA gene revolutionised microbial research.
Woese and Fox observed that the samples also contained unicellular, but anucleate microorganisms that externally resembled bacteria but were not. They called this group the archaea.
Archaea are involved in metabolism and affect the functioning of enzymes. Archaea were initially observed only in extreme conditions, such as hot springs and salt lakes, but have since also been found in different soil types, marshlands, oceans and even human intestines, for example.
Organisms could thus be divided into three categories. Eukaryota, i.e. multicellular plants, fungi and animals, have nuclei. Bacteria and archaea, in turn, are anucleate microbes that make up most of the world’s biodiversity.
“As the sequencing of DNA is becoming cheaper all the time, metagenomics allows microbes to be studied on a much larger scale and in more detail than before”, Jenni Hultman says.
The mysterious archaea may play a greater role in the formation of methane than has previously been known. Some archaea break down organic carbon into methane. But how many of such archaea are there and how effective are they as decomposers?
The data on the secrets of the microbiome collected by Jenni Hultman and other researchers is stored in the public information resources maintained by ELIXIR, the European life sciences infrastructure for bioinformatics.
Ari Turunen
19.6.2017
Article in PDF
Citation
Ari Turunen, Tommi Nyrönen, & Jenni Hultman. (2017). Microbes and climate change. https://doi.org/10.5281/zenodo.8070142
More information:
Department of Food and Environmental Sciences
http://www.helsinki.fi/food-and-environment/
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 20 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Extensive research projects are being conducted on Finnish genetic heritage and genomic data is being produced and analysed all the time. However, the national objective is to store the data produced on Finnish people in Finland, allowing analysts to combine the data with other health information. The utilisation of genomic data in health care is still in its early stages. Data analysis offers many opportunities for companies in the bio-industry, including in Finland.
Research-appropriate genetic data on the Finnish population exists fragmented all over the world in various databases and data storages with varying arrangements. Therefore, there is a need to create a domestic, secure service for the management of Finnish data that would cross organisational boundaries, be network-based and well-coordinated. By coordinating the data in different locations in just one place, the data could, with the permission of the owner, be released for legitimate purposes, such as research, product development and medication.
The human biology is very complicated, more complicated than previously thought. The expression, structure and function of genes and the building blocks of the body, or proteins, require advanced mathematical, computer science and statistical methods, i.e. bioinformatics.
New ways to study and prevent diseases are constantly being discovered through bioinformatics methods, such as gene sequencing. DNA sequencing is the starting point where the order of the four bases – adenine, guanine, cytosine and thymine (A, G, C, T) – within a DNA molecule is determined when deciphering the genetic digital code. Each ACGT base is a similar nugget of information to a computer bit, zero or one, which, as a long chain, contains the instructions for a programme.
Sequencing methods have improved and become cheaper, and this has significantly increased the possibilities of biology and medicine to produce this kind of data. The data is now being used to find out what digital messages have been written on the molecules of life for the survival of organisms.
However, data is only the first step towards interpretation. The interpretation of digital genomic data, that is, how the information stored in the genome manifests itself in the body, is still under development. In the last ten years, for example, researchers in Sweden have been creating a map (HPA Human Protein Atlas) on how genes are expressed as proteins in different cells and then combining this information with microscope images of cells. This allows you to see which gene is expressed in any given cell and is involved in the development of proteins and, hence, larger structures, such as neural fibres, hair follicles or light-sensing structures in the fundus of the eye. However, a clear, deeper level map on how molecules that are operating on a nanometre scale produce these functional, microscopic structures does not exist yet. The structure of each cell requires millions of molecules in cooperation. The building instructions stored in genomes and the resulting molecules form a self-organising network that current research tries to understand.
Finland is fairly well positioned to be an international actor in the management of genomic data, but there are too few experts in individual organisations. The data masses required to understand genomic data are large and the analysis requires specialised expertise that does not sufficiently exist in Finland yet. There is a need for cooperation in genomic data management and for more interpreters specialising in data. Finland will gain more expertise once the creation of a framework for storing Finnish genomes is achieved. Initially, this would mean a national reference database created from the data of tens of thousands of people. It would be beneficial for diagnostics, such as in improving medical treatments, as it is already possible to determine, for example, suitable and safe medication based on the patient’s genomic data.
Analysing data from molecules, cells or whole organisms requires that the data is well organised. The data produced with sequencing, microscopes, mass spectrometry or computer simulations must have common file standards and sufficient machine-readable interfaces to be followed when the data is stored. A good indicator of the degree of data organisation is if another research group can utilise the data as well as its original producers.
When data is well organised and described, it can be combined. Combining supplementary information, such as a prescription, genome and long-term treatment results, is a prerequisite for developing a deeper understanding.
Data organised in the hands of skilled analysts will help achieve breakthroughs in research. The US company GRAIL, for example, seeks to understand the underlying mechanisms of cancer. The earlier the cancer is detected significantly improves the prognosis of the disease.
The GRAIL project has involved the collection of samples from 10,000 patients and their consent for the analysis of the diverse data created from the samples. The idea is to use the cancer tumours of this group of patients to create a database against which blood samples can be screened.
Cancer tumours are usually the result of a change in the genome of a cell of the person carrying the disease, making the cell abnormal. At the cellular level, each cancer is a rather unique disease that looks like its carrier; what they have in common is the reckless growth of abnormal cells. Cancer utilises the normal regeneration and healing mechanisms of the body to selfishly spread its own genetic instructions. The genomes and the digital information contained therein of two humans are, on average, 99.5% identical. That is why the progression process of many cancers is well-known despite the individual nature of cancers. Consequently, it is justified to study how changes in individual or multiple nucleotides (ACGT) in the genome affect the balance of the cell’s molecular network so that the cell becomes a cancer cell.
In the GRAIL project, millions of unique changes in genomic data that may cause cancer are sequenced from the genomes and cancer tumours of patients. The project will create a database that allows health care professionals to detect early stages of cancer, even directly from the bloodstream. If the innovation is successful, cancer screening can be started earlier, meaning that the tumours are still microscopically small and easier to manage through, for example, medication.
Conducting similar research in Finland is possible by combining health and genomic data. The Finnish ELIXIR node, for example, has already started building the secure infrastructure necessary for the management and storage of genomic data.

There are hundreds of times more data on the information contained in DNA available for science than ten years ago. Understanding of how the information stored in the genome is transmitted at the molecular level, for example, to proteins, and further to three-dimensional functional units of cells, is growing at a rapid pace. When human biology is understood from the cellular level to the molecular level, it improves quality of life and the treatment of diseases.
One of the most important research subjects in bioinformatics is understanding the underlying mechanisms of diseases. The functional unit encoded by a gene is a protein. It is a chain of hundreds of units, or amino acids. There are 20 different amino acids. The protein chain guided by genes becomes a functional unit of the cell, such an as enzyme, only after it has folded into its three-dimensional state and can start interacting with other molecules in the cell. An incorrectly folded protein can lead to illness because it does not function as expected in the network formed by molecules important to life.
Sometimes, for example, there is a change in the genetic code at a critical point for the folding of this critical functional unit, or protein. Cells self-modify the composition of the proteins created, and thereby their structure and function. This may correct the error in the genetic code. On the other hand, what may also happen is that the protein breaks down in the cell’s own process. Most diseases can be traced back to situations where a biochemical reading error has occurred in an important part of the dynamics of the cell’s molecular network. On the other hand, this may just be a variation that only results in dietary recommendations to the person in question. The effect of molecular level changes on the data stored in the genome depends on many things, as DNA includes a “backup” of each gene from both parents. There are even several versions of some genes.
Even though the logic and knowledge on the main players in the network of biological processes are pretty much accounted for, the dynamic entity cannot yet be understood, let alone predicted or modified medically, as well as desired. Predicting the risks of contracting coronary artery disease, for example, has become more accurate thanks to the data obtained from the genome, but the understanding of molecular level events is at a stage where the components are known but there is a struggle to understand their interaction or defects occurring at the molecular level. However, molecular level understanding of diseases means more accurate and earlier diagnoses, that preventative measures can be initiated early and that those at risk, for example, may choose to change their lifestyle.
Tommi Nyrönen
Ari Turunen
21.5.2017
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2017). Storing the whole genome of the Finnish population? The data will benefit disease research. https://doi.org/10.5281/zenodo.8070146
More information:
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 20 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

As more and more data is collected about the human genome and lifestyle, it is important that you look after your personal information and are aware of the overall security of your data. Is it reasonable to put this burden on an individual? Should it be transferred to a trusted broker?
How does one ensure that this trust is kept, but at the same time, enable others relevant parties to have access to your data? This brings up the question of who collects, interprets and uses this data and do these entities have the competence to do so? The exponential growth in biological information has an impact on both individuals and communities. It will become possible to predict a person’s entire lifespan with certain genetic premises and lifestyle factors. As this information is increasing, also the possibilities to use this data for purposes other than what it was originally intended for will increase. Do we dare consume unhealthy foods anymore in the future if information is collected about it that may impact our insurance terms, for instance.
Economic and societal impacts will be felt after the next five to ten years when bioinformatics will be applied in preventive health care. For example, if a person has a genetic disposition to fall ill as a result of liver disease, which can be treated by lifestyle changes, revealing the issue to them at an early stage will probably influence the person’s lifestyle choices. Health care professionals can justify their recommendations by presenting well-known examples of life-long treatment histories from the health care system or biobank.
Open questions still remain: How and to what extent is modern biological information interpreted and used in public health care? How will/should the legislation evolve? As the need for better health care for the aging increases, so does the cost, and therefore such issues must be clarified quickly. The legislative aspects is particularly important as many insurance companies and giants of data processing, such as Google, are interested in the opportunities that are opening up.
The American 23 & me provides genetic tests for anyone, which then provides information about hundreds of medical risks related to one’s own genome. There are already many illnesses that can be analysed at the molecular level so it is therefore possible to diagnose, for example, one’s propensity for cancer, which in turn can radicalise and tailor treatments to reduce side effects related to generic “heavy” treatments. It is envisaged that such new technologies will also be able to predict changes in the state of health of an individual.
Who can, who is allowed to, and who is able to participate in the continuous observation of one’s own health? Who interprets whether a person is drifting towards a serious illness, and can this diagnosis be trusted? Whose rare disease can be cured and should it be done using public resources? Which ethical boundary conditions are used to coordinate access to the latest treatments?
Technology provides increasing opportunities for observing health and lifestyle on an individual level in real time. Different kinds of technological devices for monitoring our own health are becoming cheaper and integrated into devices that we already carry with us – mobile phones, clothes or watches. The Finnish insurance company Lähitapiola is conducting a new experiment where the company provides “Intellectual Life Insurance”. The insurance company cooperates with Polar, who produces biomonitors and collects, for instance, heart rate and lifestyle data for the application that help doctors make predictions about the person’s state of health. It is possible for the client to lower their insurance fees if certain healthy lifestyle options are met in the data given to the insurance company. Individuals would therefore benefit from lower insurance payments that encourage a healthier lifestyle. To return the favor, the insurance company accepts data as a “currency” that it utilises.
This type of data is valuable. Reliable and well organised data sources that are used in interpreting an individual’s health are currency in international commerce. In the UK, the National Health Service NHS has decided to open up the health care history of more than one million Londoners to Google. They are hoping that access to the data would enable Google’s experts to help prevent kidney diseases that are the source of great costs in public health care. It is estimated that as many as every fourth of these cases could be prevented if the risks were detected earlier and the people would change their lifestyle. This would bring about considerable savings in the public sector and improve public health.
The data that people accumulate about themselves regarding their lifestyle, e.g. engagement in sports, food and alcohol consumption, currently ends up online in very different services, or is deleted within one year . The aim of the services collecting this data is usually to gain profit by, , ”encouraging people to be engaged with their technological ecosystem”. . Connecting this type of accumulated data with third-party data sources is usually not possible. Further, using this type of data to support reliable diagnosis requires access to vast studies, so that an individual’s data retrieved from the sample can be interpreted correctly. This kind of data integration is still in its early phases.
However, the development pace is fast. Examining data collected from dogs, for example, is legally less restrictive than data from humans, and there are many services combining genetics and lifestyle for advancing dogs’ health already available (MyDogDNA). The next great favour by man’s best friend may be showing how genetic biological information should be used in health care.
Health care organisations collect data and samples from people in connection with treatments for research purposes. A medical professional is always responsible for the confidential collection of data and samples. Permission from the collector is requested if these are used for new purposes.
The prevailing practice significantly facilitates conducting studies to improve health. In the Nordic countries, centralised health care has been in use for decades, which has also been able to organise and provide high-quality data for research purposes. For example, 30 percent of Norwegian citizens have a sample in the biobank. In Finland, more than 150 million medical histories have been collected for archives from 4.3 million citizens.
There are 5.4 million people in Finland, and in 2016 nearly all medicine prescriptions go to the same archive. The Biobank Act that came into force in Finland a few years ago also ensures that responsible research use of the data is allowed without informing every citizen about the issue separately. The collection provides an excellent starting point for interpreting the connections between genetic premises and factors that happen during a person’s life, if safe and sufficiently open access to the data can be created for a large group of international, skilled analysts.
But what can we read in the data now and, above all, in the future? In the UK, Google has been given access to all patient data because it is not possible to know in advance which factors predict and explain the development of a kidney disease. But what if, when trying to predict this, it turns out that the person has an acute risk to have a heart attack? Should the person be informed about this? Nordic biobanks have studied that approximately 60 percent of people want to know about random discoveries. The rest 40 percent do not want to know. Who owns the data and samples collected from people, and who has the right to control it for example for research purposes?
Support systems for decision-making related to health care are based on constructed and maintained data sources. In international cooperation, it is possible to build more reliable data sources in light of people’s genetic premises and lifestyle and health care history than any country on its own. For this reason, we should strive for globally accessible data sources also in terms of processing and interpreting biological information related to humans. International access to data increases democracy, because the costs related to the use of the interpretations of the research data collected about people can be shared. At the same time, it is possible to support countries that would not be able to create such data services on their own. It is possible to collect measurement data about people everywhere where the internet is available – the challenge is performing the interpretation of the measurements in a reliable manner. Internationally open and secure data services would be one solution for this.
For example, the human genome has approximately 20,000 genes that guide all functions of the body. Sometimes genetic information becomes corrupted, which can lead to, for example, breast cancer. An international research group has shown in the latest studies that there are exactly 93 genes in the human genome that can change a healthy cell into a breast cancer cell when they mutate. This kind of information is crucial when designing new medication, because the proteins generated by the mutated gene are the targets of the design of the medicine molecules. The person suffering from the illness can also be diagnosed in more detail with this information.
Restricting this type of biodata only to a certain group would be wrong. Therefore there is need for open services for biological information, so that the research results are available to universities, research institutes and the pharmaceutical industry when they are needed. One of these data services is the European Genome-Phenome Archive EGA, which is a part of the European ELIXIR research infrastructure. EGA protects biological information on the old continent. EGA stores vast amounts of biomedical data material about humans and distributes the data based on licensing. Universities, research institutes as well as companies and public administration can save data into this global European service. The service has been much used, for example, in Nordic public health care for publishing long time series and research data regarding the whole population (gene pool).
During the following decades, utilising data collected from humans will be a part of society. Enlightened citizens will know how to demand for new kinds of health services. The private services sector in the field may grow quickly. However, we need international data sources and standards on which the small and medium sectors can build on and that guarantee the quality of the interpretations of the measurement data. The correlations between data sources collecting information about genetic, molecular biological and lifestyle data has only just begun.
Tommi Nyrönen
Tommi Nyrönen is a biochemist and the Director of the Finnish unit of the European infrastructure for biological information ELIXIR. He works at the IT Center for Science CSC.
16.4.2017
Read article in PDF
Citation
Ari Turunen, & Tommi Nyrönen. (2017). “Smart life insurances” offered: human biological data is only useful when interpreted correctly. https://doi.org/10.5281/zenodo.8070130
More information:
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 17 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

The Bioinformatics Unit of BioCity Turku focuses on the analysis of gene and protein data. The data analyses are useful in understanding various disease mechanisms. Cancer diseases and type 1 diabetes in adults, in particular, have been studied in the unit. The goal of the unit is to improve the diagnostics, treatment and predictability of complex diseases by combining computational, experimental and clinical research.
Bioinformatics methods are used to analyse the three-dimensional structures of proteins. This makes it possible to figure out what kinds of partially developed drugs, typically small molecules, are likely to affect the protein. By utilising this information, researchers are able to understand the normal functioning of the cell and how the protein function should be affected. The end result may be a new drug molecule that affects the target protein as desired.
“An encounter between two molecules always results in interaction. Compatible form and chemistry greatly enhance this interaction. If the encounter is strong, it can change the molecule’s potential to affect a third molecule. A signal is thus transmitted in a chain with encounters between different molecules”, says University of Åbo Akademi researcher Jukka Lehtonen who specialises in information technology in the bio-industry.
Lehtonen emphasises, however, that the molecular pairs transmitting the message are not perfectly accurate, so it is not a straightforward messaging chain. Rather, we can talk about a network of molecular interactions.
“The so-called normal functioning of cells is a delicate state of equilibrium. Medication is used to try to maintain this normal state. With diabetes, for example, the insulin function of cells has been disrupted, so medication and diet are used to replace the reduced interactions.”
“Medication is also used to try and curb signal chains that function in a harmful manner.”
In the design of drug molecules, it is important that the chain of events functions in the desired manner in all molecules. If, for example, the third molecule in the signal chain activates excessively, the drug may not have the desired effects.
“The drug is effective and there are few side effects if the structures of the binding site between the drug molecule and the protein are sufficiently unique and compatible”, Lehtonen says.
“However, there are many proteins of the same type in the human body and even the most imprecise interactions can change the administered drug molecules chemically.”
Hence, there are two parts to drug design: designing the optimal molecules for the target protein and finding compounds that, when travelling through the body, change into drug molecules without side effects.
The three-dimensional structure of a protein can be determined through X-ray crystallography. The electrons in a regular protein crystal bend the X-rays and the bending, or diffraction, can be used to calculate an electron density map. The structural model is generated by matching the atoms of the protein with the density of the electrons using computational algorithms and computer graphics.
“The crystallisation of a protein is a difficult phase. Finding the right crystallisation conditions is challenging. Some proteins do not crystallise as a whole”, says Lehtonen.
However, the number of protein structures has increased tremendously. In 1994, 1,000 structures were determined; now the number is already 100,000. The protein structures that have already been resolved are available in the PDB database.
“A significantly higher number of proteins exist and, based on other research findings, there are several potential drug targets whose structure has not yet been determined.”
If the structures of the target protein’s relatives are known, an attempt can be made to prepare a homology model.
“Relatives usually resemble each other. A theoretical model of the target’s structure can be drawn up based on a known relative. The model will inevitably resemble the original”, Lehtonen says, but points out that the model is not a result, but rather a tool.
The structural model is used to explain the experimental data collected on the functioning of the protein and to predict what may happen in an abnormal situation. The model can be used to predict, for example, what kinds of interactions different small molecules can have with the protein.
“However, the model must be assessed critically. All of its parts are not equally reliable. A structural model may depict the binding site of the drug molecule credibly even if it is otherwise uncertain.”
Lehtonen emphasises that modelling requires cooperating with research groups conducting experiments.
“Experimental arrangements that tell more about the research subject, while also revealing whether the model is reliable, are suggested based on the model. The modeller must decide whether the model can be used based on the data. The model is corrected and specified using the experimental data obtained. The cycle continues until the target is known well”, Lehtonen says.
Drug design that is based on structure utilises information on the structure the protein’s binding site and known molecules that bind to the protein, called ligands. Drug molecules are often designed to resemble a ligand. At best, researchers have a defined protein structure that includes a ligand at their disposal. A protein can also be selectively mutated, in which case it can be deduced, based on the changes in binding intensity, which amino acid residues are involved in the binding. The binding site is usually a cavity in the protein structure. The cavities in the structural model can also be outlined computationally, but identifying the authentic binding site is not automatic.
“The mode of action of the ligand, i.e. the normal function of the protein, is in itself a valuable research result. If related structures, a group of ligands that bind to them and the differences in binding intensity are known, the most significant atomic level differences can be identified through structural analysis. This will reveal what is important in the structure of the ligand.”
The potential drug molecule should, therefore, have similar parts. If there is sufficient experimental data available on the binding site in the target protein, virtual screenings performed with databases and powerful computers can be used to quickly and reliably define the potential drug candidates from a large number of molecules. This also minimises the possible side effects of the drug.
“Virtual molecule libraries can be sifted through using the created search criteria, that is, by performing a computer search that excludes all of the completely unsuitable molecules. The remaining compounds are subjected to more specific modelling so that the group of compounds to be experimentally tested is reduced to a reasonable number.”
Modelling is used to find the molecules that are likely to react correctly with the protein, and the accuracy is tested through laboratory results. This provides an answer to what the potential drug candidates are and why they work and the others do not.
“If two molecular structures are placed side by side virtually, it can be asked how strong their interaction is. The strength of forces is affected by distances between atoms and the presence of other molecules, i.e. water. Physics and chemistry have produced the observation data and theories to assess forces. When the molecules move or are transformed, the calculated forces also change. The molecules can therefore be laid out in countless ways.”
Docking is a search algorithm that calculates the force between the protein and another molecule.
“Each docking algorithm uses a different strategy for group selection. The goal is to find the optimal layout which hopefully describes how the structures actually interact. The search is quite closely limited to the assumed binding site and the permissible transformations of the molecules are small. Otherwise the search space is too large, meaning that the amount of calculation increases disproportionally.”
In bioinformatics, docking is used to determine which ligand binds the most strongly. Once there is a model for the binding site, binding style and binding strength of each ligand, a proposal can be prepared on what the new drug molecules should look like in order for them to bind to the desired target protein. There are several different computing technologies available for docking. Molecular dynamics simulation, for example, permits the free movement of molecular pairs, and the work may not take weeks. That is why efficient computing resources are required. Molecular dynamics is a computationally heavy method for docking, but the reward is a more accurate understanding of the dynamic interaction between molecules. Molecular dynamics simulations are used for a more detailed modelling of the interactions and for evaluating the interaction and stability between the protein and a partially developed drug.
“The biggest mistake in modelling is to blindly believe the answers provided by the software. What is essential is the ability to evaluate the results critically and utilising modelling for problems for which it is suited”, Lehtonen emphasises.
The Turku Bioinformatics Unit uses the cloud computing resource ePouta of the Finnish ELIXIR node in its research. It creates a transparent, local resource whose level of information security is very high. The user does not see that the computing takes place in the cloud and the data does not need to be transferred from one disk drive to another, especially via the public network. The higher information security level of ePouta is essential for research materials involving corporate secrets, for example.
“Thanks to ePouta, we have more computing capacity in the local network, which has suited us very well. In practice, our computing capacity has doubled. At the national level, CSC’s cloud is the most affordable way to create local computing resources.”
According to Lehtonen, ePouta creates a transparent, local resource. The user does not see that the computing takes place in the cloud and the data does not need to be transferred from one disk drive to another, especially via the public network. The higher information security level of ePouta is essential for some research materials.
“Since CSC is responsible for the computing resources and cloud service, it is possible to build an environment where the researcher feels comfortable at the client end. Maintaining software packages that CSC does not have is also easier this way.”
Ari Turunen
15.1.2016
Read article in PDF
Citation
Ari Turunen, Tommi Nyrönen, & Jukka Lehtonen. (2016). New drug molecules through determining the structure of proteins. https://doi.org/10.5281/zenodo.8068984
More information:
Biocenter Finland
Biocenter Finland (BF) is a distributed national research infrastructure of five biocenters in six Finnish universities:
http://www.biocenter.fi/
BioCity Turku
BioCity Turku is an umbrella organization supporting and coordinating life science and molecular medicine related research in the University of Turku and Åbo Akademi University.
http://www.biocity.turku.fi/
Turku Centre for Biotechnology
Turku Centre for Biotechnology is a joint department of the University of Turku and Åbo Akademi University, providing high-end technologies and expertise to academic and industrial researchers. .
http://www.btk.fi
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 17 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

The BBMRI (Biobanking and Biomolecular Resources Research Infrastructure) is an infrastructure established by 16 European states. Its aim is to promote the high quality, research-based use of the sample collections and associated data of European biobanks. Use of such collections assists in the development of diagnostics and treatment, as well as health promotion and disease prevention. Finland has several biobanks in operation. A common IT infrastructure is being created for these based on cooperation between the BBMRI and ELIXIR.
The BBMRI operates through national centres that coordinate the biobanks of member states. Service centres serving the customers of biobanks are also being established in member countries and under the BBMRI. BBMRI.fi is a national cooperative body belonging to the BBMRI Network; its membership is made up of Finnish biobanks.
Five biobanks were operating in Finland in 2015. New ones will be set up in the near future. More than 100,000 Finnish sample collections were transferred to the National Institute for Health (THL) in June 2015. Sample collections can be used to identify the causes of diseases and the related impact of heredity,
the environment and lifestyle changes.
A total of 50 percent of samples from the Auria Biobank are cancer samples. The Auria Biobank focuses on cardiovascular, metabolic and cancer research, as well as research into neurological diseases. The biobank was established by the University of Turku and the University Hospital Districts of Southwest Finland, Satakunta and Vaasa.
The FHRB – the Finnish Hematology Registry and Biobank – operates throughout the country and collects blood and bone marrow samples from patients with haematological diseases. Such samples are required for research into methods of treating serious haematological diseases, particularly leukaemia. The FHRB biobank is owned by the Finnish Association of Haematology, the Institute for Molecular Medicine Finland (FIMM) and the Finnish Red Cross Blood Service. The Association of Finnish Cancer Patients is also involved in its activities.
The mission of the Academic Medical Center Helsinki (AMCH) is to support research aimed at health promotion and the understanding of disease mechanisms, as well as the development of products, diagnostic methods and treatment practices used in healthcare.
The HUB Biobank focuses on urological diseases and supports research in this field, based on biobank samples. The Biobank began sample collection at the beginning of 2015. Research based on samples and data is aimed at improving the prevention, diagnostics and treatment of urological diseases. The HUB Biobank was founded by FIMM and the Hospital District of Helsinki and Uusimaa (HUS).
Biobanks manage huge and important data sets. Tasks such as the alignment and management of genome data and imaging datasets are challenging. The aim is to create a national, web-based availability service for biobank data, based on which users can search for materials suitable for research and product development.
Juha Knuuttila, Enterprise IT Architect at THL, coordinates IT cooperation between biobanks in Finland. Knuuttila views the biobanks’ IT infrastructure as central to national cooperation within the BBMRI.fi network.
”In Finland, IT infrastructure is highly developed in comparison to many other European countries. BBMRI.fi and the ELIXIR Centre in Finland are good examples of this. Both have a clear role to play. ELIXIR provides a good cloud service while the BBMRI offers specialised IT systems in support of biobank activities. The virtualised computing clusters of FIMM and CSC – IT Center for Science are available via a
cloud service. Cooperation as smooth as this is still rare at European level,” states Knuuttila.
The aim of the biobanks cooperation network is to agree on uniform practices relating to quality criteria, and to organise nationally consistent data transfer practices. Consistent ethical principles and maintaining the confidence of the persons involved in research are another important area of national biobank activity.
In terms of IT, cooperation has been initiated through database pilots. For example, pathology archives form the key sample data of most Finnish hospital biobanks and biobank projects. A national digital pathology infrastructure has been created by digitalising pathology samples from university hospital archives, based on inter-biobank cooperation.
Digitalisation fosters the use of new applications such as DNA microchip technology and the development of tools for the analysis of large data sets, thereby promoting individualised health care. These services are part of the European BBMRI infrastructure.
”The goal is to create a unified Finnish interface connecting us to the European infrastructure.”
However, much work remains to be done. With health care systems of several types in use, information is fragmented. Knuuttila believes that the greatest task lies in the harmonisation of data.
”To facilitate research cooperation, clinical data, demographic data and sample data should be combined in one place and an easily searchable format. Biobanks should therefore agree on which variables can be combined in the databases in a realistic and useful manner.”
Knuuttila believes that this would force the biobanks to work together, which would also benefit researchers and pharmaceutical companies. Knuuttila is leading the biobanks’ joint IT group.
”Obtaining patient data in a structured format would be useful to both the biobanks and hospitals.”
THL, FIMM and CSC entered into Biomedinfra cooperation to meet the need for a joint organisation to exploit genetic data obtained from biobanks. This also required joint IT solutions. The project was sponsored by the Academy of Finland and Ministry of Education and Culture.
Four different services can currently be used through common interfaces. They comprise the sample and data register (National Institute for Health and Welfare, THL), the code, consent and event register (FIMM), the research access management service, i.e. REMS (CSC) and the availability database (FIMM).
”Each has a programming interface allowing them to exchange information.”
According to Knuuttila, no one else has created a programme like the one completed by the THL, CSC and FIMM.
”Open source solutions of this kind may also be attractive to biobanks.”
Knuuttila believes that the next key tasks are the creation of a common, webbased availability service through which users can explore the availability of samples and data belonging to different biobanks.
”This and next year, the focus will be on the creation of a joint availability service.”
Ari Turunen
26.10.2015
Read article in PDF
Citation
Ari Turunen, Juha Knuuttila, & Tommi Nyrönen. (2015). BBMRI.fi: an IT Infrastructure for shared biobanks. https://doi.org/10.5281/zenodo.8068915
More information:
The National Institute for Health and Welfare (THL)
The THL is a national centre of expertise that provides reliable information in support of decision-making and action in the health and welfare sectors.
https://www.thl.fi
http://www.bbmri.fi
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 17 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Extensive data sets and databases are increasingly being used in cancer research. The research group of Sampsa Hautaniemi, Professor of Systems Biology at the Faculty of Medicine of the University of Helsinki, develops methods that can be used to integrate data from various sources, such as DNA, gene expression and protein function. When the analysis results are combined with biomedical databases, it becomes possible to generate experimentally testable predictions. This is useful in the diagnostics and design of treatment methods, for example.
Sampsa Hautaniemi worked at the Massachusetts Institute of Technology (MIT) before setting up his own research group at the University of Helsinki in 2006. Hautaniemi’s laboratory analyses complex, disease-related biological systems using mathematical methods. The analysis of data masses is not possible without computational assistance.
“Biomedical research requires databases and computational methods, especially in the interpretation phase of the results”, says Hautaniemi.
The objective of the systems biology group, which operates at Biomedicum, is to apply computational methods to medical research questions. For example, which genetic profiles affect cancer risk or what is the prognosis of a patient with a particular genetic profile? The aim is to find a unique treatment in accordance with the genomic profile for the patient.
“Our goal is to understand the behaviour of the cancer cell and look for targets that, when their activity is modified, allow cancer cells to be destroyed with minimal side effects. When wanting to treat a cancer patient, you must first understand how the tumour cells make decisions on how they grow, multiply and move. We pursue this through genome-wide measurement and mathematical methods.”
In the treatment of breast cancer, for example, it is important to be able to predict the probability of metastases emerging. Even though the treatment prognosis for breast cancer is improving all the time, metastases greatly increase the risk of disease.
“The problem is that we do not know how and which cells detach from the tumour, where they go and how they function there.”
The aim is to deduce who has a high probability of forming metastases by studying gene activity and combining data. Current measurement methods, such as microchips and new generation sequencers, generate enormous amounts of data.
“At this time, we do not yet know the main internal cell factors that affect the treatment response of cancer. That is why we use methods from different levels that measure the whole genome in research.”
In addition to DNA and RNA sequencing, such methods include, for example, epigenetics, or analysing the impact of lifestyle on gene function. Proteomics, which determines the function of proteins and their structure, is also important.
More than four billion observation points can be measured from one cancer tumour. From this mass of observations, you should be able to identify the most characteristic factors for cancer development and drug response.
According to Hautaniemi, there has been quite a change compared to the situation 10–20 years ago when the usual number of observations to be processed was a few dozen or hundred.
“In addition, databases have genome-wide data available on thousands of cancer patients. Utilising this data alongside Finnish material is important, but challenging.”
In addition to prognosis, Hautaniemi’s group also looks for suitable treatment methods based on computational analysis. Hautaniemi’s group is mapping, for example, the impact of genetic modifications on drug response. Cytostatics, which destroy cancer cells, are used in the treatment of cancer. It is important to find a suitable cytostatic because the patient does not always respond well to the given drug.
In cooperation with the group of Professor Olli Carpén, Hautaniemi’s laboratory has used genome-wide data on hundreds of ovarian cancer patients in their research. The researchers have been looking for subgroups of patients that have developed a resistance to conventional chemotherapy in which platinum derivatives and taxoids are used as cytostatics.
The research project uses hundreds of thousands of processor hours of supercomputer computing time and dozens of terabytes of storage capacity.
“For a person with a certain type of genetic profile, some medications may even be harmful, while others provide the optimal benefit.”
Hautaniemi and his group have developed methods by using data related to lymphoma together with the group of Professor Sirpa Leppä. The challenge is how to convert the data collected from genes and proteins into knowledge. Observations from clinical samples are always rather noisy and multidimensional, meaning that there are thousands of genes, proteins and potentially interesting areas of DNA. Therefore, it is essential to answer the correct and necessary medical questions so that the results are useful. The research questions can then be solved by mathematical methods.
When analysing lymphoma and ovarian cancer data, Hautaniemi’s group used the so-called deep sequencing method. The method involves DNA or RNA being divided and sequenced, after which the base sequence of the molecules is converted into a format understood by a computer. There may be hundreds of millions of short sequences converted into a computer format. According to Hautaniemi, when converting medical data into knowledge, the most significant bottleneck that is faced is the comprehension of medical questions so that they can be modified into computational problems.
To solve this problem, Hautaniemi and his group have developed a software program called GROK (Genomic Region Operation Kit). It allows questions to be converted into computational problems and solved based on the data. GROK is a universal tool and it has been used to understand the progression of prostate cancer. The study was conducted in cooperation with the laboratory of Professor Olli Jänne. The cooperation resulted in a better understanding of the function of the FoxA1 protein with the AR protein, which is the main protein affecting prostate cancer. Furthermore, the study found that a large number of FoxA1 proteins provide a poor prognosis and a small number provides a good prognosis. In future, the results can be used to prepare a treatment prognosis and for planning treatment. According to Hautaniemi, the methods developed can be applied to any kind of cancer.
“We have used the methods we have developed to study, for example, breast, prostate and ovarian cancers. Although the tumours are found in different organs, they have a significant number of similarities at the molecular level. Therefore, in future, it might be possible to use a breast cancer drug for certain subtypes of ovarian cancer, for example. Prior to this, it must be possible to characterise the subtypes of each cancer. This means that, in future, we will be able to reliably find similar cancers regardless of their location and then recommend effective medication suitable for them.”
Hautaniemi believes that cancer cell sequencing will be part of routine cancer diagnostics in future.
“We are striving to find the factors for each tumour type and individual tumour, and it is only a matter of time before we understand the biology of tumours so well that we can quickly calculate a prognosis and combinations of drugs that are likely to be effective based on their genome. Computational sciences play a key role in achieving this and utilising technology.”
These days, the amount of data produced by life science experiments doubles every few months, and the amount is still growing. The experiments also produce a completely new kind of data. The accumulation of huge amounts of data from research has created a need to systematically manage all that information. The objective of ELIXIR is to harmonise data storage, processing and analysis.
In many respects, databases are starting to be vital for life science research, but they have often been maintained alongside other research activities and dependent on fixed-term research funding. One of the main objectives of ELIXIR is to secure the funding of the most important databases containing biological research data. When the system compiling and distributing information is permanent, research groups can build their own operations on it. The ELIXIR infrastructure also provides a system and funding pathway for services developed in Finland that are significant for the whole of Europe. Everyone does not have to produce the same database on their own; instead, data that has been created once can be used effectively in multiple locations, and tasks can be shared.
“The field of bioinformatics is so vast that no single laboratory can provide all services. What the Finnish and ESFRI project infrastructures bring with them are a certain clarity and an improved flow of information. We know what is being done and planned elsewhere”, Hautaniemi says.
Ari Turunen
24.9.2015
Read article in PDF
Citation
Ari Turunen, Sampsa Hautaniemi, & Tommi Nyrönen. (2015). Fighting cancer with mathematics. https://doi.org/10.5281/zenodo.8068867
More information:
Genomic region operation kit
http://csbi.ltdk.helsinki.fi/grok/
Ovaska, Lyly, Sahu, Jänne, Hautaniemi (2013): Genomic region operation kit for flexible processing of deep sequencing data
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

The research groups of Jukka Jernvall and Petri Auvinen at the Institute of Biotechnology are investigating the genomes of different species and the structures of populations. The objective is to understand when the species arose and diverged from one another. The groups are particularly interested in the Saimaa ringed seal, whose full genome will be determined.
The Saimaa ringed seal is an excellent research subject for the study of genetic diversity, isolation and inbreeding. The Saimaa ringed seal has not been in contact with other seal species in more than ten thousand years. Its eyes, brain and skull are different from those of other types of ringed seal. The Saimaa ringed seal developed from a seal population that probably came from the Baltic Sea to Lake Ladoga before moving to the Saimaa archipelago.
“If a Ladoga ringed seal was transferred to Lake Saimaa, it might not survive. The Saimaa ringed seal has adapted to the murky waters containing humus and the maze-like archipelago”, says Petri Auvinen, Laboratory Director of the Institute of Biotechnology.
The DNA Sequencing and Genomics Laboratory of the Institute of Biotechnology specialises in gene sequencing, or determining the order of base pairs in DNA. The laboratory has sequenced the entire genome of several organisms, starting from the Lactococcus piscium bacteria that spoil cold food. Gene expression is also studied at the laboratory through sequencing. Key events in the evolution of organisms include cellular division and differentiation, which are highly temporally and spatially regulated.
Cellular differentiation takes place in stages. Sometimes a gene is switched on and sometimes it stops functioning. This active functioning is called gene expression. When gene expression can be measured, it is possible, for example, to monitor which genes start to function when a tree prepares for winter. The EST (Expressed Sequence Tag) technology provides information on the location and function of a gene. By identifying the base pair sequence of genes, a tag can be provided for each expressed gene. Currently, the RNA-Seq method is mainly used to study gene function.
The aim of the researchers of the Institute of Biotechnology at the University of Helsinki is to obtain a reference genome of the highest quality possible from the Saimaa ringed seal. A reference genome is a digital sequence database on the full base pair sequence of a single species, compiled from one individual in the case of the Saimaa ringed seal and from numerous genomes in the case of humans. Collecting a good reference genome requires the use of various advanced technologies.
The reference genome and deviations in individual genomes enable the efficient study of the population. In the STR (Short Tandem Repeat) method, a specific locus on DNA where a few base pairs in a row are always repeated is compared with two or more DNA samples. The DNAs of individuals are clearly distinguished with STR. Mitochondrial DNA, in turn, can be used to trace the maternal lineage of individuals back thousands of years. The rapid development of DNA sequencing technologies has enabled the identification of single-nucleotide polymorphisms (SNP), providing a very accurate estimate on the differences between individuals. This method is also used in the Saimaa ringed seal genome project. The data collection requires a lot of storage space and computing power, provided by CSC – IT Center for Science via the ELIXIR infrastructure.
The genome of the Saimaa ringed seal is 2.5 billion base pairs in length, the same as the canine genome. In determining the genome of the Saimaa ringed seal, the group of Academy Professor Jukka Jernvall focuses on studying the teeth of seals while the group of Petri Auvinen focuses on population history and genome structure. Once the genome has been determined, the genome of the Saimaa ringed seal will be compared with the genomes of the ringed seals in Lake Ladoga, the Baltic Sea and the Arctic Ocean.
The researchers are collecting data on the connections between the genotype (genetic factors) and phenotype (environmental factors) together with researchers from the universities of Oulu and Eastern Finland. A lot of data on developmental biology is obtained by analysing teeth. Once a tooth erupts, it will no longer develop or change due to the environment. However, there is huge variation in teeth. That is why it is studied as to which genes have affected unusual teeth. The teeth of the crabeater seal, for example, have become very polymorphic due to evolution and function like the baleen of whales because the seals eat krill.
“We have computer models of all ringed seal skulls. We can create accurate phenotypes and look for the probable genes that caused a particular tooth. Gene function can be modelled on a computer and analyse which areas of the genome could affect the tooth.”
A different skull or teeth indicate adaptation or specification, adjustment to different conditions. Because the orbits of the Saimaa ringed seal are different from those of other, even closely related ringed seals, it can be concluded that, for example, it has adapted to murky and maze-like waters.
The groups of Auvinen and Jernvall have access to the DNA of the only known hybrid between a ringed seal and a grey seal in the world. In 1929, Skansen Zoo was the birthplace of a cub from whose tooth Auvinen managed to isolate DNA. The offspring of a huge grey seal and a small ringed seal only lived for a short time. The teeth and skull of the hybrid indicate an intermediate form. According to Auvinen, it would probably be the equivalent of a hybrid between a chimpanzee and a human. It is now possible to compare why a specific kind of tooth or skull develops.
Auvinen considers this significant research also for human evolution because it is not known when modern humans differentiated into their own species. Hybrids have also occurred during human evolution. There have been findings of human skull fragments that are a cross between Cro-Magnon and Neanderthal. 2–5% of Europeans carry genes passed down from Neanderthals. Furthermore, the skeleton of a human subspecies named the Denisovan was found in the Denisova Cave in Siberia. It became extinct 40,000 years ago, earlier than its cousin, the Neanderthal. When DNA was isolated from the finger of the Denisovan’s skeleton, it was found that Tibetans have Denisovan genes. One hereditary gene helps Tibetans survive in a high altitude climate.
The researchers of the Institute of Biotechnology want to find out whether the Saimaa ringed seal is its own species or a subspecies. The researchers know exactly for how many generations the ringed seal has been isolated in Lake Saimaa. The population of the Saimaa ringed seal is small. There were only 140 individuals left in the 1980s, now the number is 320. By comparing the samples from Lake Saimaa, the Baltic Sea and Lake Ladoga to the reference genome of the Saimaa ringed seal, it is possible to study what kind of a population has passed through a so-called bottleneck.
Nowadays, there are also computational methods that make it possible to determine reasonably accurately, even from a single genome, the kind of a population its ancestors have lived in. The bottleneck phenomenon faced by a population refers to an event where a large part of the population is destroyed or only a small number of individuals establish a new group, such as the people who once arrived in Finland. The reason behind the destruction may be changes in the environment or a transition to a new environment, which can prevent reproduction.
Studying the genetic history of the Saimaa ringed seal is also helpful for human genome research. Bottlenecks can increase inbreeding and thus also affect the disease heritage. In Finland, bottlenecks have given rise to about forty hereditary diseases in the population that are much common here than anywhere else. Finnish genetic bottlenecks have included the adoption of agriculture 4,000 years ago and the spread of settlements to the northern and eastern Finland in the 16th century.
“It is now possible to examine the impact of disease genes on population structure and the bottlenecks caused by nature and humans. The Finnish disease heritage is interesting in this respect. It can be determined what the disease heritage carried by Finns was like when they went through a bottleneck”, says Auvinen.
Creating a reference genome comes with many benefits. The reference genome data can always be reused. The better the reference genome is, the easier it is to analyse new data that can be compared to the data of the reference genome.
For example, analysing the reference genome of birch accelerates and enhances wood research for the needs of industry and medicine. New properties that affect the quality and quantity of wood can be looked for in the birch genome. This data can also be utilised in research on other wood species.
“Unlike birch, it will take 10 years to determine the properties of, for example, poplar and eucalyptus. Birch can be genetically modified. Since birch can be made to bloom up to three times a year, new properties can be introduced to birch in one to two years. These techniques can also be applied to other wood species. The genetic model of birch can be used, for example, in the study of eucalyptus”, says Petri Auvinen.
The birch reference genome project was also followed by industry representatives. Thanks to genetic data, birch properties can be refined and the forest industry can use the wood for purposes other than timber.
New applications include nanomaterials, wood processing industry side streams and, for example, hemicellulose. Auvinen also mentions the betulin in birch bark that has been reported to have anti-cancer and even antiviral effects. Betulin has already been used to create medicinal creams. Striving to produce birches with more betulinic acid using conventional breeding methods is also a possibility.
Ari Turunen
10.8.2015
Read article in PDF
Citation
Ari Turunen, Petri Auvinen, & Tommi Nyrönen. (2015). Saimaa ringed seal aids the study of population genomes. https://doi.org/10.5281/zenodo.8068837
More information:
Institute of Biotechnology
Institute of Biotechnology is an independent research unit belonging to the Helsinki Institute of Life Science (HiLIFE) at the University of Helsinki
http://www.biocenter.helsinki.fi/bi/dnagen/index.htm
CSC – IT Centre for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 17 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

An invention of the Finnish doctors Johan and Mikael Lundin provides an effective solution for the analysis and storage of tissue section images.
The volume of research data is increasing enormously year after year, requiring a continuously active approach from software developers. It must be possible to analyse large amounts of data with software that does not jam the workstation. Johan Lundin, Research Director at the Institute for Molecular Medicine Finland (FIMM), studies and develops image-based diagnostics using machine vision solutions. In future, it will become possible to produce individualised disease prognoses and treatments by combining various data sources, genetic data, tissue data and clinical patient data. This has been applied especially in the treatment of breast, prostate and colon cancer.
When he was working at the Helsinki University Hospital in the early 2000s, Lundin became frustrated with how difficult it was to process large tissue section images at the workstations. The size of tissue section images is 1–2 gigabytes, so storing them on your own hard drives does not make sense. Rotating the images is also slow. With his brother Mikael, Johan Lundin started thinking about a functional software solution for the problem.
The brothers developed a fully web-based software program, the essential components of which are an efficient image server and a web interface that works with all browsers. With their compression algorithm, images take up less space and load quickly. A two gigabyte sample image can be compressed to the size of half a gigabyte. The tissue sample is stored in the cloud and large amounts of data can be processed quickly and easily from your own workstation.
The online microscope service can be used with all browsers and tablets, including smartphones. WebMicroscope® is also compatible with the image formats of different microscope manufacturers. WebMicroscope enables the study of very extensive materials and is also ideal for collaborative projects as a joint management and analysis space for digitised images.
“There has been a growing interest in the service. Doctors, researchers and teachers are shifting to digital microscopy. An online cloud-based service is a progressive solution for the users of digital microscopy all over the world”, says Kaisa Helminen, CEO of the service’s provider, Fimmic. Helminen is a trained biochemist and has previously worked for several companies in the bio-industry.
Fimmic was established in 2013 and the commercialisation of the service started the following year. Fimmic’s customers include universities, research institutes, pharmaceutical companies and companies conducting external quality control. External quality control is enhanced when samples can be sent for analysis virtually instead of mailing samples on glass slides to laboratories.
Fimmic uses the cPouta cloud service of CSC’s data centre as a partner in providing these services. It offers the WebMicroscope users their own server, a high-speed bandwidth and a massive amount of storage space. This ensures that the service works as efficiently as possible. WebMicroscope is also suitable for biobanks for tissue sample management. The service can be customised to suit a particular biobank.
Microscope scanners are expensive devices with the price typically varying between 150,000 and 300,000 euros. However, the number of scanners is increasing and, when images are scanned, the most convenient and least expensive solution for many users is to store them directly in the cloud.
“If a customer does not have the opportunity to use a scanner, the samples can be sent to us for scanning. We will store the digitised samples directly in the customer’s WebMicroscope account”, Helminen says.
Through the WebMicroscope portal, users can share their own microscope images with different research groups and partners around the world. This is an important feature because in drug design, for example, the rapid sharing of test results between research groups and pharmaceutical companies is a prerequisite for breakthroughs. Research related to drug development is one of Fimmic’s focuses.
With a traditional microscope, only a small portion of a sample can be examined at one time. A microscope scanner takes a picture of the sample with a large objective, digitising the entire sample in detail. With WebMicroscope, the resulting image can be viewed easily and quickly, regardless of location.
“You can select a section of the tissue sample to be examined, similarly to Google Maps, and only look at a part of it, quickly moving to another spot. The image is not saved on the workstations, but is rather loaded over the network directly from the image server.”
All Finnish universities teaching medicine use WebMicroscope for educational purposes at anatomy and pathology courses. WebMicroscope allows digitised samples to be easily shared with students along with other documents and videos.
You can secure your own pages with a password and the software can also be used to complete exams. The virtual samples can be viewed using a tablet or a smartphone in distance education, or on a large screen in the classroom. The application is ideal for multi-touch screens that utilise multiple points of contact. Massive tissue section images can then be viewed easily and quickly on a large touch screen even with a larger group.
A microscope scanner produces a lot of data. There may be millions of observation points to examine, the processing of which requires computing power and good algorithms. Fimmic plans to further develop the software and introduce quantitative image analysis tools, algorithms. According to Kaisa Helminen, the number of potential research subjects that algorithms can be used for is huge.
“Machine vision algorithms are based on signal processing. With dozens or even hundreds of images, the machine is taught to identify a particular signal from the background, for example stained cells from other tissue. Screening is case-specific and it varies how different samples have been processed. An algorithm is just as good as it has been taught to be.”
All of this requires computing power obtained, for example, from the supercomputers of CSC – IT Center for Science.
“A lot of computing power is required because the images being studied are so-called whole slide images. Smaller sections of these may, of course, be selected for analysis, but a lot of computing power is still required so that the analysis would not take too much time”, Kaisa Helminen notes.
Ari Turunen
1.8.205
Read article in PDF
Citation
Ari Turunen, Kaisa Helminen, & Tommi Nyrönen. (2015). Webmicroscope stores tissue samples in the cloud. https://doi.org/10.5281/zenodo.8068745
More information:
Fimmic
CSC – IT Center for Science
CSC – The Finnish IT Center For Science is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centralised IT infrastructure.
http://www.csc.fi
https://research.csc.fi/cloud-computing
ELIXIR
ELIXIR builds infrastructure in support of the biological sector. It brings together the leading organisations of 17 European countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish
centre within this infrastructure.
https://www.elixir-finland.org
http://www.elixir-europe.org

Would you have thought that the beloved tail-wagging pet resting on your couch could serve as a source for human genetic discoveries? Few people know or even come to think that the genome and diseases of dogs are 95% the same as those of humans. The genetic research conducted by Professor Hannes Lohi at the University of Helsinki brings forwards significant information regarding the eye, bone and neurological diseases of both dogs and humans. The field of study represented by Lohi is promoted by a Europe-wide bio and medical research infrastructure (ELIXIR) for which Finland is a co-founder.
The “Eureka!” moment occurred about ten years ago when research fellow Hannes Lohi pinpointed the epilepsy gene of miniature dachshunds with his research group in Toronto. At the same time elsewhere, the gene was also found in humans. This coincidence was the starting point of the cross-disciplinary canine genetic research led by the professor at the Faculty of Veterinary Medicine and Medicine of the University and Helsinki and the Folkhälsan Research Center. Since 2006, DNA samples from almost 50,000 Finnish dogs have been collected in the DNA bank established by Lohi.
“Dog breeds provide a genetically excellent structure especially for behavioural studies and research into canine and human diseases in general. What animal species is socially gifted, shares the same environment and is exposed to the same pathogens other than man’s best friend?”, Lohi realised at the time.
Lohi noted that inbreeding within dog breeds, in particular, facilitates the identification of disease genes.
“It is easier to discover genes from bloodlines using smaller study cohorts. Compared with the mice and rats typically used in studies, dogs are closer to humans also in terms of vital functions due to their size”, Lohi says.

The spectrum of the canine genetic research led by Lohi is extensive. The subjects include eye diseases, autoimmune diseases, neurological diseases as well as skeletal muscle diseases. The group has identified several new disease genes in dogs from factors causing, for example, epilepsy, dwarfism and anxiety disorders. With the genetic areas found, conditions such as anxiety disorders, from which about 5% of the human population will suffer at some point during their life, gain a new research basis for the study of, for example, the genetic background and environmental factors in obsessive-compulsive behaviours.
“We look for the gene causing a disease in a dog breed and, at the same time, the breed provides a canine model for identifying the disease mechanism of human diseases”, Lohi says, describing the benefits of the research.
The group identified the CNGB1 gene that causes retinal degeneration and, at worst, blindness in Papillon dogs. The same gene has been found in human patients. One in ten people over the age of 65 suffers from this disease during retirement. The condition involves blind spots that limit the area of sharp vision, preventing the renewal of a driving licence, for example.
“With the further development of partially developed drugs, the degeneration of the human retina could be treated externally with gene therapy, for example, by applying to the retina a cream containing viruses carrying normal gene copies that would correct the functioning of the cells and may correct vision”, Lohi describes the possibilities.
“After identifying the gene, it becomes possible to study the disease mechanism and make comparisons between humans and dogs. The gene may not always be the same in humans and the mutation can be located elsewhere, in another gene of the cell pathway. Understanding the gene function and disease mechanism are prerequisites for inventing treatments for the disease. On the other hand, when a mutation is found, it is possible to develop a genetic test for dogs and see which dogs are carriers of the disease. This allows dog breeders to quickly benefit from the research”, Lohi says.
He is involved in Genoscoper Laboratories Ltd, a company that, under his leadership, has built a unique and affordable genome-wide genetic test for dogs, MyDogDNA, which tests the dog’s carrier status for over 100 diseases and traits in one go, as well as genomic diversity and structure.
“The genetic diversity of dogs has been weakened by breeding. The number of dogs carrying disease genes has increased, and because many diseases arise in adulthood, sick dogs will have already been used for breeding. To counter the negatives, breeding may lead to the gene causing the disease becoming more common in a particular dog breed. The candidate gene is more easily identified in dogs than in humans and with fewer samples.”
A large number of veterinarians and dog lovers around Finland have not been enthusiastic about participating in a DNA sampling effort for the benefit of a passing project. The aim of the research group is to build a separate, extensive sequence and variant database for Finnish dogs and cats, similar to the one that already exists for humans (1000 Genomes).
“Genetic research has always been the flagship of Finnish science. We have uniquely accurate health information on patients, including family trees. Equivalent pedigree databases and health data are available on dogs, and soon also on cats. Few countries have such a good, centralised system”, Lohi says.
“There are 400 breeds of dogs. At present, a total of 700 diseases have been depicted in dogs and more are found all the time. The aim is to have a database with the entire genome of each breed sequenced. This will speed up genetic discoveries”, Lohi says.
Lohi believes that the benefit of a large sequence database is a kind of consensus. This is achieved once hundreds or thousands of genomes have been sequenced and the large number of variants can be accurately mapped. There may be many diseases in the same breed.
“For example, if the genomes of 1,000 dogs from 50 breeds have been sequenced into the database, it will include an estimated 25 million variants from the different breeds. The database will facilitate future projects in that a small family of dogs or cats can be studied with just a few of the individual animals sequenced to provide a sufficiently reliable result on the correct disease variant. The variants of a dog patient are compared with the variants of the thousand samples in the database and, if a particular variant is found in the patient but not in the reference samples in the database, it can be inferred to be disease-causing. After this, the matter is confirmed using a larger file.”
“An efficient and nationally significant database will help us catch disease genes faster. As things are now, you have to do a lot work in research to obtain a sufficient picture of the location of a variant in the chromosomes. Going forward, a sample will be taken, the entire genome sequenced and compared directly with the variants in the database.”
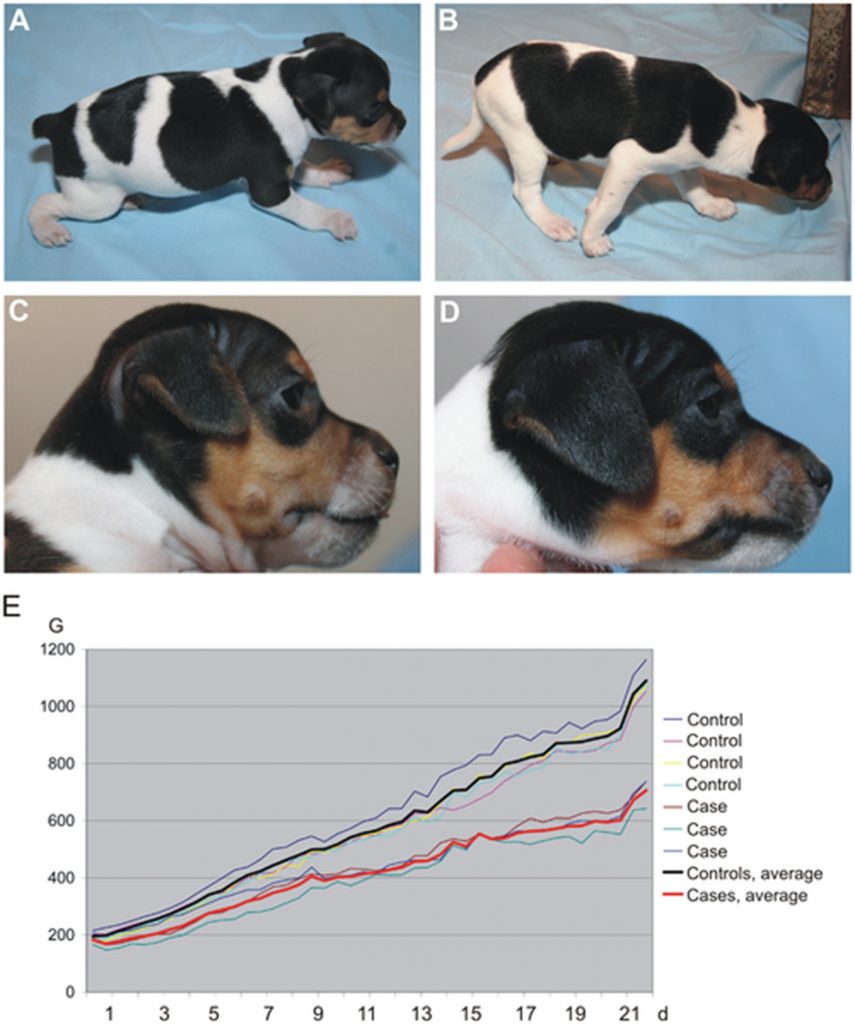
It is estimated that new biotechnical methods will produce a million times the amount of data produced today by 2020. Lohi states that large amounts of computing resources are needed for both the methods and tools used in research.
“Before, short sections of the genome were sequenced. Now, genome lists are so long that managing them manually is completely impossible. If 200 dogs are studied and the entire genome, i.e. 39 pairs of chromosomes, is sequenced from each dog, the analysis would take several months with the traditional method. A single genome affords hundreds of gigabytes of raw data.”
“As we have shifted from the traditional Sanger method of sequencing to Next-Generation Sequencing (NGS) of the entire genome, huge quantities of data are being analysed using new methods. The genome is first split into sections in the database, sequenced and assembled. The sequencing of a genome involves the processing of three billion pairs of genes for humans and 2.5 billion pairs for dogs as well as different variants and insertions that complicate the interpretation of the sequence”, Lohi says, describing the challenges of the research data.
“After the variants have been identified, it is examined whether the variant is pathogenic. Computing resources are required at this stage, too. Bioinformatics tools can be used to predict which amino acid change the variant causes in the genome. After that, the effects of the amino acid change are studied more closely, switching to use protein-level tools and various algorithms.”
The research group pinpointed the gene causing retinal degeneration in Papillon dogs with six sick and 14 control animals. The genetic defect was identified using exome sequencing technology, analysing all of the protein-coding areas at once. Many disease-causing mutations are located in the exome, even though it only accounts for 1.5% of the genome. This technology, which is used especially to find disease forms present in the database, led to the identification of a mutation carried by almost one in five Papillon dogs in their genome.
Lohi’s research group participated as a pilot organisation in a project of CSC – IT Center for Science exploring what kinds of materials are created for researchers with extensive computing capacity and memory space. The aim of the project was to pilot models and solutions for the kinds of resources needed by researchers in the ELIXIR research infrastructure.
Tiina Autio
15.7.2015
Read Aarticle in PDF
Citation
Tiina Autio, Hannes Lohi, & Tommi Nyrönen. (2015). Pups and pooches behind genetic discoveries in human diseases: Canine genetic research benefits from ELIXIR databases. https://doi.org/10.5281/zenodo.7923092
More information:
CSC – IT Center for Science
is a non-profit, state-owned company administered by the Ministry of Education and Culture. CSC maintains and develops the state-owned, centra- lised IT infrastructure.
https://research.csc.fi/cloud-computing
ELIXIR
builds infrastructure in support of the biological sector. It brings together the leading organisations of 21 Euro- pean countries and the EMBL European Molecular Biology Laboratory to form a common infrastructure for biological information. CSC – IT Center for Science is the Finnish centre within this infrastructure.

Science costs money, and the bill is paid by society. However, what happens to the data generated in research? Data is the capital of life sciences, and it should be invested right.
The data of life science research has become a focal point of European science policy in the 21st century. International research uses and produces enormous amounts of data. The confidential storage and further use of the constantly expanding and increasingly complex life science data raise questions.
How and where should the data be stored? How can the data, such as genetic databases, be stored securely? How is the data distributed? The international ELIXIR project is tackling these issues.
ELIXIR provides solutions for how to open up research data in a way that promotes science, and who has access to the data. Finland has been at the core of the ELIXIR project since its inception in 2007. In small European countries, cooperation in data storage and distribution is a prerequisite for competitiveness in life sciences. Small countries should share their once generated materials, rather than producing them repeatedly at different research universities. Life science research requires reference materials, and ELIXIR is a channel for distributing them.
For example, if a Finnish research group studies the heritability of Parkinson’s disease, it is vital that they have access to the genetic material produced by other European research laboratories. Discovering mechanisms is demanding, and researchers need at least a reference point on how a healthy human genome works.
When materials produced by Europeans are available, Finns do not have to build reference material in addition to research sample material. That would be extremely expensive and delay the research results for a decade.
However, using international genetic materials at a Finnish university does not just happen instantly like that. Similarly to when people travel from one country to another, moving data requires infrastructures and agreements between countries. Foreign research universities need to be certain of the identity of the party using the data. Life science data is often associated with information security provided by law. There must also be efficient communications links between the countries through which the huge masses of data can be transmitted. The open Internet broadband is not enough. Furthermore, the recipient must have access to storage space and software with which it can process the material.
The management, storage and distribution of data do not always invoke the same passion and enthusiasm as scientific breakthroughs. Bureaucratic language is deceiving, however. The slowly built network enables scientific breakthroughs, but is also in itself a project comparable to scientific innovation.
The antithesis of international cooperation and data sharing is the huge waste of resources of competing European universities conducting the same basic research. The end result would benefit society significantly less; if everyone was reinventing the genome, research would cost more and produce less. Europeans would drop out of the international competition in life sciences and the bio-industry.
Finland is represented in ELIXIR by CSC – IT Center for Science in cooperation with the Institute for Molecular Medicine Finland (FIMM) of the University of Helsinki and the National Institute for Health and Welfare.
“ELIXIR is a big deal for Finnish life sciences, and it will become even bigger in the future”, says head of the Finnish ELIXIR node Tommi Nyrönen from CSC.
“Through ELIXIR, Finnish biomedicine has access to huge data sets. In future, we will gain more precise information on the rare aberrations in the Finnish genome, for example, as we can compare it to international reference material. This will also allow us to prepare more accurate treatment plans.”
The guiding light of ELIXIR is that research data is the capital of science. Reliable storage and distribution of materials are prerequisites for scientific productivity.
Building and maintaining the infrastructure actually only costs a fraction of the cost of the research itself. Moreover, as for the scientific results that the infrastructure yields – according to Nyrönen – the same party that finances science also benefits from it: society.
“When European genetic researchers are able to use each other’s materials back and forth, the result is more accurate information on the mechanisms of disease and health. In cooperation, the scientific results will also circulate to treatments faster”, says Tommi Nyrönen.
Opening the genome of the Finnish silver birch can be beneficial, for example, in curbing the birch plague epidemic in England. Comparison can be used to identify, for example, why the defence mechanisms of English birches are weaker than those of the silver birch.
The serious illness of a Finnish child acquires a more precise description from a combination of gene mapping and comparative data. When the defective gene is known, planning the treatment can begin. European countries that export wine can cooperate in investigating the genetics of grape diseases and, through breeding, gain competitive advantages in relation to wine producers elsewhere in the world.
ELIXIR also benefits businesses. The project has shared, for example, the canine genome, based on which researchers from the University of Helsinki have developed a commercial application. It allows dog breeders to select the healthiest of their breeder dogs and only breed those that do not carry disease genes, for example, for articular diseases.
Even though ELIXIR encourages cooperation, sharing and openness, the openness of data materials is nonetheless limited. Some of the material is public, some is not. In any case, we are not talking about the open web.
ELIXIR balances between high information security and openness. The most sensitive data in the network is open to those with rights for research purposes. Separate practices have been created between ELIXIR countries for the identification of researchers and granting of “access passes”.
Elina Kuorelahti
5.6.2015
Read article in PDF
Citation
Elina Kuorelahti, & Tommi Nyrönen. (2015). Life science in European cloud. https://doi.org/10.5281/zenodo.8176710